An innovating Statistical Learning Tool based on Partial Differential Equations, intending livestock Data Assimilation
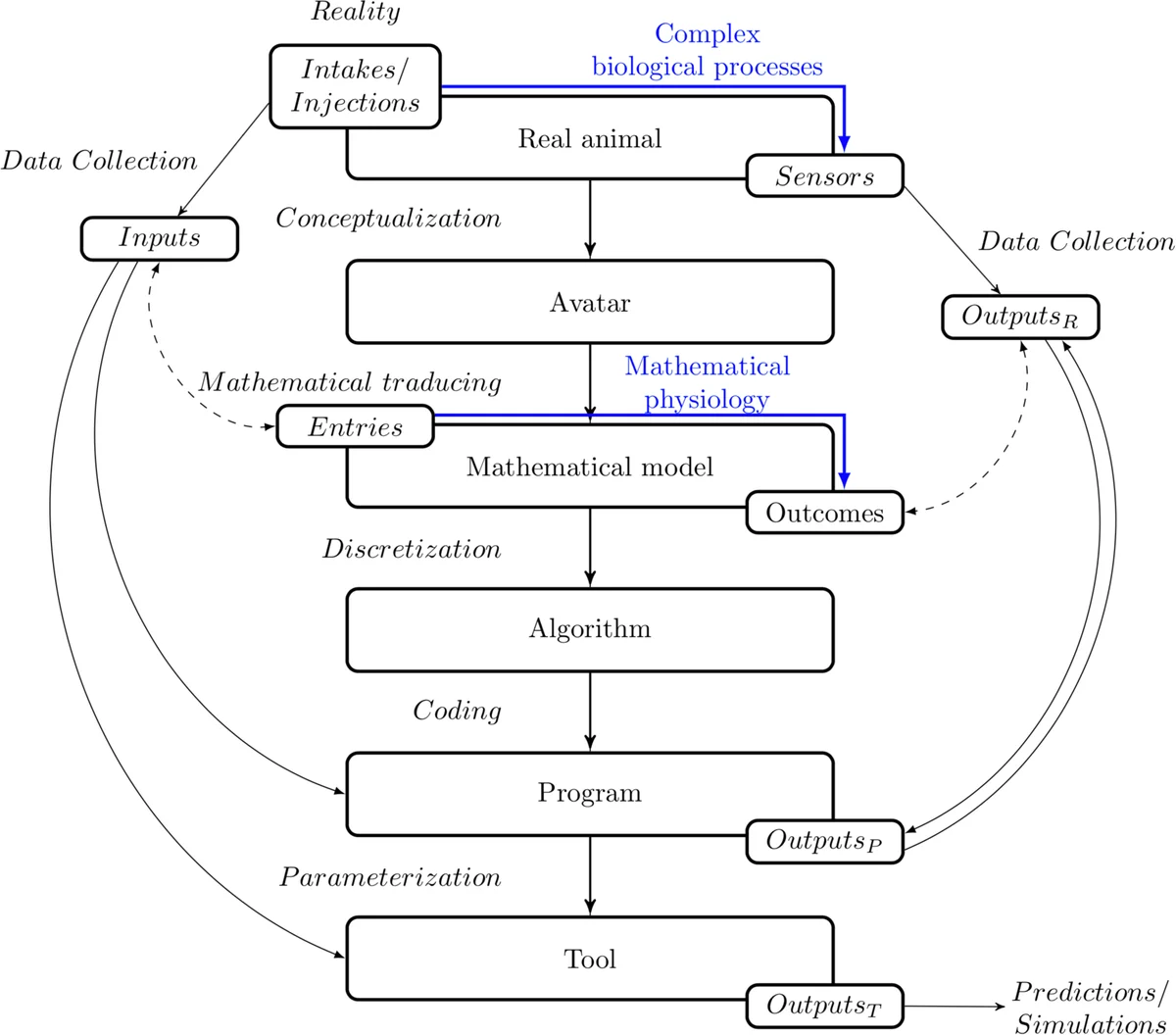
The realistic modeling intended to quantify precisely some biological mechanisms is a task requiering a lot of a priori knowledge and generally leading to heavy mathematical models. On the other hand, the structure of the classical Machine Learning algorithms, such as Neural Networks, limits their flexibility and the possibility to take into account the existence of complex underlying phenomena, such as delay, saturation and accumulation. The aim of this paper is to reach a compromise between precision, parsimony and flexibility to design an efficient biomimetic predictive tool extracting knowledge from livestock data. To achieve this, we build a Mathematical Model based on Partial Differential Equations (PDE) embarking the mathematical expression of biological determinants. We made the hypothesis that all the physico-chemical phenomena occurring in animal body can be summarized by the evolution of a global information. Therefore the developed PDE system describes the evolution and the action of an information circulating in an Avatar of the Real Animal. This Avatar outlines the dynamics of the biological reactions of animal body in the framework of a specific problem. Each PDE contains parameters corresponding to biological-like factors which can be learnt from data by the developed Statistical Learning Tool.
💡 Research Summary
The paper addresses the need for accurate, parsimonious predictive tools in livestock production, where data are often scarce and biological processes are complex. Traditional approaches fall into two categories: mechanistic models that require extensive prior knowledge and result in heavy mathematical formulations, and black‑box machine‑learning models that demand large datasets to achieve acceptable performance. To bridge this gap, the authors propose a “Data‑Model Coupling” framework that embeds a lightweight, parameterized partial differential equation (PDE) system within a statistical learning pipeline.
The core concept is the construction of an “Avatar” – an abstract representation of the animal that captures the global dynamics of information (nutrients, drugs, metabolites) circulating through the body. Two density fields, a forward flux Φ_f and a backward flux Φ_b, evolve over a one‑dimensional spatial domain
Comments & Academic Discussion
Loading comments...
Leave a Comment