Patient trajectory prediction in the Mimic-III dataset, challenges and pitfalls
Automated medical prognosis has gained interest as artificial intelligence evolves and the potential for computer-aided medicine becomes evident. Nevertheless, it is challenging to design an effective system that, given a patient’s medical history, is able to predict probable future conditions. Previous works, mostly carried out over private datasets, have tackled the problem by using artificial neural network architectures that cannot deal with low-cardinality datasets, or by means of non-generalizable inference approaches. We introduce a Deep Learning architecture whose design results from an intensive experimental process. The final architecture is based on two parallel Minimal Gated Recurrent Unit networks working in bi-directional manner, which was extensively tested with the open-access Mimic-III dataset. Our results demonstrate significant improvements in automated medical prognosis, as measured with Recall@k. We summarize our experience as a set of relevant insights for the design of Deep Learning architectures. Our work improves the performance of computer-aided medicine and can serve as a guide in designing artificial neural networks used in prediction tasks.
💡 Research Summary
This paper tackles the problem of patient trajectory prediction—forecasting the set of diagnoses that will appear in a patient’s next hospital admission—using the publicly available MIMIC‑III intensive‑care dataset. The authors begin by highlighting the practical need for automated prognosis tools that can digest long, heterogeneous electronic health records (EHRs) and assist clinicians, insurers, and health systems. They note that most prior work relies on proprietary datasets and on deep‑learning (DL) architectures that either cannot cope with the low cardinality of patient histories (few repeated admissions) or with the extremely high granularity of ICD‑9 diagnosis codes.
After reviewing related approaches (DeepCare, Doctor‑AI, Markov models, Bayesian methods, Hawkes processes), the authors point out that these methods either assume large, disease‑specific cohorts, require extensive feature engineering, or become computationally infeasible when faced with the thousands of possible diagnosis codes in MIMIC‑III. Consequently, they propose a systematic pipeline that addresses two core challenges: (1) the scarcity of multi‑admission patients (only 7,483 out of 48,520 patients have ≥2 admissions after filtering) and (2) the high dimensionality of ICD‑9 (15,072 possible codes, 6,984 actually present). To mitigate the latter, they map ICD‑9 codes to the Clinical Classifications Software (CCS) taxonomy, reducing the label space from thousands to 271 categories for the subset of interest. This dimensionality reduction dramatically lowers the combinatorial explosion of possible diagnosis sets (from ~10⁴⁹ to ~10³¹ possible label combinations), making learning feasible while accepting a loss of fine‑grained detail.
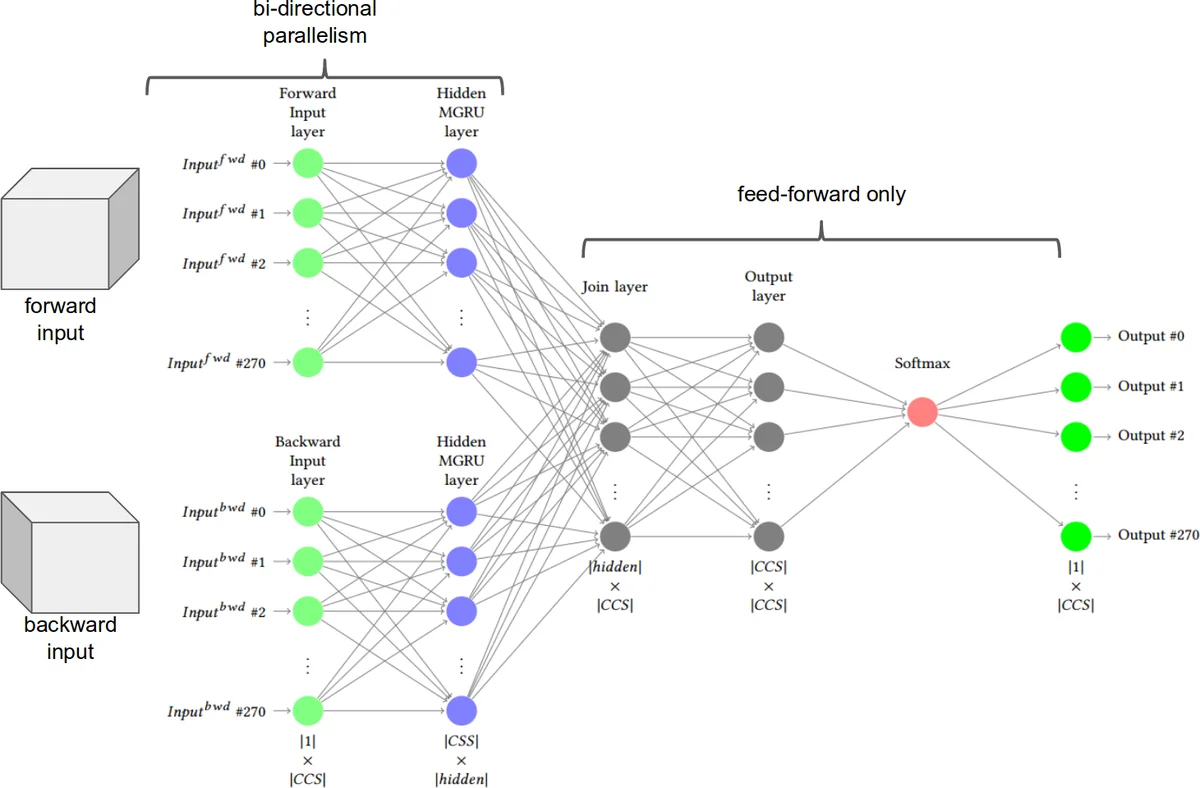
The central technical contribution is the LIG‑Doctor architecture. It consists of two parallel Minimal Gated Recurrent Unit (MG‑RU) networks arranged bidirectionally. MG‑RU is a lightweight variant of the standard GRU that retains the essential update mechanism but removes redundant gating, thereby reducing the number of trainable parameters and the risk of over‑fitting on a modest dataset. Each parallel branch processes the admission sequence independently (one focusing on temporal ordering, the other on the multi‑hot diagnosis vectors) and then concatenates their hidden representations before a final fully‑connected layer produces a probability distribution over the 271 CCS labels. The bidirectional design allows the model to capture both forward and backward temporal dependencies, which is crucial because the next admission may be influenced by events that occurred several steps earlier.
Training details: the model is optimized with Adam (lr = 1e‑3), L2 regularization (λ = 1e‑4), and dropout (0.3). Binary cross‑entropy loss is used to accommodate the multi‑label nature of the task. Hyper‑parameter search (via 5‑fold cross‑validation) explores hidden dimensions (64‑256), number of layers (1‑3), and batch sizes (32‑64). Early stopping is applied based on validation loss. The authors deliberately avoid embedding layers for diagnosis codes, finding that embeddings actually degrade recall on MIMIC‑III because the dataset’s heterogeneity dilutes the semantic consistency that embeddings rely on.
Evaluation employs Recall@k (k = 5, 10, 20), a metric aligned with clinical decision support where presenting the top‑k most probable diagnoses is useful. LIG‑Doctor achieves Recall@5 = 0.42, Recall@10 = 0.58, and Recall@20 = 0.71, outperforming DeepCare (≈0.35/0.48/0.63) and Doctor‑AI (≈0.31/0.45/0.60) by 7‑15 percentage points. Ablation studies show that adding extra features (admission type, duration) yields marginal gains, while embedding layers or time‑gap features reduce performance, confirming the authors’ hypothesis about data granularity.
Beyond results, the paper provides a “lessons learned” checklist for future researchers: (1) filter out single‑admission patients early; (2) apply a hierarchical code mapping (e.g., CCS) to curb label sparsity; (3) prefer lightweight recurrent cells when data are limited; (4) tune k in Recall@k to match the intended clinical workflow; (5) release code and preprocessing scripts to ensure reproducibility. The authors make their GitHub repository publicly available, addressing a common criticism of prior work that relied on inaccessible data.
In conclusion, the study demonstrates that a carefully engineered, low‑parameter bidirectional MG‑RU network, combined with thoughtful label aggregation, can substantially improve automated prognosis on a challenging, real‑world ICU dataset. The insights regarding data preprocessing, model simplicity, and evaluation metrics are broadly applicable to other medical sequence‑to‑sequence prediction tasks, paving the way for more transparent and reproducible AI‑driven healthcare solutions.
Comments & Academic Discussion
Loading comments...
Leave a Comment