Multi Scale Curriculum CNN for Context-Aware Breast MRI Malignancy Classification
Classification of malignancy for breast cancer and other cancer types is usually tackled as an object detection problem: Individual lesions are first localized and then classified with respect to malignancy. However, the drawback of this approach is …
Authors: Christoph Haarburger, Michael Baumgartner, Daniel Truhn
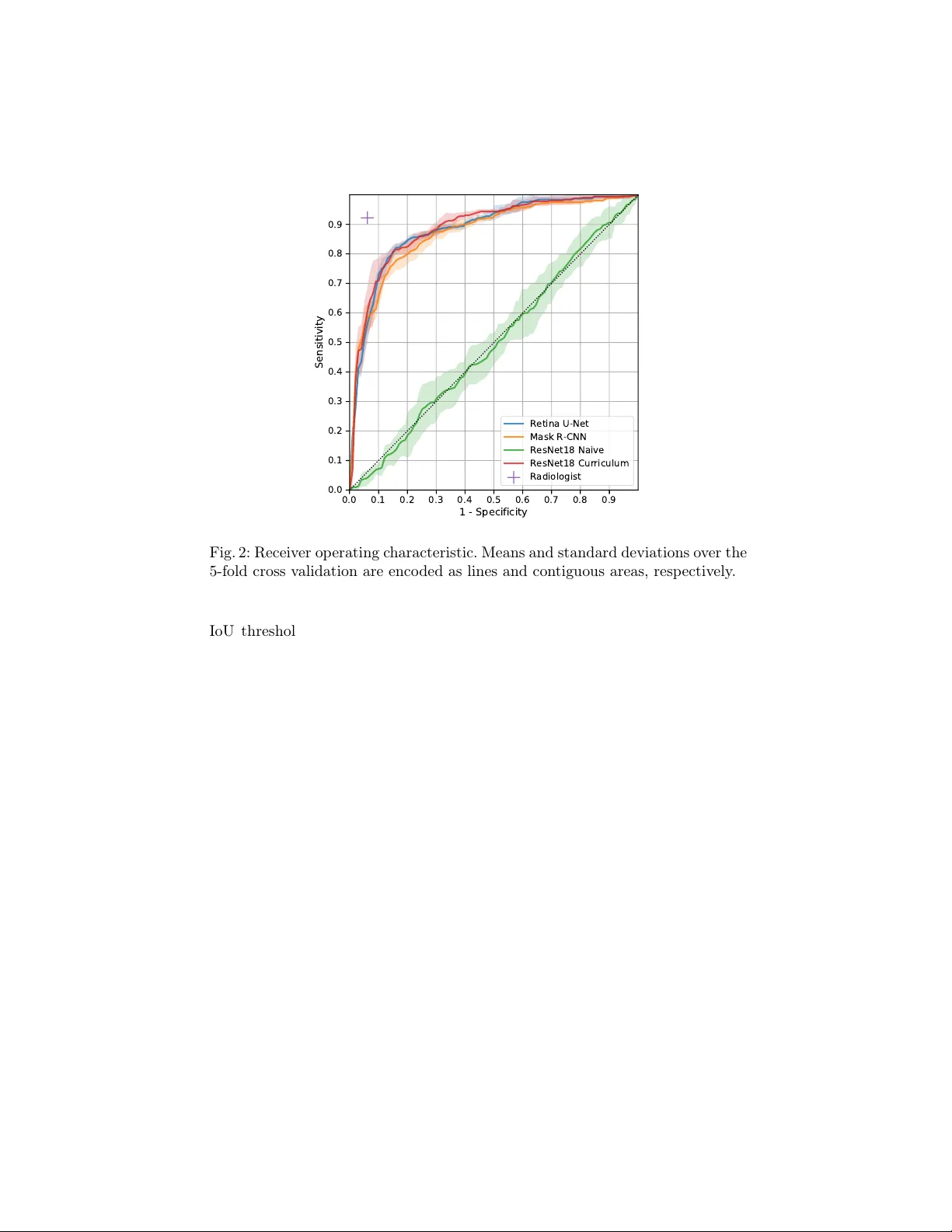
Multi Scale Curriculum CNN for Con text-A w are Breast MRI Malignancy Classification Christoph Haarburger 1 , Mic hael Baumgartner 1 , Daniel T ruhn 2 , 1 , Mirjam Bro ec kmann 2 , Hannah Sc hneider 2 , Simone Sc hrading 2 , Christiane Kuhl 2 , Dorit Merhof 1 1 Institute of Imaging and Computer Vision , R WTH Aac hen Universit y , Germany 2 Dpt. of Diagnostic and In terv. Radiology , Univ ersit y Hospital Aachen, Germany christoph.haarburger@lfb.rwth-aachen.de Abstract. Classification of malignancy for breast cancer and other can- cer t yp es is usually tackled as an ob ject detection problem: Individual lesions are first localized and then classified with respect to malignancy . Ho wev er, the dra wback of this approach is that abstract features in- corp orating sev eral lesions and areas that are not lab elled as a lesion but contain global medically relev ant information are thus disregarded: esp ecially for dynamic contrast-enhanced breast MRI, criteria such as bac kground parenc h ymal enhancemen t and location within the breast are imp ortant for diagnosis and cannot b e captured by ob ject detection approac hes properly . In this w ork, we propose a 3D CNN and a multi scale curriculum learning strategy to classify malignancy globally based on an MRI of the whole breast. Th us, the global context of the whole breast rather than individ- ual lesions is taken into account. Our prop osed approac h does not rely on lesion segmentations, whic h renders the annotation of training data m uch more effectiv e than in curren t ob ject detection approaches. A chieving an AUR OC of 0.89, we compare the performance of our ap- proac h to Mask R-CNN and Retina U-Net as well as a radiologist. Our p erformance is on par with approaches that, in contrast to our metho d, rely o n pixelwise segmentations of lesions. Keyw ords: Breast cancer · Lesion detection · Lesion classification. 1 In tro duction Detection of anatomical structures or lesions in medical images is mostly solv ed b y ob ject detection algorithms that rely on pixelwise segmentations of individual ob jects. How ever, there is not alwa ys a clear consensus among clinicians on which ob ject is considered a suspicious lesion that should b e segmen ted. Moreo ver, seg- men tations are sub ject to high inter-rater v ariability [18]. Both of these aspects limit the qualit y of training data and a reliable and meaningful ev aluation. The degree of natural v ariation cannot b e captured by interpreting tumors as ob jects with hard b oundaries. The latter do es mak e sense in classical computer visi on, but for biomedical applications, a more holistic approach that takes the global 2 C. Haarburger et al. c ontext of a lesion into account is crucial. Another limitation of the detection and instance segmentation approach lies in the high cost asso ciated with lab el- ing large datasets. As a consequence, most datasets are rather small, prev enting that deep learning algorithms unfold their full p otential and limiting the pow er of ev aluation results. Finally yet importantly , clinicians demand a decision sup- p ort at patien t-level rather than a classification of individual ob jects/regions of in terest to efficiently integrate computer-aided diagnosis algorithms into clinical w orkflow. Related W ork. A naiv e approach to incorp orating global context would b e a classification of whole axial slices instead of detecting individual ob jects. Ho w- ev er, this approac h has pro duced po or results so far because it is a needle- in-ha ystack kind of problem [17]. Other works hav e adapted ob ject detection algorithms such as Mask R-CNN [3] and RetinaNet [12] for lesion detection and classification [8]. In [7], diffusion-weigh ted breast MR images are classified for malignancy b y setting all vo xels outside the lesion segmentation to zero. Lotter et al. [13] prop osed a curriculum learning strategy for mammogram classification that concatenates features maps from patc h lev el for a global breast malignancy classification. In a similar approach K oshrav an et al. [9] prop ose a detection and malignancy classification algorithm for lung nodules by sup erimp osing a grid on CT scans and by classifying each grid cell as b enign or malignant. Maicas et al. [16] proposed a reinforcement learning approach for breast MRI lesion detection using b ounding b oxes. Sev eral other authors hav e prop osed metho ds for lesion classification based on bounding b oxes [1,2,21]. In [15], a curriculum learning strategy is prop osed and extended to classification of whole v olumes in [14] sho wing promising results. Zhu et al. [22] prop ose a m ultiple instance learning loss for whole mammogram classification. In [20], a multi scale CNN for detection of lesions in breast ultrasound images is prop osed. Finally JÃ ďger et al. [6] prop osed Retina U-Net, whic h adv ance s RetinaNet [12] by lev eraging segmen tation supervision for breast lesions. W e introduce a CNN that is trained on several scales in a curriculum learning strategy , whic h is highly efficient for small datasets. Moreov er, our 3D CNN does not rely on pixelwise segmentations or b ounding b oxes but rather lesion cen ter p oints. This allo ws for efficient anno- tation and lev erages the whole lesion context for classification at patien t level. Con tributions. W e introduce a simple 3D CNN for breast cancer malignancy classification that do es not need to detect lesions individually and do es not rely on pixelwise segmen tations or b ounding b oxes. Moreov er, we prop ose a multi scale curriculum learning strategy that efficien tly trains the net w ork on tw o scales, starting on patc h scale and contin uing training on the whole breast in- cluding all global con text. W e pro vide a comparison with other state of the art metho ds: Naive whole breast classification, Mask-R-CNN and Retina U-Net. Finally , in order to maximize repro ducibilit y , comparisons and adaption, w e pro- vide all co de 1 including implementations of netw ork arc hitectures, prepro cessing and curriculum training in PyT orc h. 1 https://github.com/haarburger/multi- scale- curriculum Curriculum CNN for Brea st MRI Malignancy Classification 3 2 Image Data Our dataset consists of dynamic contrast-enhanced (DCE) MR images of 408 patien ts from clinical routine at our institution. Images w ere acquired on a 1.5 T Philips Scanner using a standardized proto col consisting of a T2-weigh ted turb o spin ec ho sequence and a T1-w eighted gradient ec ho sequence acquired as a dynamic series [10]. DCE T1-weigh ted images were acquired b efore and every 70 s after injection of contrast agen t (gadobutrol) for four p ost-contrast agen t time points. The acquisition matrix was 512 × 512 , yielding an in-plane resolution of 0 . 6 × 0 . 6 mm and slic e thic kness was 3 mm. In order to allow a comparison of our approach with approac hes that rely on an auxiliary segmen tation such as Mask R-CNN and Retina U-Net, all lesions were manually segmen ted on every slice b y a radiologist with 13 years of experience in breast MRI. Out of the 408 patients, 305 had malignant and 103 had b enign findings. Malignancy w as determined b y biopsy; diagnoses of b enign lesions were v alidated b y follow up for 24 mon ths. Since man y of the malignant findings w ere only presen t in one breast, the ov erall ratio of malignant and benign samples at breast lev el (rather than patient lev el) in the whole dataset is 40.4%/59.6%. All images w ere resampled to 512 × 512 × 32 vo xels. No bias field correction was p erformed. Finally , we cropp ed all images by removing v oxels that only contain air or tissue from the thorax, leading to images of a spatial resolution of 512 × 256 × 32 v oxels. 3 Metho ds 3.1 Multi Scale Curriculum Netw ork The netw ork architecture w e prop ose consists of tw o basic comp onents as de- picted in Fig 1: 1) A Backbone net work that consists of a 3D CNN to generate feature maps. 2) Classification head that p erforms a classification based on the aggregated feature maps pro vided b y the t wo previous comp onents. F or the Back- b one we initially ev aluated man y different ac hitectures including ResNets [4] and DenseNets [5] of differen t depths, U-Net [19] and F eature Pyramid Net work [11]. Since choice of the backbone architecture did not hav e a significan t impact on the o verall performance for the final model, we focus on ResNet18 in this w ork, b ecause of its efficiency . Multi Scale Curriculum T raining. T raining is conducted in t wo stages: – Stage 1: Classification of 3D lesion p atches with size 64 × 64 × 4 , where each patc h con tains at least one lesion as determined by the lo cation of the lesion cen terp oints. If at least one malignant lesion is contained in the patch, the patc h lab el is set to malignant and benign otherwise. – Stage 2: Classification of 3D v olumes con taining a whole breast with size 256 × 256 × 32 . T o account for the changed input size, the net work arc hitec- ture is modified b y in tro ducing an additional adaptive a verage p o oling la y er b et ween the original a verage po oling and fully-connected la yer. All net work parameters are trained during Stage 2. 4 C. Haarburger et al. C 1 C 2 C 3 C 4 C 5 S t a g e 1 S t a g e 2 S t a g e 1 S t a g e 2 M a l i g n a n c y ? M a l i g n a n c y ? 6 x 6 4 x 6 4 x 4 6 x 2 5 6 x 2 5 6 x 3 2 1 4 4 x 1 x 1 x 1 1 4 4 x 4 x 4 x 8 1 4 4 x 1 x 1 x 1 1 8 1 8 3 6 7 2 1 4 4 Fig. 1: Netw ork architecture: Residual Blo cks are named C i as in [4] with c hannel dimensions indicated at the b ottom of each blo ck. Con volutions are indicated in blue, p o oling op erations in red. F rom all MRI scans, left and righ t breast were fed into the netw ork separately . The fiv e time p oints from the dynamic T1-w eigh ted series and the T2-weigh ted series w ere fed into the CNN in the channel dimension, whic h leads to an input v olume of 6 × 256 × 256 × 32 ( channels, x, y , z). The net work was trained with batc h size 4, Adam optimizer with default parameters, instance normalization, leaky ReLU activ ation functions and a learning rate of 10 − 4 and 10 − 5 for Stages 1 and 2, resp ectiv ely . F or data augmen tation, we mirrored all images and rotated around the z axis by ± 15 ◦ . T raining, hyperparameter tuning and testing w ere p erformed in a 5-fold cross v alidation, using 261 patien ts (64%), 65 patien ts (16%) and 82 patien ts (20%) of the data for training, v alidation and testing, resp ectively . Since eac h single breast is fed into the netw ork separately , splitting at patien t level guaran tees that b oth breasts of a single patient are exclusiv ely contained in only one of the three data subsets. 3.2 Comparison Metho ds Naiv e ResNet. In this approach, we naively train a v anilla 3D ResNet18 to directly predict malignancy globally without multi scale curriculum learning. Since only a very small fraction of v o xels b elongs to a lesion, this approac h is a needle-in-a-ha ystack kind of problem. It serves as a baseline for the multi scale curriculum learning approac h. Mask R-CNN. Mask R-CNN [3] is a widely used tw o-stage detection framew ork that lev erages sup ervision information from a segmentation auxiliary task. Retina U-Net. Based on the RetinaNet one stage detection framework [12], Retina U-Net combines RetinaNet with enhanced segmen tation supervision us- ing the U-Net arc hitecture. Curriculum CNN for Brea st MRI Malignancy Classification 5 4 Results On a Nvidia GeF orce 2080 Ti GPU, training of Stage 1 and 2 takes ab out 4 and 6 hours, resp ectiv ely . Prediction of a single breast can b e p erformed in under 100 ms. In order to sho w the b enefit of multi scale curriculum learning, we ev alu- ated Mask R-CNN and Retina U-Net as described ab ov e as well as a naiv e 3D ResNet18 without curriculum training (Stage 2 only) and the prop osed approac h with multi scale curriculum learning. In addition, a radiologist with 2 years of exp erience in breast MRI rated the images with resp ect to malignancy . T est p erformance is pro vided in T ab. 1. The highest AUR OC is ac hieved by the ra- diologist, follow ed by ResNet18 Curriculum and Retina U-Net. Mask R-CNN p erforms slighly worse. The naive 3D image classification approach achiev ed a v ery po or p erformance that is not significantly differen t from random guessing. A UROC A ccuracy #P arameters Mask R-CNN [3] 0.88 ± 0.01 0.77 ± 0.03 3.91M Retina U-Ne t [6] 0.89 ± 0.01 0.82 ± 0.02 3.90M ResNet18 Naive 0.50 ± 0.04 0.45 ± 0.05 2.66M ResNet18 Curricul um 0.89 ± 0.01 0.81 ± 0.02 2.66M Radiologist 0.93 0.93 - T able 1: T est p erformance of the comparison metho ds and prop osed approac h o ver a 5-fold cross v alidation. In Fig. 3, class activ ation maps for ResNet18 Curriculum are shown. Regions with high activ ations for malignancy are mark ed in red, in order to pro vide clinicians with enhanced guidance b y our algorithm. 5 Discussion Our prop osed multi scale curriculum training enabled successful training of the relativ ely simple 3D ResNet18 from an AUR OC of 0.5 to 0.89. In other experi- men ts that are out of the scop e of this w ork, we found that the particular c hoice of the backbone architecture had negligible effect on the p erformance. On our dataset, the p erformance of our approach is on par with Retina U-Net [6] and ev en exceeds the p erformance of Mask R-CNN [3]. Since w e did not tune our mo dels extensiv ely with resp ect to ensembling, the performance in comparison with the rep orted results in [6] seem reasonable to us. W e h yp othesize that the high p erformance of our model at least partly arises from the high amount of con text and global information that is pro vided. Both Retina U-Net and Mask R-CNN incorp orate more mo del parameters and hyperparameters such as the 6 C. Haarburger et al. 0.0 0.1 0.2 0.3 0.4 0.5 0.6 0.7 0.8 0.9 1 - Specificity 0.0 0.1 0.2 0.3 0.4 0.5 0.6 0.7 0.8 0.9 Sensitivity Retina U-Net Mask R-CNN ResNet18 Naive ResNet18 Curriculum Radiologist Fig. 2: Receiver operating characteristic. Means and standard deviations o ver the 5-fold cross v alidation are encoded as lines and contiguous areas, respectively . IoU threshold. Most imp ortantly , these approac hes rely on pixelwise segmen- tations for all individual lesions. Our approach on the other hand only needs a coarse lo calization (i.e. co ordinate) for Stage 1 training and only one global lab el per breast for Stage 2. Since in most clinical sites p erforming breast MRI, a global BIRADS classification p er breast is assessed and provided along with the image data, global lab els are relatively c heap to generate. This w ould allow a Stage 2 training with m uch more training data and more thorough ev aluation with muc h more test data in the future. Curriculum learning on a series of tasks as proposed in [15] ac hieved v ery similar performance on a breast MRI dataset. Our approach has some similarities with [13]: Stage 1 training is v ery similar, yet p erformed in 3D in our case. In Stage 2, Lotter et al. [13] concatenate and p o ol feature vectors of individual (2D) regions while w e perform direct predictions of whole images in 3D. Our wor k has sev eral limitations: Firstly , our dataset is mono cen tric and ev en though to our best knowledge it is the largest breast MRI dataset for computer aided diagnosis, it is limited in size. How ever, extension is relatively easy since only global lab els are required for Stage 2 training and assessment of test p erformance. F or Stage 1 a coarse lo calization that has to be mark ed man ually is still required. In comparison to the diagnostic performance of a breast radiologist, our mo del is inferior to h uman p erformance. F or future work, we will expand the size of our Stage 2 dataset. Since breast MRI e xperts are rare, it would b e in teresting to assess whether our algorithm Curriculum CNN for Brea st MRI Malignancy Classification 7 (a) (b) (c) (d) Fig. 3: Class activ ation maps for (a): Malignant case that was correctly classified b y our algorithm and the radiologist. (b): Benign case that was classified cor- rectly by the CNN and incorrectly b y the radiologist. (c): Malignant case that w as classified incorrectly b y b oth CNN and radiologist. (d): Malignan t case that w as classified incorrectly by the CNN and correctly b y the radiologist. Images ha ve been masked. (including the activ ation maps) can b e applied to teach radiologists with limited exp erience. Moreov er, it would b e interesting to assess whether a combination of an exp ert rater with our algorithm could further improv e the ov erall p erfor- mance. 6 Conclusion W e presen ted a simple 3D CNN arc hitecture that is able to p erform breast cancer malignancy classification based on magnetic resonance images using a m ulti scale curriculum learning strategy . The netw ork architecture predicts malignan t breast cancer globally for a whole 3D v olume without scoring individual lesions. This approac h pro vides the whole spatial context of a breast to the net w ork, yielding state of the art p erformance and without the need for lesion segmentations. A c knowledgemen ts The authors thank Paul JÃ ďger for making Medical Detection T o olkit [6] pub- licly a v ailable, whic h w as a great help when comparing our method with Mask R-CNN and Retina U-Net. References 1. Amit, G., Hadad, O., Alp ert, S., Tlusty , T., Gur, Y., Ben-Ari, R., Hashoul, S.: Hybrid mass detection in breast mri combining unsup ervised saliency analysis and deep learning. In: MICCAI. pp. 594–602 (2017) 2. Dalmış, M.U., V reemann, S., K o oi, T., Mann, R.M., Karssemeijer, N., Gub ern- Mérida, A.: F ully automated detection of breast cancer in screening MRI using con volutional neural net works. Journal of Medical Imaging 5 (01), 1 (jan 2018) 8 C. Haarburger et al. 3. He, K., Gkioxari, G., Dollár, P ., Girshic k, R.: Mask R-CNN. In: ICCV. pp. 2961 – 2969 (2017) 4. He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: (CVPR) (2016) 5. Huang, G., Liu, Z., W einberger, K.Q., v an der Maaten, L.: Densely Connected Con volutional Netw orks. In: CVPR. pp. 4700 – 4708 (2017) 6. Jaeger, P .F., Kohl, S.A.A., Bick elhaupt, S., Isensee, F., Kuder, T.A., Schlem- mer, H.P ., Maier-Hein, K.H.: Retina U-Net: Embarrassingly Simple Exploita- tion of Segmentation Supervision for Medical Ob ject Detection (2018), http: 7. Jäger, P .F., Bic kelhaupt, S., Laun, F.B., Lederer, W., Heidi, D., Kuder, T.A., P aech, D., Bonek amp, D., Radbruch, A., Delorme, S., Sc hlemmer, H.P ., Steudle, F., Maier-Hein, K.H.: Rev ealing hidden p otentials of the q-space signal in breast cancer. In : (MICCAI). pp. 664–671 (2017) 8. Jung, H., Kim, B., Lee, I., Y o o, M., Lee, J., Ham, S., W o o, O., Kang, J.: Detection of masses in mammogram s using a one-stage ob ject detector based on a deep con volutional neural net work. PLOS ONE 13 (9), 1–16 (09 2018) 9. Khosra v an, N., Bagci, U.: S4ND: Single-Shot Single-Scale Lung No dule Detection. In: MICCAI. pp. 794–802 (2018) 10. Kuhl, C.K.: The Curren t Status of Breast MR Imaging Part I. Choice of T ec hnique, Image Interpretation, Diagnostic A ccuracy , and T ransfer to Clinical Practice. Ra- diology 244 (2), 356–378 (2007) 11. Lin, T.Y., Dollár, P ., Girshick, R., He, K., Hariharan, B., Belongie, S.: Feature Pyramid Net w orks for Ob ject Detection. In: CVPR. pp. 2117–2125 (2017) 12. Lin, T.Y., Goy al, P ., Girshick, R., He, K., Dollár, P .: Fo cal Loss for Dense Ob ject Detection. In: ICCV. pp. 2980 – 2988 (2017) 13. Lotter, W., Sorensen, G., Co x, D.: A Multi-Scale CNN and Curriculum Learning Strategy for Mammogram Classification. In: MICCAI W orkshop DLMIA (2017) 14. Maicas, G., Bradley , A.P ., Nascimento, J.C., Reid, I., Carneiro, G.: Pre and Post- ho c Diagnosis and Interpretation of Malignancy from Breast DCE-MRI (2018), 15. Maicas, G., Bradley , A.P ., Nascimento, J.C., Reid, I., Carneiro, G.: T raining Med- ical Im age Analysis Systems lik e Radiologists. In: MICCAI. pp. 546–554 (2018) 16. Maicas, G., Carneiro, G., Bradley , A.P ., Nascimento, J.C., Reid, I.: Deep rein- forcemen t learning for active breast lesion detection from dce-mri. In: MICCAI. pp. 665–6 73 (2017) 17. Maier-Hein, K.: h ttp://on-demand.gputechconf.com/gtc-eu/2018/video/e8481/ accessed 2019-03-06 18. Menze, B.H., Jak ab, A., et al.: The multimodal brain tumor image segmentation b enc hmark (BRA TS). IEEE TMI 34 (10), 1993–2024 (10 2015) 19. Ronneb erger, O., Fischer, P ., Bro x, T.: U-net: Conv olutional netw orks for biomed- ical ima ge segmen tation. In: MICCAI. pp. 234–241 (2015) 20. W ang, N., Bian, C., W ang, Y., Xu, M., Qin, C., Y ang, X., W ang, T., Li, A., Shen, D., Ni, D.: Densely deep supervised net works with threshold loss for cancer detection in automated b reast ultrasound. In: MICCAI. pp. 641–648 (2018) 21. Y an, K., Bagheri, M., Summers, R.M.: 3d context enhanced region-based con vo- lutional neural netw ork for end-to-end lesion detection. In: MICCAI. pp . 511–519 (2018) 22. Zh u, W., Lou, Q., V ang, Y.S., Xie, X.: Deep m ulti-instance netw orks with sparse lab el assignment for whole mammogram classification. In: MICCAI. pp. 603–611 (2017)
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment