Visualisation of Brain Statistics with R-packages ggseg and ggseg3d
There is an increased emphasis on visualizing neuroimaging results in more intuitive ways. Common statistical tools for dissemination, such as bar charts, lack the spatial dimension that is inherent in neuroimaging data. Here we present two packages …
Authors: Athanasia M. Mowinckel, Didac Vidal-Pi~neiro
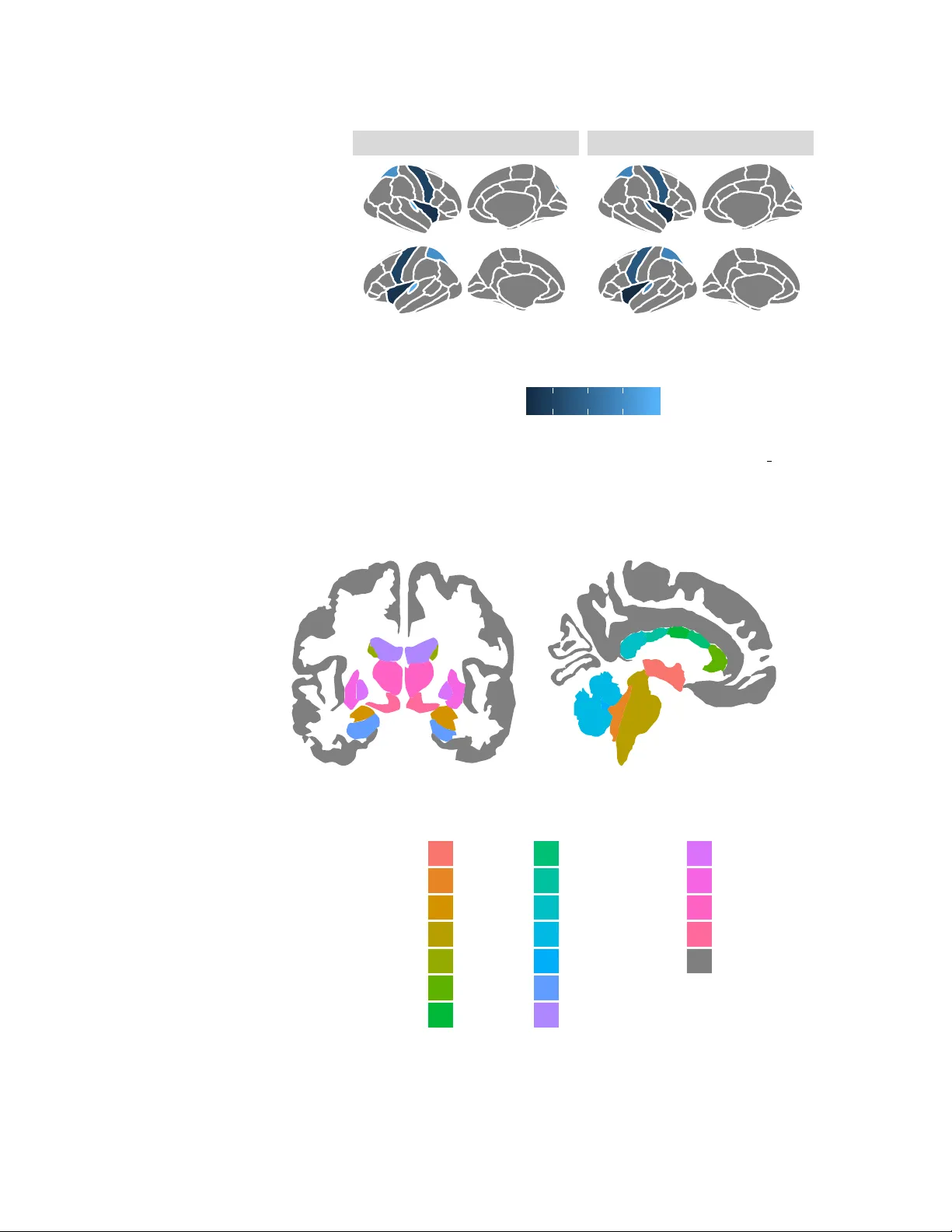
Visualisation of Brain Statistics with R-pac kages ggseg and ggseg3d Athanasia M. Mowinc kel 1 and Didac Vidal-Pi ˜ neiro 1 1 Center for Lif espan Changes in Brain and Cognition, University of Oslo, PO . box 1094 Blindern, 0317 Oslo, Norway Corresponding author : Athanasia M. Mowinc kel 1 Email address: a.m.mowinckel@psykologi.uio.no ABSTRA CT There is an increased emphasis on visualizing neuroimaging results in more intuitive wa ys. Common statistical tools for dissemination, such as bar char ts, lack the spatial dimension that is inherent in neuroimaging data. Here we present two pac kages for the statistical software R, ggseg and ggseg3d , that integrate this spatial component. The ggseg and ggseg3d packages visualize pre-defined brain segmentations as both 2D polygons and 3D meshes, respectiv ely . Both packages are integr ated with other well-established R-pac kages, allowing g reat flexibility . In this tutorial, we present the main data and functions in the ggseg and ggseg3d packages f or brain atlas visualization. The main highlighted functions are able to display brain segmentation plots in R. Fur ther, the accompanying ggsegExtra -package includes a wider collection of atlases, and is intended for community-based efforts to dev elop more compatible atlases to ggseg and ggseg3d . Over all, the ggseg -packages facilitate parcellation-based visualizations in R, improv e and ease the dissemination of the results, and increase the efficiency of the workflows. 1 INTRODUCTION V isualization is increasingly important for accurate guidance and interpretation of neuroimaging results, as current research is able to generate a high amount of data and outcomes. For Magnetic Resonance Imaging (MRI), neuroimaging software provides whole-brain information by using many small units of space ( > 100.000). Nonetheless, this data is often grouped and summarized into a limited number of regions using predefined brain parcellation atlases. Brain parcellations segment the brain into a finite set of meaningful neurobiological components, which reflect one or more brain features either based on structural or connectivity properties (Eickhoff et al. [2018]). The use of brain atlases is widespread as these facilitate interpretation and minimize the amount of data, hence reducing problems with multiple comparisons. This enables replicability and data sharing in otherwise computationally expensi ve analyses, which are often performed in specialized software en vironments such as R (R Core T eam [2019]). MRI data provides good spatial resolution and thus an optimal representation has to respect spatial relationships across regions. Results from brain atlas analyses are most meaningfully visualized when projected onto a representation of the brain, thus it is desirable that any visual representation tak es this relation into account. The projection of data onto brain representations provides clear points of reference - especially when the reader is unfamiliar with the atlas - eases readability , guides interpretation, and con ve ys the spatial patterns of the data. Adopting the grammar of graphics implemented in ggplot2 (W ickham [2016]), one can plot neuroimaging data directly in R with se veral tools such as ggBrain (Fisher [2019]) and ggneuro (Muschelli [2017]; see neuroconductor [2018] for curated neuroimaging packages for R). Y et these tools display whole-brain image files and are not well-suited for representing brain atlas data. In this tutorial, we introduce two packages for visualizing brain atlas data in R. The ggse g and g gse g3d – pluss the complimentary ggse gExtr a – packages include pre-compiled data sets for different brain atlases that allow for 2D and 3D visualization. The two-dimensional functionality in ggse g is based on polygons and ggplot2 -based grammar of graphics (W ickham [2016]), while the 3D functionality in ggse g3d is T able 1. T able of currently av ailable atlases in the ggseg, ggseg3d, and the ggsegExtra R-packages. Polygon and mesh refer to 2D and 3D brain atlas representations, respectiv ely T itle Item Mesh Polygon citation Automated probabilistic reconstruction of white-matter pathways tracula None ggsegExtra Y endiki et al. [2011] Desikan-Killiany Cortical Atlas dkt ggseg3d ggseg Desikan et al. [2006] Desterieux cortical parcellations desterieux ggsegExtra None Destrieux et al. [2010] Freesurfer automatic subcortical segmentation of a brain v olume aseg ggseg3d ggseg Fischl et al. [2002] Genetic topography of brain area morphology chenAr None ggsegExtra Chen et al. [2013] Genetic topography of brain thickness morphology chenTh None ggsegExtra Chen et al. [2013] Harvard-Oxford Cortical atlas hoCort None ggsegExtra Makris et al. [2006] HPC - Multi-modal parcellation of human cerebral cortex glasser None ggsegExtra Glasser et al. [2016] ICBM white matter parcellation icbm ggsegExtra None Mori et al. [2005] JHU parcellation jhu None ggsegExtra Hua et al. [2008] Local-Global 17 Parcellation of the Human Cerebral Cortex schaefer17 ggsegExtra None Schaefer et al. [2017] Local-Global 7 Parcellation of the Human Cerebral Cortex schaefer7 ggsegExtra None Schaefer et al. [2017] Parcellation from JHU jhu ggsegExtra None Hua et al. [2008] Parcellation from the Human Connectome Project glasser ggsegExtra None Glasser et al. [2016] White matter tract parcellations tracula ggsegExtra None Y endiki et al. [2011] Y eo 17 Resting-state Cortical Parcellations yeo17 ggsegExtra ggsegExtra Y eo et al. [2011] Y eo 7 Resting-state Cortical Parcellations yeo7 ggsegExtra ggsegExtra Y eo et al. [2011] based on tri-surface mesh plots and plotly (Sie vert [2018]). Both packages present compiled data sets, tailored functions that allow brain data integration and plotting, and other minor features such as custom colour palettes. The data featured in the packages are de- riv ed from two well-known parcellations: the Desikan-Killany cortical atlas (DKT ; Desikan et al. [2006]), which cov ers the cortical surface of the brain, and the Automatic Segmentation of Subcortical Structures (aseg; Fischl et al. [2002]), which covers the subcortical structures. Both atlases are implemented in sev eral neuroimaging softwares, such as FreeSurfer (Fischl et al. [1999], Dale et al. [1999], Fischl and Dale [2000]), and are commonly used in relation to developmental changes, disease biomarkers, genomic data, and cognition (Amlien et al. [2019], W alhovd et al. [2005], Pizzagalli et al. [2009]). The ggse gExtra package contains a collection of precompiled atlases (currently 15 additional atlases) and it is frequently updated. A summary of all a vailable atlases compatible with the ggse g -packages as of December 2019 can be found in T able 1. 2 TUT ORIAL This tutorial will introduce the ggse g , ggse g3d , and ggse gExtra packages and familiarize the reader with the main functions and the general use of the packages. The tutorial will focus on the two main functions: ggseg() for plotting 2D polygons and ggseg3d() for plotting 3D brains based on tri-surf ace mesh plots. 2.1 Plotting pol ygon data (ggplot2) ggse g is the main function for plotting 2D data. By default, the function automatically plots the DKT atlas (see Figure 1). The ggseg() -function is a wrapper for geom polygon() from ggplot2 , and it can be built upon and combined like an y ggplot -object. The image plot consists of a simple brain representation containing no extra information. Hence, ggse g plots can be easily complemented with an y of the a vailable ggplot2 features and options. W e recommend users to get familiarized with ggplot2 (W ickham [2016]). The package is currently only av ailable through github, but we expect to submit the ggse g -package to The Comprehensiv e R Archiv e Network (Hornik [2012]) in 2020. # remotes::install github("LCBC-UiO/ggseg") library (ggseg) library (dplyr) library (tidyr) # Figure 1 ggseg () 2/17 left right hemisphere Figure 1. By default ggseg() will plot the dkt atlas in grey shaded polygons. In addition to the standard options for ggplot2 polygon geoms, the function also has se veral options for plotting the main brain representations. These options are atlas-specific. For cortical atlases, such as the dkt , one can stack the hemispheres, display only the medial or lateral side, choose either one or both hemispheres, or any combination of hemisphere and vie w (see Figure 2 for e xamples). For subcortical atlases, such as the aseg atlas, the options are more limited but one can often choose between axial, sagittal, and coronal views. # dkt dark theme p1 <- ggseg ( position = "stacked" ) + theme dark () + labs ( title= " " ) # dkt classic theme p2 <- ggseg ( position = "stacked" ) + theme classic () + labs ( title = " " ) # dkt medial view med <- ggseg ( view = "medial" ) + labs ( title = " " ) # dkt left hemisphere left <- ggseg ( hemisphere = "left" ) + labs ( title = " " ) # aseg default theme p3 <- ggseg ( atlas= aseg) + labs ( title = " " ) # dkt left medial alone combo <- ggseg ( view = "medial" , hemisphere = "left" ) + labs ( title= " " ) # Combine plots to Figure 2 cowplot :: plot grid (p1, med, combo, p2, left, p3, labels = c ( "A: dkt - dark" , "B: dkt - medial" , "C: dkt - combo" , "D: dkt - classic" , "E: dkt - left" , "F: aseg" ), hjust = -.05 ) 2.1.1 Using o wn data with fill and colour ggseg() accepts any argument you can supply to geom polygon() , and therefore is easy to work with for those familiar with ggplot2 functionality . Standard arguments like fill that floods the segments with a colour , or colour that colours the edges around the segments are typical arguments to provide to the function either as a single setting value or within the ggplot2 mapping function aes . T o use color 3/17 left right lateral medial side hemisphere A: dkt − dark left right hemisphere B: dkt − medial left hemisphere C: dkt − combo left right lateral medial side hemisphere D: dkt − c lassic left hemisphere E: dkt − left left midline right hemisphere F: aseg Figure 2. ggse g plots can be used with any ggplot2 feature such as standard scales and themes. For cortical atlases, one can supply special ggse g options to determine, for example, hemisphere, view , or position. A: dkt atlas, stacked with dark theme ; B: dkt with medial view only; C: dkt atlas with only left medial display; D: dkt atlas, stacked, with classic theme; E: dkt atlas with left hemisphere only; F: complete aseg atlas 4/17 left right hemisphere area banks superior temporal caudal anterior cingulate caudal middle frontal cuneus entorhinal frontal pole fusiform inferior parietal inferior temporal insula isthmus cingulate lateral occipital lateral orbitofrontal lingual medial orbito frontal middle temporal para central parahippocampal pars opercularis pars orbitalis pars triangularis pericalcarine post central posterior cingulate pre central precuneus rostral anterior cingulate rostral middle frontal superior frontal superior parietal superior temporal supramarginal temporal pole transverse temporal NA Figure 3. Supplying ‘area’ to the fill option in ggseg() , will use the column ‘area’ from the accompanying dataset to create a discrete colour palette o ver the segments in the atlas. The dkt atlas default palette corresponds to the FreeSurferColorLut scheme. palettes corresponding to those used in the original neuroimaging softwares one can use atlas-specific ‘brain’ palette scales (Figure 3). # Figure 3 ggseg ( mapping= aes ( fill = area), colour= "black" ) + scale fill brain ( "dkt" ) + theme ( legend.justification= c ( 1 , 0 ), legend.position= "bottom" , legend.text = element text ( size = 5 )) + guides ( fill = guide legend ( ncol = 3 )) Most users will use ggseg() to display - using a color scale - some descripti ve or inferential statistics, such as mean thickness or brain-cognition relationships across the different brain regions. Y et, before projecting the statistics onto the segments, one should explore the structure of the atlas data sets. The atlas data set structure will help users understand what incoming statistical data needs to look like. Note that each atlas corresponds to a unique data set. All data sets hav e a similar structure and contain key information regarding the atlas, the re gion names, and the coordinates for the segment polygons. # Look at the top 5 rows of the dkt dataset head (dkt, 5 ) In any atlas, the column ‘label’ is particularly useful for combining the data of interest with the ggse g -polygons. The column ‘label’ contains the label (re gion) names as in the original neuroimaging software. For example, the DKT atlas label column matches the region names from Freesurfer statistics table outputs. Y et, the data in ggse g is in a long format - that is each region has a single row - and any 5/17 data of interest needs to be in this same format. Often data sets are organized in wide format, in which subjects are represented by rows and each different data v ariable is represented in a separate column, and thus need to be rearranged to work with ggseg. See below an e xample of wide-to-long con version. # Create a mock freesurfer output freesurfer stats <- data.frame ( id = c ( 10 : 12 ), lh superiortemporal = c ( 3.32 , 4.1 , 3.5 ), lh precentral = c ( 2.3 , 2.5 , 2.1 ), lh rostralmiddlefrontal = c ( 3.3 , 3.2 , 3.1 ) ) freesurfer stats # Wrangle wide format data into long format freesurfer long <- freesurfer stats %>% gather (label, thickness, - id) freesurfer long ## id lh superiortemporal lh precentral lh rostralmiddlefrontal ## 1 10 3.32 2.3 3.3 ## 2 11 4.10 2.5 3.2 ## 3 12 3.50 2.1 3.1 ## id label thickness ## 1 10 lh superiortemporal 3.32 ## 2 11 lh superiortemporal 4.10 ## 3 12 lh superiortemporal 3.50 ## 4 10 lh precentral 2.30 ## 5 11 lh precentral 2.50 ## 6 12 lh precentral 2.10 ## 7 10 lh rostralmiddlefrontal 3.30 ## 8 11 lh rostralmiddlefrontal 3.20 ## 9 12 lh rostralmiddlefrontal 3.10 Data in long format can then be used directly with the ggseg() -function, as the ‘label’ column corresponds in name and content with the ‘label’ column in the atlas data of dkt . The data must include a column that has the same column name and at least some data matching the values in the corresponding column in the atlas data. In the next example we create some data with 4 rows, with an ‘area’ and ‘p’ column, representing the results of a hypothetical analysis. The ggseg() -function will recognise the matching column ‘area’, and merge the supplied data into the atlas using dplyr -joins. W e use the ‘p’ column as the column flooding the segment with colour . The appearance of the plot can then be modified similarly to any other ggplot2 graph using functions such as scales, labs, themes, etc., as seen in Figure 4 # Make some mock data someData = data.frame ( area = c ( "transverse temporal" , "insula" , "pre central" , "superior parietal" ), p = sample ( seq ( 0 ,. 5 ,. 001 ), 4 ), stringsAsFactors = FALSE ) # Figure 4 ggseg ( .data= someData, mapping= aes ( fill= p)) + labs ( title= "A nice plot title" , fill= "p-value" ) + scale fill gradient ( low= "firebrick" , high= "goldenrod" ) If the results are only in one hemisphere, but you still want to plot both of them, make sure your data.frame includes the column ‘hemi’ with either ‘right’ or ‘left’. In this case, data will be merged into the atlas both by ‘area’ and by ‘hemi’. For more information about adapting data and vie wing only one hemisphere or side, the package vignettes contains more elaborate information. 6/17 left right hemisphere 0.18 0.20 0.22 0.24 p−value A nice plot title Figure 4. Supplying data through the ‘.data’ option in ggseg() enables use of columns in the supplied data to aesthetical arguments (such as ‘fill’). The ggse g plot can be used with any other polygon compatible function from ggplot2 or ggplot2 e xtentions, for instance adding title, changing the legend name and the colour scheme with standard ggplot2 functions. 2.1.2 Creating subplots There is often the need to plot a statistic of interest in different groups (e.g. thickness or brain - cognition relationships in young or older adults). This may be obtained also with ggseg() , using ggplot2 ’ s facet wrap or facet grid , with three guiding rules: 1) as before, data needs to be in long format with a column indexing which group the ro w corresponds to (group data should appear in seperate rows, not in separate columns). 2) The data needs to be grouped using dplyr ’ s group by() function befor e providing the data to the ggseg() -function. The ggseg() -function will detect grouped data, and adapt it to facet ‘s requirements. 3) Apply facet wrap or facet grid to the plot having used the abov e two rules. An example of this can be seen in Figure 5, where a mock data set including summary statistics for two groups (’Y oung’ and ‘Old’) is used when faceting a ggse g -plot. All the concepts described abov e also work with the aseg atlas for subcortical structures, except for ‘hemisphere’ and ‘view’ ar guments that are superfluous in subcortical atlases (Figure 6). # Make some mock data someData = data.frame ( area = rep ( c ( "transverse temporal" , "insula" , "pre central" , "superior parietal" ), 2 ), p = sample ( seq ( 0 ,. 5 ,. 001 ), 8 ), AgeG = c ( rep ( "Young" , 4 ), rep ( "Old" , 4 )), stringsAsFactors = FALSE ) %>% group by (AgeG) # Figure 5 ggseg ( .data= someData, colour= "white" , position = "stacked" , mapping= aes ( fill= p)) + facet wrap ( ˜ AgeG, ncol= 2 ) + theme ( legend.position = "bottom" ) # Figure 6 ggseg ( atlas= "aseg" , mapping= aes ( fill= area)) + theme ( legend.justification= c ( 1 , 0 ), legend.position= "bottom" , legend.text = element text ( size = 5 )) + guides ( fill = guide legend ( ncol = 3 )) 2.2 Plotting 3D mesh data Representing brains as 2D polygons is a good solution for fast, efficient, and fle xible plotting, and can be easily combined with interacti ve apps such as Shiny (Chang et al. [2019]). Y et, brains are intrinsically 3-dimensional and it can be challenging to recognize the location of a region as a flattened image. This 7/17 Old Young lateral medial lateral medial left right side hemisphere 0.1 0.2 0.3 p Figure 5. T o visualize data corresponding to different groups, one needs to use group by or similar befor e plotting with ggseg() for facetting to work. left midline right hemisphere area 3rd ventricle 4th ventricle amygdala brain stem caudate CC anterior CC central CC mid anterior CC mid posterior CC posterior cerebellum cortex cerebellum white matter hippocampus lateral ventricle pallidum putamen thalamus proper ventral DC NA Figure 6. The aseg atlas showing subcortical structures has some distinct dif ferences from the dkt . For instance, there is no option to sho w only a single hemisphere. Furthermore, rather than showing lateral and medial surfaces, it sho ws an axial and sagittal slice. 8/17 problem is exacerbated in atlases that represent subcortical features as they are 3-dimensional, while cortical structures, such as grey matter structures, can be flattened to 2-dimensions. Hence, here we also provide the ggse g3d package to plot, vie w , and print 3D-atlases in R. g gse g3d is based on tri-surface mesh plots using plotly (Sie vert [2018]). The data structure is more complex than the ggplot2 polygons, and includes additional options for brain inflation, glass brains, camera locations, etc. As ggse g3d is based on plotly , the resulting brain atlases are interacti ve, which guides interpretation, and is useful for public dissemination. W e recommend users to familiarize themselv es with plotly (Sie vert [2018]) when using this function. Out-of-the-box, ggseg3d() plots the dkt 3d atlas in ‘LCBC’ surface, but there are two more surfaces a vailable for cortical atlases (Figure 7) The ‘LCBC’ surf ace consists on a semi-inflated white matter surface based on the fsaverage5 template subject. All [...] 3d atlases include a colour column that based on the color scheme used in the source neuroimaging software. Figure 7. The three surface options provided in ggse g3d atlases. From left to right: the ‘white’ surface is the white matter surface, ‘LCBC’ surface is the semi-inflated white matter surf ace (inflated over 10 iterations), and the ‘inflated’ surface is a inflated grey matter cortical surf ace as provided by the FreeSurfer software. The 3D-atlas data is stored in nested tibbles. Each cortical atlas has data sets for three dif ferent surfaces (see Figure 7) and the two hemispheres. Only one surface is available for subcortical atlases as inflation procedures are irrelev ant. The ‘ggseg 3d’ column includes all necessary information for ggseg3d() to create a 3D mesh-plot, and should not be modified by the user . The additional 3D-atlases in ggse gExtra ha ve the same data structure. It is important to note that the coordinates in the plot (X, Y , Z) are not any type of radiological coordinate system, b ut arbitrary Cartesian plot coordinates. # remotes::install github("LCBC-UiO/ggseg3d") library (ggseg3d) 2.2.1 External data suppl y Similarly as in the 2D-atlas, the user will use ggseg3d() to display through a colour scale some descriptiv e or inferential statistic. If the data is not already in the correct long format or uses similar naming as the atlas, the users should inspect the atlas data for a specific surface (and hemisphere, if desired), and then unnest(ggseg 3d) it to see how the atlas data is or ganised. # Select surface and hemisphere, and then unnest to inspect the atlas data dkt 3d %>% filter (surf == "inflated" & hemi == "right" ) %>% unnest (ggseg 3d) %>% head ( 5 ) Note the mesh column, which contains lists. Each list corresponds to a region and contains 6 vectors required to create the mesh of the tri-surface plot. It should also be noted that the ‘label’, ‘annot’ and ‘area’ columns could provide matching values for your own data. Similarly to the ggseg() -function, the ‘label’ column should match the region names used in the original neuroimaging software while ‘area’ and ‘annot’ provide alternativ e/secondary names. It is thus important to match your regional identifiers with those used in the atlas. T o colour the segments using a column from the data, a column name from the data needs to be supplied to the colour option, and providing it to the text option will add another line to the plotly hov er information. 9/17 # Figure 8 left ggseg3d ( .data = someData, atlas = dkt 3d, colour = "p" , text = "p" ) # Figure 8 middle ggseg3d ( .data = someData, atlas = dkt 3d, colour = "p" , text = "p" , palette = c ( "#ff0000" , "#00ff00" , "#0000ff" )) %>% pan camera ( "right lateral" ) %>% remove axes () # Figure 8 right ggseg3d ( .data = someData, atlas = dkt 3d, colour = "p" , text = "p" , na.colour = "black" ) %>% pan camera ( "right lateral" ) %>% remove axes () Figure 8. Left: Supplying data to ggseg3d() works similarly to ggseg() . Since ggseg3d() is based on plotly rather than ggplot2 , most aesthetic adaptations must be set in the main function. Here we set the parcellation colours with the ‘colour’ column and accompanying mouse-ho ver text with the ‘te xt’ column. Middle: The palette for ggse g3d needs to be specified directly in the main plot call. The ‘palette’ option takes vectors of colours either as HEX-codes or R-colour names. Right: The colour of the NA values can also be changed through the option ‘na.colour’. 2.2.2 Customizing colour s and the colour bar Y ou can provide custom colour palettes either in hex or R-names, as seen in Figure 8. Colours will be ev enly spaced when creating the colour-scale. A palette may also be supplied as a named numeric vector , where the vector names are the colours that users wish to use, and the numeric values are the breakpoints for each colour (e.g. c("red" = 0, "white" = 0.5, "blue" = 1) ). This way the users can control the minimum and maximum v alues of the colour scale, and also ho w the gradient is applied. If another colour than the default gray is w anted for the NA regions, supply ‘na.colour’, either as HEX colour or colour name. This option only takes a single colour . 2.2.3 Ad ding a glass brain Subcortical atlases include cortical surfaces and other landmark structures for visualization purposes only . One can control the opacity of the these NA structures, to improv e visualization. Glass brains can be added to provide a frame of reference for the subcortical structures with the function add glassbrain() (Figure 9) , which takes three extra ar guments: hemisphere, colour , and opacity . # Figure 9 ggseg3d ( atlas = aseg 3d, na.alpha= .5 ) %>% add glassbrain ( "left" ) %>% pan camera ( "left lateral" ) %>% remove axes () 10/17 Figure 9. For subcortical structure visualization, one can add a glass brain to the plot. This will help with locating the structures relativ e to the cortex, and make the plot easier to interpret. The glass brain is controlled by three options: opacity , hemisphere, and colour . ggsed3d() is based on plotly and thus additional plotly functionalities can be used to modify and improv e the 3D atlas representations. In addition to Carson Siev ert’ s book on plotly in R ([2018]), we recommend resources for modifying axes in 3D plots (Damiba [2019a]), the basic introduction to tri-surface plots (Damiba [2019b]), and this tutorial on tri-surface plots with plotly in R (Riddihiman [2016]). Finally , we recommend orca command line tool to sav e ggse g3d atlas snapshots. 2.3 Ad ditional atlases The ggse g and ggse g3d packages have tw o atlases each, which are 2D and 3D v ariations of the same main atlases: the dkt (Desikan et al. [2006]) and aseg (Fischl et al. [2002]) atlases. These are, howe ver , only two among man y meaningful ways of segmenting the brain into dif ferent regions. Thus, the ggse gExtra package is a repository containing additional with additional data sets for plotting with the ggse g and ggse g3d packages. There is an ever -increasing amount of new atlases being created, as research and methods in neuroimaging analysis progresses. The ggse gExtra -package is intended to be expanded as a community-ef fort, as ne w and informati ve atlases are published. A small collection of the atlases currently in the ggse gExtra package may be viewed in Figure 10 11/17 Figure 10. Four example data sets from the g gse gExtra library , plotted with the ggseg() and ggseg3d() functions. 3 DISCUSSION The main aim of the ggse g , ggse g3d , and ggse gExtra packages is to ease and streamline visualization of brain atlas data in R, by gathering a collection of atlases from sev eral scientific sources and providing customized plotting functions. In this tutorial, we introduced the packages to the readers by presenting some use examples and highlighting the main functions and options that are av ailable. As visualization tools, these packages add up to manifold functionalities such as ggBrain (Fisher [2019]) and ggneuro (Muschelli [2017]) in R, and software-specific image vie wers such as FSLe yes (McCarthy [2019]) and Freevie w (Dale et al. [1999]). In this regard, we do not aim to compete with software-specific visualizations or advocate for the superiority of the ggse g -packages as visualization tools. After all, flattened 2D polygons do not rely on a meaningful brain coordinate system and the units of information in 3D meshes are limited to the number of parcellations. On the contrary , we believe the ggse g niche among visualization tools resides in its simplicity and its ability to be combined with statistical analysis pipelines. The possibility to serve as an interacti ve tool for dissemination and reproducibility when combined with other technologies, such as Binder (Project Jupyter et al. [2018]) or Shiny (Chang et al. [2019]), is an added benefit. This is exemplified in the online supplementary information of V idal-Pi ˜ neiro et al. ([2019]). The three ggse g -packages contain three main features: 1) a collection of 2D-polygon and 3D mesh brain parcellation atlases. The atlas data include the necessary coordinates for plotting, and include other information that should be recognizable for users. 2) ggseg() and ggseg3d() functions for visualization. Both functions are flexible and well-adapted to their en vironment and can be combined with any additional argument from g gplot2 and plotly , respec- tiv ely . 12/17 - ggseg() is a wrapper function for geom polygon from ggplot2 and it can be built upon like any ggplot2 object. - ggseg3d() is a plotly wrapper function for tri-surface mesh plots which prints 3D atlases. 3) Complimentary features – e.g. color scales - and functions such as as ggseg atlas() and as ggseg3d atlas() to con vert data in the correct atlas format. These functions pro vide users with the possibility of adapting the plots to their wishes, and also makes it possible to create and contribute to the atlas repository in g gse gExtra . The foundations of the ggse g -packages trace back to the necessity of visualizing and exploring the lifespan trajectories of cortical thickness across diff erent brain regions (see supplementary information in V idal-Pi ˜ neiro et al. ([2019]) ) . That is, ggse g appears with the need to inspect and display brain information over time - i.e. including a spatial dimension and a time-varying factor - overcoming the constrains of printed journals and classical 2D plots (e.g. bar plots). The current state of science requires researches to share the results of studies in both high detail and in an intuitiv e manner, as it permits communication to wide audiences and f acilitates reproducibility . Hence, we belie ve this tool conforms to the essence of open science and in vite users to improv e the code, provide e xamples, or tutorials, and contribute to the atlas collection according to their o wn interest and needs via the public ggse g GitHub repository, ggseg3d GitHub repository, and ggse gExtra GitHub repository. Finally - while the ggse g -packages are circumscribed to brain parcellations - we believe that the structure and functions of the package can be easily applied to any scientific field that benefits from data being displayed across the spatial dimension. W e encourage readers to borro w the package functionalities and adapt it to their respectiv e fields and structures of interest, such as has already been done with the gganatogr am -package (Maag [2018]). 4 PLANNED P A CKA GE IMPRO VEMENTS In the ggse gExtra github wiki, we offer a pipeline to create and supply atlases for 2D plotting. At the moment, the creation of atlas for ggseg is con voluted and dif ficult and requires manual intervention. W e are in the process of designing a simple, straightforw ard pipeline to facilitate the creation of ne w ggseg (2D) atlases. The aim is to create a set of functions that will call specialized tools like FSL(W oolrich et al. [2009]), Freesurfer(Fischl et al. [1999], Dale et al. [1999], Fischl and Dale [2000]) and Imagemag- ick(Ooms [2019]) to detect the polygon vectors or the mesh se gments giv en an MRI image containing a parcellation specification, and organize these into valid ggse g and ggse g3d atlases. W e encourage users to contribute to the g gse gExtra brain atlas repository by including additional brain atlases. 5 CONCLUSION V isualization is a fundamental aspect of neuroimaging to explore and understand data, guide interpretation, and communicate with colleagues and the general audience. In this tutorial, we hav e introduced the ggse g - packages, tools for visualizing brain statistics through brain parcellation atlases in R. This visualization tool easily combines with interactiv e routines as well as with di verse statistical analysis pipelines. W e hope this tool and tutorial proves useful to neuroscientists and inspires others to apply the functions in a wide variety of fields and structures. 6 A UTHOR CONTRIBUTIONS Didac V idal-Pi ˜ neiro generated the idea for the tool, and the initial scripts for plot visualization. He has also been responsible for con verting images from neuroimaging data to ggseg-lik e data (e.g. polygons and mesh data). Athanasia M. Mowinckel adapted the initial scripts and made the functions into package format and has continued de veloping the functions with the aim of increasing user-friendliness. She is also responsible for conceiving and adding the mesh-plot functionality through plotly , and developing the pipeline for making that possible. A. M. Mowinck el wrote the first draft of the paper , and both have since critically edited it. 7 CONFLICTS OF INTEREST The authors declare that there were no conflicts of interest with respect to the authorship or the publication of this article. 13/17 8 A CKNO WLEDGEMENTS W e thank John Muschelli for package code comments and its adaptation to neuroconductor ([2018]), and Richard Beare for the first community created atlas, and base code comments. W e are indebted to A. M. W inkler scripts for con verting neuroimaging data (Freesurfer) into .ply files, a necessary step for neuroimaging to mesh-plots con version, and to Inge Amlien for the ‘LCBC’ surface files and helpful contributions to the ggseg3d pipeline. W e are indebted to those users that activ ely contributed to the package and all the individuals that work ed on the framework on which the package is sustained (e.g. R- project, neuroimaging software). Finally , we thank Anders M. Fjell (AMF) and Kristine B. W alhovd (KBW) for their encouragement and financial support. 9 FUNDING This work is funded by EU Horizon 2020 Grant ‘Healthy minds 0-100 years: Optimizing the use of European brain imaging cohorts (Lifebrain)’, with grant agreement 732592. The project has also receiv ed funding from the European Research Council’ s Starting (grant agreements 283634, to A.M.F . and 313440 to K.B.W .) and consolidator Grant Scheme (grant agreement 771355 to KBW and 725025 to AMF). The project has receiv ed funding through multiple grants from the Norwegian Research Council” 10 PRIOR VERSIONS Athanasia Monika Mo winckel also has se veral tutorials on her blog reg arding ggse g creation and function- ality (Mowinck el [2018a], Mowinck el [2018b]). REFERENCES Inge K. Amlien, Markus H. Sne ve, Didac V idal-Pi ˜ neiro, Kristine B. W alhovd, and Anders M. Fjell. Elaboration Benefits Source Memory Encoding Through Centrality Change. Scientific Reports , 9(1): 3704, March 2019. ISSN 2045-2322. doi: 10.1038/s41598- 019- 39999- 1. URL https://doi. org/10.1038/s41598- 019- 39999- 1 . W inston Chang, Joe Cheng, JJ Allaire, Y ihui Xie, and Jonathan McPherson. shiny: W eb Application F ramework for R , 2019. URL https://CRAN.R- project.org/package=shiny . R package version 1.3.1. Chi-Hua Chen, Mark Fiecas, E. D. Guti ´ errez, Matthew S. Panizzon, Lisa T . Eyler , Eero V uoksimaa, W esley K. Thompson, Christine Fennema-Notestine, Donald J. Hagler , T erry L. Jernigan, Michael C. Neale, Carol E. Franz, Michael J. L yons, Bruce Fischl, Ming T . Tsuang, Anders M. Dale, and W illiam S. Kremen. Genetic topography of brain morphology . Pr oceedings of the National Academy of Sciences , 110(42):17089–17094, 2013. ISSN 0027-8424. doi: 10.1073/pnas.1308091110. URL https://www.pnas.org/content/110/42/17089 . Anders Dale, Bruce Fischl, and Martin I. Sereno. Cortical surface-based analysis: I. segmentation and surface reconstruction. Neur oImage , 9(2):179 – 194, 1999. Joseph Damiba. plotly axes. https://plot.ly/r/axes/ #modifying- axes- for- 3D- plots , 2019a. Accessed: 2019-04-01. Joseph Damiba. plotly tri-surf. https://plot.ly/r/trisurf/ , 2019b. Accessed: 2019-04-01. Rahul S. Desikan, Florent S ´ egonne, Bruce Fischl, Brian T . Quinn, Bradford C. Dickerson, Deborah Blacker , Randy L. Buckner , Anders M. Dale, R. Paul Maguire, Bradle y T . Hyman, Marilyn S. Albert, and Ronald J. Killian y . An automated labeling system for subdi viding the human cerebral corte x on mri scans into gyral based regions of interest. Neur oImage , 31(3):968 – 980, 2006. ISSN 1053-8119. doi: https://doi.org/10.1016/j.neuroimage.2006.01.021. URL http://www.sciencedirect.com/ science/article/pii/S1053811906000437 . Christophe Destrieux, Bruce Fischl, Anders Dale, and Eric Halgren. Automatic parcellation of hu- man cortical gyri and sulci using standard anatomical nomenclature. Neur oImage , 53(1):1 – 15, 2010. ISSN 1053-8119. doi: https://doi.org/10.1016/j.neuroimage.2010.06.010. URL http: //www.sciencedirect.com/science/article/pii/S1053811910008542 . 14/17 Simon B. Eickhoff, B. T . Thomas Y eo, and Sarah Genon. Imaging-based parcellations of the human brain. Nature Reviews Neur oscience , 19(11):672–686, November 2018. ISSN 1471-0048. doi: 10.1038/s41583- 018- 0071- 7. URL https://doi.org/10.1038/s41583- 018- 0071- 7 . Bruce Fischl and Anders M. Dale. Measuring the thickness of the human cerebral cortex from magnetic resonance images. Pr oceedings of the National Academy of Sciences of the United States of America , 97(20):11050–11055, 2000. Bruce Fischl, Martin I. Sereno, and Anders Dale. Cortical surface-based analysis: Ii: Inflation, flattening, and a surface-based coordinate system. Neur oImage , 9(2):195 – 207, 1999. Bruce Fischl, David H. Salat, Evelina Busa, Marilyn Albert, Megan Dieterich, Christian Haselgrov e, Andre v an der K ouwe, Ron Killiany , David Kennedy , Shuna Kla veness, Albert Montillo, Nikos Makris, Bruce Rosen, and Anders M. Dale. Whole brain segmentation: Automated labeling of neuroanatomical structures in the human brain. Neur on , 33(3):341 – 355, 2002. ISSN 0896-6273. doi: https://doi.org/10.1016/S0896- 6273(02)00569- X. URL http://www.sciencedirect.com/ science/article/pii/S089662730200569X . Aaron Fisher . ggBr ain: ggplot Brain Imag es , 2019. R package version 0.1. Matthe w F . Glasser , T imothy S. Coalson, Emma C. Robinson, Carl D. Hacker , John Harwell, Essa Y acoub, Kamil Ugurbil, Jesper Andersson, Christian F . Beckmann, Mark Jenkinson, Stephen M. Smith, and David C. V an Essen. A multi-modal parcellation of human cerebral cortex. Natur e , 536:171 EP –, 07 2016. URL https://doi.org/10.1038/nature18933 . Kurt Hornik. The comprehensive r archive network. WIREs Computational Statistics , 4(4):394–398, 2012. doi: 10.1002/wics.1212. URL https://onlinelibrary.wiley.com/doi/abs/10. 1002/wics.1212 . Ke gang Hua, Jiangyang Zhang, Setsu W akana, Hangyi Jiang, Xin Li, Daniel S. Reich, Peter A. Calabresi, James J. Pekar , Peter C.M. van Zijl, and Susumu Mori. T ract probability maps in stereotaxic spaces: Analyses of white matter anatomy and tract-specific quantification. Neur oImage , 39(1):336 – 347, 2008. ISSN 1053-8119. doi: https://doi.org/10.1016/j.neuroimage.2007.07.053. URL http://www. sciencedirect.com/science/article/pii/S105381190700688X . Jesper Maag. gganatogram: An r package for modular visualisation of anatograms and tissues based on ggplot2. f1000r esear ch , 2018. URL https://f1000research.com/articles/7- 1576/ v1 . V ersion 1: A waiting peer revie w . Nikos Makris, Jill M. Goldstein, David Kennedy , Steven M. Hodge, V erne S. Caviness, Stephen V . Faraone, Ming T . Tsuang, and Larry J. Seidman. Decreased volume of left and total anterior insular lobule in schizophrenia. Schizophr enia Resear ch , 83(2):155 – 171, 2006. ISSN 0920-9964. doi: https: //doi.org/10.1016/j.schres.2005.11.020. URL http://www.sciencedirect.com/science/ article/pii/S0920996405004998 . Paul McCarthy . Fsleyes, April 2019. URL https://doi.org/10.5281/zenodo.2630502 . Susumu Mori, S. W akana, Peter C M v an Zijl, and L.M. Nagae-Poetscher . MRI Atlas of Human White Matter . Elsevier Science, 2005. ISBN 9780080456164. URL https://www.elsevier.com/ books/mri- atlas- of- human- white- matter/mori/978- 0- 444- 51741- 8 . A.M. Mowinck el. Get the brain animated! - Dr .Mowinckels blog. https://drmowinckels.io/ blog/get- the- brain- animated/ , 2018a. Accessed: 2019-04-01. A.M. Mowinck el. Introducing the ggseg r -package for brain segmen- tations - Dr .Mowinck els blog. https://drmowinckels.io/blog/ introducing- the- ggseg- r- package- for- brain- segmentations/ , 2018b. Ac- cessed: 2019-04-01. A.M. Mowinck el and D. V idal-Pi ˜ neiro. ggseg package website. https://lcbc- uio.github.io/ ggseg/ , 2019a. Accessed: 2019-11-26. 15/17 A.M. Mowinckel and D. V idal-Pi ˜ neiro. ggseg3d package website. https://lcbc- uio.github. io/ggseg3d/ , 2019b. Accessed: 2019-11-26. J. Muschelli, A. Gherman, J. P . Fortin, B. A vants, B. Whitcher , J. D. Clayden, B. S. Caff o, and C. M. Crainiceanu. Neuroconductor: an R platform for medical imaging analysis. Biostatistics , Jan 2018. John Muschelli. ggneur o: Plotting Functions for Neur oimaging Data in ’ggplot2’ , 2017. R package version 0.5.0. Jeroen Ooms. magick: Advanced Graphics and Image-Pr ocessing in R , 2019. URL https://CRAN. R- project.org/package=magick . R package version 2.2. Diego A. Pizzagalli, A vram J. Holmes, Daniel G. Dillon, Elena L. Goetz, Jeffrey L. Birk, Ryan Bogdan, Darin D. Dougherty , Dan V . Iosifescu, Scott L. Rauch, and Maurizio Fav a. Reduced caudate and nucleus accumbens response to rew ards in unmedicated individuals with major depressive disorder . American J ournal of Psychiatry , 166(6):702–710, 2009. doi: 10.1176/appi.ajp.2008.08081201. URL https://doi.org/10.1176/appi.ajp.2008.08081201 . PMID: 19411368. Project Jupyter , Matthias Bussonnier , Jessica Forde, Jeremy Freeman, Brian Granger , Tim Head, Chris Holdgraf, Kyle K elley , Gladys Nalvarte, Andre w Osherof f, M Pacer , Yuvi Panda, Fernando Perez, Ben- jamin Ragan K elley , and Carol W illing. Binder 2.0 - Reproducible, interactiv e, sharable en vironments for science at scale. In Fatih Akici, David Lippa, Dillon Niederhut, and M Pacer , editors, Pr oceedings of the 17th Python in Science Confer ence , pages 113 – 120, 2018. doi: 10.25080/Majora- 4af1f417- 011. R Core T eam. R: A Language and En vir onment for Statistical Computing . R Foundation for Statistical Computing, V ienna, Austria, 2019. URL https://www.R- project.org/ . Riddihiman. plotly trisurf2. https://moderndata.plot.ly/ trisurf- plots- in- r- using- plotly/ , 2016. Accessed: 2019-04-01. Alexander Schaefer , Ru K ong, Ev an M Gordon, T imothy O Laumann, Xi-Nian Zuo, A vram J Holmes, Simon B Eickhoff, and B T Thomas Y eo. Local-Global Parcellation of the Human Cerebral Cortex from Intrinsic Functional Connectivity MRI. Cerebr al Cortex , 28(9):3095–3114, 07 2017. ISSN 1047-3211. doi: 10.1093/cercor/bhx179. URL https://doi.org/10.1093/cercor/bhx179 . Carson Siev ert. plotly for R , 2018. URL https://plotly- r.com . D. V idal-Pi ˜ neiro, N. Parker , J. Shin, L. French, AP . Jacko wski, AM. Mowinck el, Y . Patel, Z. Pausov a, G. Salum, Ø. Sørensen, KB W alhovd, T . Paus, and AM Fjell. Cellular correlates of cortical thinning throughout the lifespan. bioRxiv , page 585786, January 2019. doi: 10.1101/585786. URL http: //biorxiv.org/content/early/2019/03/23/585786.abstract . Kristine B. W alhovd, Anders M. Fjell, Ivar Reinv ang, Arvid Lundervold, Anders M. Dale, Dag E. Eilertsen, Brian T . Quinn, David Salat, Nikos Makris, and Bruce Fischl. Effects of age on volumes of cortex, white matter and subcortical structures. Neur obiology of Aging , 26(9):1261 – 1270, 2005. ISSN 0197-4580. doi: https://doi.org/10.1016/j.neurobiolaging.2005.05.020. URL http://www. sciencedirect.com/science/article/pii/S0197458005001673 . Hadley W ickham. ggplot2: Ele gant Graphics for Data Analysis . Springer-V erlag New Y ork, 2016. ISBN 978-3-319-24277-4. URL https://ggplot2.tidyverse.org . Mark W . W oolrich, Saad Jbabdi, Brian Patenaude, Michael Chappell, Salima Makni, T imothy Behrens, Christian Beckmann, Mark Jenkinson, and Stephen M. Smith. Bayesian analysis of neuroimaging data in FSL. Neur oImage , 45(1):S173–S186, March 2009. doi: 10.1016/j.neuroimage.2008.10.055. URL https://doi.org/10.1016/j.neuroimage.2008.10.055 . Anastasia Y endiki, Patricia Panneck, Priti Sriniv asan, Allison Stev ens, Lilla Z ¨ ollei, Jean Augustinack, Ruopeng W ang, David Salat, Stefan Ehrlich, T im Behrens, Saad Jbabdi, Randy Gollub, and Bruce Fischl. Automated probabilistic reconstruction of white-matter pathways in health and disease us- ing an atlas of the underlying anatomy . F rontier s in Neur oinformatics , 5:23, 2011. ISSN 1662- 5196. doi: 10.3389/fninf.2011.00023. URL https://www.frontiersin.org/article/10. 3389/fninf.2011.00023 . 16/17 Thomas Y eo, B. T ., Fenna M. Krienen, Jorge Sepulcre, Mert R. Sabuncu, Danial Lashkari, Marisa Hollinshead, Joshua L. Roffman, Jordan W . Smoller , Lilla Z ¨ ollei, Jonathan R. Polimeni, Bruce Fischl, Hesheng Liu, and Randy L. Buckner . The organization of the human cerebral cortex estimated by intrinsic functional connectivity . J ournal of Neur ophysiology , 106(3):1125–1165, 2011. doi: 10.1152/ jn.00338.2011. URL https://doi.org/10.1152/jn.00338.2011 . PMID: 21653723. 17/17
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment