Fragment Graphical Variational AutoEncoding for Screening Molecules with Small Data
In the majority of molecular optimization tasks, predictive machine learning (ML) models are limited due to the unavailability and cost of generating big experimental datasets on the specific task. To circumvent this limitation, ML models are trained…
Authors: John Armitage, Leszek J. Spalek, Malgorzata Nguyen
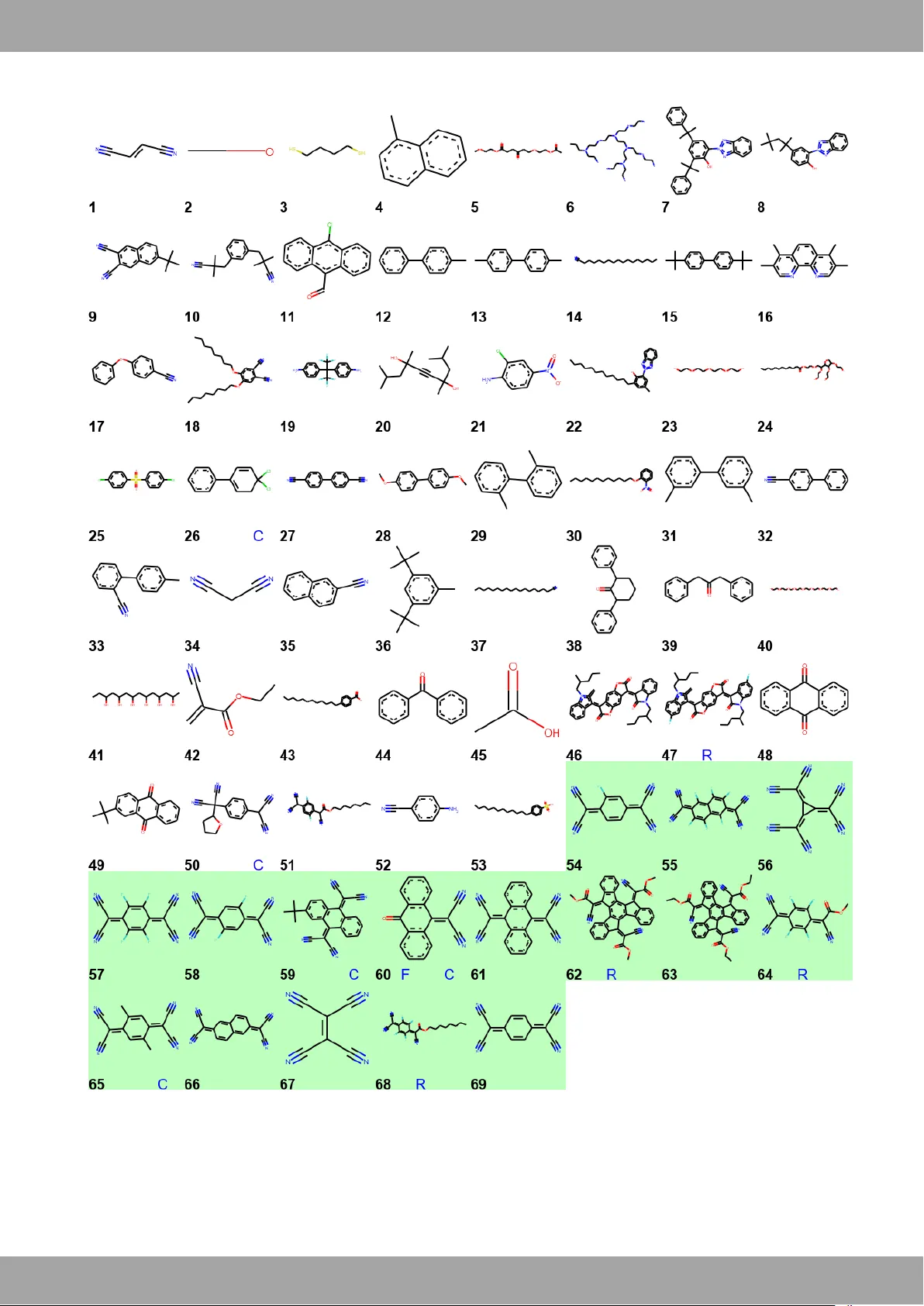
Fragment Grap hical Variatio nal Auto E ncodi ng for Scree ning Molecul es with Small Dat a a. De par t m ent of P h ysic s , Ca v en di sh La b orat o ry C am bri d ge Un iv er si ty , J J T ho m son Av enu e , Cam b ri dg e , CB 3 0H E. E- m ail: H . S. h s 22 0 @ca m .a c .uk b. M at eri al Re sea r ch In stit ut e , Que en M a ry , U ni ver sit y of L on don , Mil e En d Ro a d , Lon d on E 1 4 N S . E-m ail : C.B .N c .b .nie ls en @ qm ul . ac. uk c. Sol ar & P ho to vol tai c s En gi ne eri ng Re se ar ch C ent re , Divi si o n o f P h y sical S cie n ce an d En gin e eri n g, Ki ng A b dull a h Uni ve r sit y o f S ci en ce an d T ech nol o g y ( KA US T) , Thu w al 23 9 55 -6 90 0 , Ki n gd om o f Sa u di A ra bi a . El ect ro ni c Su ppl e me nt a ry I nfo rm ati o n (E SI) * a vail a bl e: [ h tt ps: / /git hu b. co m/ O E- FE T/ Fra G VA E ]. Fragment Graphical Variational AutoEnc oding for Scr eenin g Molecules with Smal l Data John Ar mitage a , Leszek J. Spalek a , Malgorzata Nguyen a , Mark Niko lka a , Ian E. Jacobs a , Lore na Marañón b , Iyad Nasral lah a , Gui llaume Sch weiche r a , Ivan Dimo v a , D imit rios Si matos a , Iai n McCulloch c , Chr istian B. N iel sen b , Gareth Conduit a and Henn ing Sirrin ghaus a In the majority of molecular optimizatio n tasks, predictive machin e learning ( ML) m odels are limited due to the unavailabili ty and c ost of genera ting big experimen tal datasets on the specific task. To circu mvent t his limita tion, ML models a re tr ained on big the oretical d atas ets or experi mental indicat ors of molecular suita bility that ar e eit her p ubli cly avail able o r i nexpensiv e to acquir e. These a pproaches produce a set of can didate molecules which have to b e ranked using limited e xperimental data or expert kn owledge. Und er th e assu mption th at stru cture is r elated t o functi onality, here w e use a molecular fragm ent-bas ed gra phical autoenc oder t o ge nerate unique struct ural finger prints t o e fficiently sear ch through the ca ndidate s et. We d emonstrate t hat fra gment -bas ed gr aphic al autoencoding re duces the err or in predicting physical c haracteristi cs s uch as the solubili ty and par tition c oefficient in the smal l data regim e com pared t o ot her extended circula r finger prints and s tring based a pproach es. W e f urther demonstra te that t h is ap proach is capa ble of providing insight int o real w orld molecula r optimizati on proble ms, s uch a s s earching for stabil ization a dditives i n organic semiconduct ors by accurately predicting 92% of test molecules given 69 tr aining examples . This task is a m o del example of black box molecular optimizati on as there is mini mal th eoretical and experimental knowledg e t o acc urately predict the suitability of t he additives. Introduction A sign ific ant at tribute o f orga nic mol ecules is their almo st infinite chemica l struct ural variat ions w hich exhibit a r ange of tunable propert ies 1,2 . The cha lleng e of molecu lar opt imizat ion is to eff iciently find the appropr iate molecu lar st ructure for a particul ar t ask. In pract ice, whi le cert ain att ributes can b e simulated, t his is not tru e for all t asks d ue to incompl ete theory or int ractab le comput ation . In t hese c ases, m olecu lar optimizati on is driven by e xpens ive and time -cons umin g empirical me asureme nts an d not by an alytic al pred icti ons. A promisi ng rout e to impro ve the searc hing e ffic iency of molecular struc tures is t o aug ment co mputation al an d experiment al discove ry of novel m aterials usi ng mac hine learning t echn iques. Machine learning (ML) p rovides a ro ute t o effici ently obtain a mapping from the fe atures of experime nt s to the ir outcome s through stat istical t echnique s. ML algorit hms have alre ady been us ed t o predi ct valid organic molecules for bot h pharm aceutic al and organ ic el ectronics app licati ons. The se approache s focus on pre d icting valid mo lecules bas ed on bi g data from e ither know n dat abases or relevant theoret ical calculat ions. For exam ple, ML techn iques are app lied to big theoret ical and expe riment al dat abases to pre dict met ric s such as drug likeli ness a nd partition c oeffic ient, wh ich ar e strong theoret ical indicat ors t o pre -scree n the drugs 3 – 5 . I n organic elect ronics , mach ine le arni ng has been use d to efficient ly pr oduc e t heoretica l ind icators fo r dat asets w hich are too l arge to be e xha ustively sc reened w ith quant um simulati ons 6,7 . In b oth of these e xam ples, the ML is not learning fr om l arge experi mental dat asets but from large theoret ical d atabases gene rated by accur ate t heoretic al represent ations . H owever, ver y oft en su ch the oret ical m odel s do not e xist and man y of the ex isting ones are incom plete, as they generate indic ators of vali d molecul ar s tructur es but are not able to efficient ly model the compl ete material syste m. Assuming the t heoretic al mo dels are v alid, t hey can pre -scree n many mole cular struct ures; howeve r, t he rem ain ing set of molecules , t he can did ate set (Figur e 1), w ill still n eed t o b e screened , base d on e mpir ical data or e xpert kn owled ge. Fragment Grap hical Var iational AutoEnc oding for Scr eening Molecules with Sm all Data arXiv 2 | J . A r m i t a g e , 2 0 1 9 A clear t heore tical ga p in dr ug discover y is w hether a d rug ha s any side e ffects, as acc urat e simul ations of t he co mplet e biologica l syst em are intract able. S imilar ly, i n org anic electronics, pre dicting the morpho log y of org anic semicon ductors pure ly on che mica l struct ure is high ly pro ne t o error. U nfortun ately, unlike other areas of applied mac hin e le arni ng w ithout the oretica l m odels, suc h as imag e recogniti on, natural language processing and financ e , experiment al data for mo lecular disco very is limit ed and extremel y c ost ly to acquire. As most molecul ar optimizat ion problems do not have a valid and accu rate theo retical mod el for all r elevant asp ects, this sm all d ata reg ime is t he b ottlenec k of mos t m olecular disc overy a ppl ications and t his is w hy molecular opt imizat ion is an extremely dif ficult problem. T o support t his ef fort, our object ive is t o pro vid e a n int uitive struc tural lat ent space base d on m olec ular fr agments , w hic h reduces the a mount of inf ormat ion requ ired to f ind appropriat e molecula r struc tures in t he candid ate s et. Fragment s are subgraphs of molec ular struc tures commo nl y used as b asis fu nct ions in or gani c chem istry. In machine le arni ng, e fficie nt encodi ngs of data c an b e achieved t hr ough a proces s know n as autoe ncod ing, a n unsuperv ised le arni ng alg orithm . An autoen coder c onsists of a neural n et work t hat le arns how t o copy the inp ut to its outp u t, however, the nu mber of neurons repre senting t he inp ut at one of the l ayers in t he p ipeline is reduce d, resu lting i n a force d dimensi onality redu ction , Figure 1 8 . This technique is used in image and n atural lang uage proces sing to gener ate compresse d repres ent ations o f im ages an d t exts 9 . Here w e train a gr aph ical a utoenco der t o ge nerate an ef ficient lat ent space re prese ntation of our ca ndidat e molec ules in relat ion to other molec ules in t he set . T his appro ach d iffers f rom traditiona l che mica l techni ques, w hich att empt t o make a fingerprint sys tem for all pos si ble mo lecular s truct ures inst ead of a spec ific set. Assumi ng a m olec ular s tructur e is n ot randomly rel ated to functio nality, the design of a smoot h struc turally s orted s pace shou ld als o pe rmit a sm ooth map ping of de scriptors ont o pr opert ies. This reduc es the Nyq uis t criterion resulting in less informat ion requir ed to accurat ely model pr opert ies. Hence , a sorted spa ce w ould incre ase t he search e fficiency of any bl ack box optim ization tec hnique 10 . In s umm ary, t he pr ima ry h ypothe sis in this work is that graphicall y aut oenc oding can did ate mo lecul ar gr aphs produc es e fficient fingerp rint s of candid ate molec ules in th e small d ata regime . As this st ructure -focused app roac h will n ot be ab le to captur e all know n q ualitat ive the oret ical or experiment al knowledg e, this app roac h shoul d be us ed as a n unbiase d qua ntit ative struct ure activity relat ionsh ip met hod to aid a c ollabor ative dec ision -ma king pr ocess. T his a pproac h would help to augm ent the screen ing of molecu lar structur es by pro viding a n unb iased plausibi lity of s ubsequ ent mol ecule s given the s mall amount o f establ ished data. To validate our primary h ypothesis t hat graphical aut oenc oded represent ations are app ropriat e for molecular fi nger prints i n the small d ata reg ime, w e com pare t he pred ictions of our graph -based met h od t o stand ard che mica l and s tring -base d molecular fingerpri nt s in both t heor etical and experi ment al dataset s. Large t heoretic al datasets are use d to gener ate robust s tatis tics of sim ilar sma ll dat asets un der t he assum ptio n Figure 1 : Pipeline of clustering molecular candidat es bas ed on structures using a molecular autoencoder. The graphical en coder reduces the dimensionali ty of a graph representati on of a molecule to a specific point in a continuous latent space. The decod er samples the same point in the latent space to rebuild the same graph. By t raining the encoder and decoder to learn how t o decompose and r econstruct molec ules in a reduced dimen sional space, the algorithm l earns how to efficiently represent molec ular graphs relative to oth er molecular graphs in the candidate s et. As molecules wit h simi lar fragments are located clos er in the latent space and assumi ng structur e is related to functionalit y , minimal data i s req uired to label regions of the m olecular space (green and blu e) in order to predict unknown regions (tan). arXiv Fra gment Gra phical Vari ational Aut oEn c oding f or Screening M olecules wi th S mall Data J . A r m i t a g e , 2 0 1 9 | 3 that t heoretical dat abas es are an accur ate representatio n of practica l e xperi ments. To demons trate gr aph ical aut oencodin g in smal l exp eriment al datas ets, we used this techn ique t o search for molecul ar add itives in organic se miconduct ors . I n organic electro nics the major lim iting fact or for de vice applicat ion is t hat s olution pr ocessed devic es ha ve p oor stability. Howe ver, rece nt work has dem onstrat ed t hat the format ion of t raps r espons ible for device degr adation can be stabilized usi ng bot h liquid and solid additiv es. As the specific mecha nism is un clear , he re w e use ML t o augm ent t he se arc h for new molecul ar addit ives. Fragmen t Graph ical Au toenco ding An es ta blished appr oach to autoe ncode m olec ular graphs is t o avoid gra phs altog ether a nd co nvert t he gr aph int o a one dimensi onal r epresent atio n, su ch as a s tring base d on S MILE S (simplifie d m olecular -inp ut line -entry system) or t rees 4,5 . A known prob lem with t he st ring ap proach is t hat s ma ll variations in the m olecular st ructure c an result in larg e modificat ions of the s tring 5 . Tree st ructures are m ore ro bus t but t he enco ding is st ill depende nt on an arb itrary trace an d start ing node 5 . T his e ncod ing sc heme result s in mult iple arbitrar y en cod ings for a given m olecu lar st ructure, w hich could result in a more complic ated m olecular sp ace, undesir able for sm all dat asets, w hich we avo id by encodin g graphicall y. Interes t in the e ncod ing of molec ular gra phs h as e xplode d, resulting in a class of ne ural net work archit ectures know n as Message P assing Neu ral Netw orks (MPNN s), ca pable of generating uniq ue enc odi ngs of molec ular s t ructure b y exploiting Ba nach's f ixed poi nt theorem 11 . T he challeng e of applying t hese t echn iques to sma ll data s et appl ications is t o ensure t hat the model do es not overf it, as t he unique encoding of larger st ructures requires de ep MPNN. F or example, t o enc ode a fragme nt with a m aximum deg ree of 3, one r equi res 3 gr aphi c m essage pas sing iterat ions and additio n al la yers to rel ate the out put of t he MPNN t o t he depend ent variab le. As eac h l ayer has hundr eds of param et ers this proces s is p rone to over fitting 7 . To t rain a dee p MP NN on a s mall d ataset , here we e xpl oit a major challeng e of gr aphical autoenco ders, dire ct autoenco ding is intract a ble for re asona bly sized gr aphs 12 . T o make t he p roblem t ractab le, a c omm on appro ach is t o perform a sequenc e of discrete decis ions t o rec onstruct an undirected gr aph t race . In this case t he ne xt graph in t he sequence is dependent only on the current stat e of the g rap h and not t he hist ory of the gra ph trace. This results i n o rders of magnitud es more t raining e xamples f or t he MPNN enc oder f or each m olecular grap h in the train ing set. What is uniq ue in t his work is that the gr aph is r econst ructe d fragme nt by fragm ent ; he nce, t his proce dure is ca lled Fragment Gaphic al Variat ional A utoEncodi ng (Fra GVAE). T he smallest fragment is a n e xtende d c onne ctivity fing erpri nt (ECFP) wit h a radius of 1, whic h is a node at om connecte d to neighb oring no des 13 . As thes e frag ment s are s mall, t hey can b e directly de coded from fragm ent lat ent s pace ( ), unlike graphs larger t han 6 nodes 14 . A pr operty of these fragment s i s that each of t hem must be inc luded onc e an d o nly onc e whe n rebuilding t he f inal st ructure t o all ow sampl ing w itho ut replace ment. To reco nstruct the large graph, here w e randomly select a nucl eati ng fragment with a nu mber of dangling bonds (cy an) t hat ca n accept fragm ent s to e xpand t h e struc ture. In an ite rative a ppro ach, t he c orrect ne ighborin g fragme nt from t he frag ment bag, based on larger radiu s fragme nts ind icative of the c onne ctivit y of s mall e r fragm ent s in a separ ate latent space , is connect ed t o t he e merging molecule st ructure. The full mol ecul ar gr aph is enc oded in th e combine d latent sp ace , Figure 1. Tr aining a n etwork t o Figure 2 : FraGVAE autoencod er overview: The graph is decomposed into a bag of fragments, enc oded to a la tent space ( ) and then decoded to re produce the fragment bag. Secondly the connectivity of fragments is encoded to a latent space ( ) and, using the b ag of fragments and , the molecular struc tur e is reconstructed . Steps 1-4 demo nstrat e the first i terations of adding fragments/rings to a random starting frag men t t o reco nst ruct the molecular structu re. Fragment Grap hical Var iational AutoEnc oding for Scr eening Molecules with Sm all Data arXiv 4 | J . A r m i t a g e , 2 0 1 9 autoenco de a molecula r graph with N un ique fr agment s w hich can c onnect t o every other frag ment resu lts in N ! trai ning examples to autoen code a sing le m olecular stru cture. T hi s property he lps the aut oencode r t o be ro bust t o overfit tin g even wit h a sm all number of trainin g exa mples. In cont rast a similar s tring b ased a pproach w ould have 1 traini ng e xample. The deco mposit ion o f ibupro fen to c ircular fragment s cent ere d on at oms ( green no des) w it h a r adius o f 1 wit h t he ir neighb oring atoms (cy an nodes/ bonds) and rec onstruct ion process can b e seen in F igur e 2. This iterati ve proc ess is terminat ed when all d angl ing bon ds are cons idered, re sultin g in t he reprod uct ion o f the origi nal m olecular st ructure. All dangling bo nds are re mov ed by e ither con necting t o a terminat ing frag ment or connect ing to anot her d angling bon d on t he emerg ing m olec ule form ing a ring. To check the validit y of each propos ed subgr aph (graph index ) in iterat ion , the propose d sub grap h is enc oded using the s ame enco ding neur al message pass ing net work to c reate with add itional labe ls to atoms an d bo nds s ignif ying unk nown conn ections, w hic h produc es an enco ding for . Deep learn ing is the n used to determine the ran k of the next possi ble subg raph g iven the pre vious select ed fragment ( ) and the connect ivity o f the fina l subst ructure . By usin g ci rcular fragm ents w ith r adius 1 w he n de coding t h e connectiv ity, one c an ex ploit la rger c ircular f ragme nts . For example, cons ider the c ombinat ion o f t wo fr agments a long an edge. Since edges are i nherentl y one di mensi onal, the combinat ion of t wo circ ular fr agments w ith radius o f 1 centered on at oms alwa ys res ult s in a ne w circular fragm ent centered a round a bo nd w ith rad ius 1 (Figur e 3). The circul ar fragme nts centered on bon ds with radius 1 mus t be include d once a nd o nly once from a b ag w hen rebu ilding t he m olecu lar graph. As t hese bond fragm ents have set rad ii, un directe d neural mes sage pass ing net works are c apable of p rodu cing unique fi ngerpr ints, s imilar to E CFP, e ach tim e a frag ment i s added to the eme rging grap h 15 . Interestingly , most funct ion al groups h ave a set circ ular rad ius aro und a partic ular poi nt , such as ami nes, s ulfones , nitri les and m any m ore. Thi s sugges ts that this fragment -base d scheme c ould be a n appropriat e bas is set t o descr ibe a molecul ar funct ion al s pace . Furt hermore, i f eac h fragme nt cent ered on a n atom wit h radius 1 is unique, know ing the set of frag ment s centered o n bonds with r adius 1 is s uff icient infor matio n t o re const ruct the graph. H owever, most fragment bags with radi us 1 are n ot unique, t herefore higher r adius informati on is re quir ed. T his would n ot be true if the nodes in the fragme nts w ere clust ers of at oms, simi lar t o the Jin et al. J uncti on T ree Autoenc oder 5 . As fragment clust ers tend to be uni que for reaso nably size d molecular grap hs (30 at oms ), knowi ng t he set of un ique molecular fr agme nts cent ered on edges w ould be s ufficie nt informat ion to reconst ruct the connec tivit y o f t he clust ers. Furt hermore, using cl ust ers w ould p revent the trans form atio n of e mergi ng molecula r structures converting betw een different t opologies . He re we focus on a dif ficult reconst ruction tas k o f using at oms as no des. How ever, it should be not ed t hat this clust er-b ased fragment ap pro ach is a logical e xtension of t his sche me a nd sh ould be e xplored in more deta il. Unfortun ately, by incor poratin g only a s ingle f ragme nt into the emerging stru cture, the c ontinu ous add ition of fr agment s doe s not nec essitat e equiv alent un ique i dent ifiers for larger radiu s fragme nts beyon d a rad ius of 1 in t he emerg ing st ructure due to t he presence of dang ling b onds co mpa red to the enc ode d graph. This is e xemplified by a non -circular frag ment whic h cannot be uni quel y ide nt ified as a si ngle c ircul ar fr agme nt around an y atom or bond w ith any r adius Figur e 3. This is because t he d ecode r does n ot hav e know ledge of the l arg er molecular st ructure b eyond t he dan gling bon d in t he em ergin g graph. T his is i mport ant as when t he decoder compares t he encoding of the origina l and e mergin g graph the dec od er compare s p artial t o co mplet ed circ ular f ingerp rints. Thi s means t he aut oencod er l earns how p arti al non -circul ar fragme nts are sub fragme nts of larger cir cula r fragment s. W it h the assumpt ion it is e asier for t he dec oder t o com par e complete circu lar fragme nts, w e bias the t raining data t o favour the additio n of fr agments t o bonds, whi ch increas es th e maximum radius of c ircular frag ments . H owever, as t he molecule is g ener ated fr agment by fr agme nt, it is t rivial t o determine the ma ximum circular r adius o f each fr agme nt. T o directly enc ode ring fr agme nts, w e inco rpor ate a n orthogo nal MPNN dedi cated t o t he commun icatio n of mes sages o nl y along b onds in ring s, making the net work capab le of generating a u nique fi ngerpr int for each ring frag ment. One out standing issue of this approac h is that it is p oss ible t o build mo lecul arly in valid st ructure s. For ex ample , t his metho d could gen erate a molecule with a sing le d angling aromat ic bond and inapp ropr iate a rom atic ri ngs. T he fragme nt bag could also be inc ompl ete to recons truct a molecul e. Most of thes e failure s ar e av oided b y hard co ding rules t o preve nt generatio n an d ad dition of cert ain ch emicall y implaus ibl e fragme nts. To avoid an i ncom plete fragment b ag, one coul d train the fragme nt dec oder t o always deco de a complet e set or excluding fr agme nts w hich do not ha ve a com plement ary fragme nt. A l imitat ion o f t his m od el is that it does not co ntain any geom et ric inf ormati on, hence , it is unable to d istinguis h stere oisomers. Th is co uld pos sibly b e addres sed by i nclu ding Figure 3 Exa mples of circular f ragments centred on atom s and bonds with radius 1 and 2 and a non-circu lar frag ment which cannot be uniquely i dentified from any bond/atom with a set radius. The ML alg orithm learns how t o describe circular fingerprints such that non-circular fragment are related appropriat ely. arXiv Fra gment Gra phical Vari ational Aut oEn c oding f or Screening M olecules wi th S mall Data J . A r m i t a g e , 2 0 1 9 | 5 geometric informat ion (s uch as rel ative d istance and c hira lit y ) and me ssage p assing d irectly t o the next neares t nei ghbors . Even t hough it is poss ible for t he autoen coder to n ot decomp ose an d re const ruct the molecule perfect ly, th e purpose here is t o us e a n u nsuper vised learnin g algorit hm to remove redu ndant info rmation whi le descr ibing the relatio n betw een fragment s. In d oing so t his appr oac h produces a n orthogon al and comp lete represe ntat ion o f all fragme nts in a reduced basis s et compare d to standar d ECFP. Due to l imite d computat ional reso urces w e ha ve n ot ex plored th e optimizati on procedur e for selectin g m odel hy perp aramet ers that t rade o ff bet ween com pleteness , orthon orma lity and basis se t size which sh ould be e xplored in future w ork. In thi s work w e select int uitive dime nsional ity reductio ns and hyperp aramet ers. F or example , in ou r organi c a dditive s dataset we reduce t he dimens i ona lity of t he ECFP w ith maximum rad ius 3 from 9 3 7 to 3 0 using Fra GVAE wh ich is l ess than t he numbe r of tra ining examp les (6 9). Results Predicti ve perfo rman ce of che mic al finge rprints in sm all c alculate d datasets Here w e compare t he predi ctive perf orma nce of variou s chemic al fingerp rints traine d on r andom sm all su bsets (10 t o 100) an d t ested on ran dom larger s ubs ets (500 t o 100 0) of big dataset s. T he molecul ar finger prints m ethods inc lude extended -con nectivity fingerprint s, ChemV AE, ran dom FraGVAE an d F raGVAE . E xtende d-connec tivit y fi ngerprint s ECFP is a s tanda rd circul ar fragment b ased fing erpri nt technique use d by c hemist s, w hich prov ides a b inar y identi fier for e ach uniqu e circle fragme nt of a set rad ius 16 . C he mVAE is the standard st ring based molec ular aut oenc oder use d for automatic chem ical desig n co mmonly cited in t he literature 4 . ChemV AE conve rts the s implif ied mo lecular -input line -entr y syst em (S MILES ) repres ent ations of a mole cule t o a on e -h ot encoding which is a utoenco ded using s tand ard n atur al language pr ocessing t echniq ues. FraGV AE with fixed rando m small weig ht s w ere chosen, a s graph ica l conv olutions wit h fixed small random we ights can be an appropri ate fingerpri nt and the di fference bet ween t he r andom and t he tra ine d FraGVAE can be att ributed t o the Fra GVAE mode l learnin g 15 . Here w e p redict the the oretica l octan ol/wat er p artit ion coefficient (logP) , quantit ative eff ective dr ug score (QED ) and synthet ic acc ess ibility score (S AS) from 250 ,000 ran dom struc tures from t he Zinc1 5 datab ase calc ulated b y RDKIT 4,17 ,1 8 . The Z inc15 dataset w as ch osen as logP, QED and S AS are we ll establis hed e xperime ntal in dicat ors o f su itable m olecu lar struc tures wit h robus t the oretical mo dels 3 . In a dditio n, mo st predictive perfor manc es of the a utoenc oded latent s pace models are t est ed o n the sam e ran dom Z inc15 dat aset fro m Gomez et. al. using 10 -fol d cros s validati on 4,5, 17,19,2 0 . Autoenco ders report t heir pre dictive per formance of l og P, QED and S AS as the y are typica lly used as ge nerat ive mo dels t o generate new mole cular st ructures with optim al va lues. I n additio n w e a lso c omp are the e xper imenta l s olubi lity of molecules in aq ueo us solut ions from t he ESOL datasets which are co mmo nly used to be nch mark no vel MPNN i n big dat a applicat ions 21,2 2 . Prior to t raining t he pr edict ive mode ls, w e t rain F raGVAE t o reconst ruct mo lecu les in the d ataset s, which include bot h th e training and tes t data. In most ML mode ls one a lway s separates t he t raining and t he test s et, here we train our autoenco der t o rec onst ruct molec ules in bot h the t raini ng an d test set (candidate se t). Training the autoenc oder in this manor allows us t o us e unsupervis ed learning to s ort the tes t set molecules i n rel ation to other molecules in the tra inin g s et. This appr oac h red uces t he amo unt of in form ation required t o find appr opri ate mo lecules in t he tes t set . To comp are the pred ictive pe rfor mance of c hemi cal fingerpr ints traine d on s mall dat asets , we t rain a nd t est a Figure 4: Predicative perf ormance of chemical fing erp rints p redicting logP, SAS and Q ED as a function of training size de pendence. The shaded area correspon d to one standard deviat ion of the error. Fragment Grap hical Var iational AutoEnc oding for Scr eening Molecules with Sm all Data arXiv 6 | J . A r m i t a g e , 2 0 1 9 number of rando m forest mode ls on r andom subset s of th e dataset and com pare the root mean squ ared err or (RMS E), a standard practice, of dif ferent fingerprint ing techniques 22 . I n this small dat a reg ime we directly co mpare ECFP, a tra ined FraGVAE , F ra GVAE wit h fixed r and om s mall w eight s an d ChemV AE fingerpr ints. ChemV AE is a st ring base d molecu l ar autoenco der rep orted by Gomez e t. a l. 4 The ma ximu m number o f basis vect ors for the F raGV AE m odels was selecte d to be on the or der of a hundr ed dime nsions w hich i s comparab le to Che mVAE and t he nu mber of train ing e xam ples in the s mall d ata regime . Spe cific det ails ca n be seen in supplem entary inf ormati on*. The s pecif ic b asis vectors us ed in each model w as determine d in situ by ranking the Pe arson coefficient of each basis vector and selecting t he top nu mber of vectors with t he larg est Pe arson coe fficient which r educe s the RMS E in thre efold cross v alidat ion. These results demonst rate that the FraGVAE fingerpr ints have the bes t predi ctive pe rform ance in the s mall data r egime w hen predicting the logP fr om t he Zinc 15 datab ase and the mole cule solubilit y i n t he s mall d ataset s reg ime bet ween 10 t o 1 00 molecular s truct ures com pared to all other fingerpri nt techniques . T o illust rate this po int t he Fra GAVE mode l require s approxi mately 42 and 60 training exa mples to have the same error that ECFP have with 1 00 exampl es w hen predict ing logP from Zinc 15 an d aque ous s olubil ity f rom E SOL datas ets respect ively. Furthermor e , the area u nder ECFP RMSE curve for F raGVAE is the only model that is co nsis tently p ositive. It is possible the error cou ld be f urther redu ced by tra ining th e FraGVAE method only on the e xamp le used in t he train ing an d test set subs ets instead of the complete d ataset . T his was not atte mpted as this w ould requ ire a larg e numbe r of Fr aGVA E models to deve lop val id stat istics. This sug gests t hat graphic al autoenc oders are p ossibly w ell suited for small dataset s comp ared t o s tand ard finge rprint s, for exam ple ECFP. T his t echniq ue c ould be used in t he larg e data regime as we ll, how ever, MPNN t rained to directl y predict a metric have bee n de monst rated as appro priate for large data applicat ions when there is suf ficient data to av oid over fitting 22 . FraGV AE did n ot reduc e the RMS E err or in th e SAS and QED predict ion in the small d at a re gime; however , it clearly did n ot subs tantiall y incre ase the error sugges ting it is a competit ive fingerp rint techniq ue. For a direct c omparis on of the predictive perf orma nce o f FraGVAE to other autoencoder s in the literature in the larg e dat a r egime, p lease see th e supplem entary informat ion * . Screen ing molecul ar additives for organic s emicon ducto r s with neural p assin g netwo rk fing erp rints To demons trate that gra phical autoenc oding is a reas onabl e strat egy in a real-w orld s ituati on, we dem onstrat e t his approach i n a mo lecul ar o ptimizat ion pr oble m: searc hing f or molecular add itives for organic sem iconduc tors. I n org anic electronics, the relev ant m aterial prope rties such as mob ilit y, electrolu minesce nce, quant um yi e ld an d photov oltai c efficiency are increment ally i mpr oving 23 – 26 . Un fortun ately, t he poor st ability due t o e xtrinsic env ironment al s pecies, su ch a s water and o xygen c ontami natio n, is a well -doc ument ed pheno menon that is i ncreasi ngly limit ing indus tri al applicat ions 27 . O ne p ossible ro ute t o s olve t his prob lem is t o incorporat e liquid or solid -st ate molec ular addit ives whic h improve t he o perati onal and environ mental st ability of conjugat ed po lymers used in field- effec t t ransist ors and diodes 25,2 6,28 . The underlying mechan ism of these addit ives is not entirel y underst ood, but it is be lieved t o be r elated to a n inter actio n with wate r in voids in t he po lymer 25 . F or solvent add itives, w e know that t he for mation of aze otropes pl ays a key ro le i n remov ing w ater relat ed t raps. For solid s tate addit ives on t he other hand, t he mec hanis m is less wel l u nderst ood. Th e mecha nism for the solid state additives is not clear as do pin g and non -dop ing additives im prov e de vice stab ilit y character ist ics, and dir ect spect roscopic e vide nce of t he additives in t he film is chal lengi ng d ue to the sm all samp le siz e and low impur it y de nsit y. It is cha llenging to prob e these v oids and to de termine the exact mo rphology of the mater i al syst em, phys ical i nteractions a nd possib le chem ical byprod ucts to generat e a cle ar e xperiment al pict ure of t he pro ces s. To f ind new mo lecular a dditives , t here is insuffici ent corre latio n al data, theoretica l knowl edge and ex pe rime ntal techniques to create an in disput able mode l o f the mat erial s ystem. To f in d new molecul ar struct ures, one uses e xpert knowl edge t o search for ne w molec ular additiv es, w hich ca n be biase d. H ere we augme nt t his appro ach b y using our F raGV AE mode l to provide a quant ifia ble u nbi ased o pini on as t o whether or not a molecular addit ive w ould impr ove st ability in an org ani c electronic app lication. In order t o use the Fra GVAE as a qu antifiabl e unb iase d opinion, 66 molec ular addit ives w ere t este d (Figure 5). T hey were chosen bas ed on c ost and va riety of functi onal gr oup s which are all known to sh ow interact ion with w ater spe cies (este r, nitri le, phen yl, am ine, nitro , q uinone, s ulfonic a cid, ether, alcoh ol and halog en grou ps). To deter mine whethe r the additives were c apable of i mproving d evice st ability in organic semic onductors , the y were test ed in top -g ate bott om-cont act organ ic f ield effect t rans istors (OF ETs ), w here the org anic sem icond uctor w as the amor pho us pol ymer indacenod ithiop hene - co -be nzoth iadiazo le co polymer (ID TBT). Table 1: Area u nder E CFP R MSE curv e be tween 10 a nd 100 trai ning exam ples. Model Zinc15 logP Zinc15 SAS Zinc15 QED ESOL ChemVAE -18.4 -11.8 -1.41 -11.9 Rnd FraGVAE 5.49 -6.7 0.59 7.9 FraGVAE 8.26 1.74 0.06 17.4 arXiv Fra gment Gra phical Vari ational Aut oEn c oding f or Screening M olecules wi th S mall Data J . A r m i t a g e , 2 0 1 9 | 7 Figure 5: A ll additives in the cross vali dation set w ith their corresponding i dentificati on number (corresponding numb er appears below structure). Molec ules highlig hted by a green sq uare i mproved device characteristi cs. Mole cules marked wi th the blue characters F , R, E a nd C were inacc urately classified by FraGVAE, random FraGVAE, ECFP a nd ChemVAE respectively determine d by LOOCV. Fragment Grap hical Var iational AutoEnc oding for Scr eening Molecules with Sm all Data arXiv 8 | J . A r m i t a g e , 2 0 1 9 During the fabricat ion, the m olecular add itives wer e incorporat ed int o t he devi ce by b len ding the ad ditive solut ions into the IDT-BT so lutions. Due to a large numbe r of addit ives and l ong f abricat ion proce ss, w hich c auses v ariati ons bet wee n batches , w e c lassified t he additives in a Bo olean ( bin ary) manne r to compare the result s. It shou ld be n oted thoug h, that all additives were t est ed against a reference and on ly i n the cas e of a stat istical ly signi ficant improve ment is an addit ive consider ed t o be functio nal. T herefore , the additives wer e classified as functiona l additiv es if they were able t o impr ov e device chara cterist ics throug h a ny proces s, where v oltag e thres hold was re duced by 5V. T he impr ovem ent w ith t he additio n of the add itive T CNQ (red) com pared t o a refere nce sample wit hout additiv es (bla ck) is de monst rated i n Figure 7 , where t he addit ive decre ases the v oltage t hreshol d from -1 9 to -6V. During the sear ch f or new additi ves, we discovere d the soli d - stat e additives unde rgo a c hemical react ion w ith wat er, w hic h correlat es wit h impro ved device c har acterist ics. U n fortun atel y, the che mical re action is non -trivi al and direct evidence of t his reaction occ urring i n the film is ch allenging ( more inform atio n can be see n in t he s uppleme ntary sec tion* F igures 2 t o 8). As a conseque nce, w e do not ha ve a st rong t heoret ical underst anding o f this pro cess. Ins tead, we use a quanti fied struc ture-based approach as an unbias ed opinio n based o n a ll empirical evide nce. Table 2: Predictive performance of dif ferent ch emical fingerprinting t echniques on cross validation and tes t set. NPV and PPV are t he negati ve a nd positive predication values, i.e. t he pe rcentage of ad ditives co rrectly labe lled as n egative and positi ve respectively. ROC-AUC i s the area under the rec eiver op erating characteristics curve. Cross Vali dation Test Method PPV NPV ROC-AUC PPV NPV ROC-AUC ChemVAE 0.94 1 0. 94 0 1 0.60 ECFP 1 1 0.99 0.33 1 0.63 Rnd FragV AE 0.94 0.96 0.94 0. 5 0.67 0.67 FragV AE 0.94 1 0.995 1 0.83 0.90 The comp let e set of all mol ecular s truct ures t ested, the ir identifi cation number and their class ification of whet her th e molecular add itive c ould im prov e devices charact eris tics (highlight ed i n gree n) ar e s een in Figure 5 . B ased on t his s ma ll dataset , we w ould like t o ext rapolate and find n ew m olecul ar struc tures given o ur sm all amount s of dat a. T o ext rapol ate from our gi ven dat a, he re w e t rain s imple linear logisti c regress ion mod els, w hich were opt imized using leave -on e-out cross validat ion. Opt imizat ion was p erfo rmed v ia a grid s earc h. Molecula r f ingerprint tec hniqu es in clud e Fra GVAE, ran do m weight FraGVAE, E CFP and Ch emVAE. T o gener ate the FraGVAE f inger print, we t rai n the F raGVAE t o reco nst ruct a ll molecules in both the tra ining and tes t s et s. F or t he ChemVA E model ther e is on ly a si ngle tra ining example for e ach molecular cand idate, w e use t he Che mVAE m odel tra ined o n the Zinc1 5 dat aset alo ng wit h our ca ndid ate st ruct ures. To test the performance o f t he mode l, we selected a s et of molecular ad ditives ba sed on ch emical intuit ion t o act as appropriat e m olecular additives . The mo lecul es in the test set and t he ir corresp onding r efere nce num ber and cl assific atio n can be seen in Figu re 7. In particu lar w e select HAT -CN 6 (#7 7), DDQ (#78) and Ch loranil (#7 1), w hich are all well -know n electron acce ptors for org anic el ectronics. T he a dditives # 7 2, Figure 7: Top-gat e bottom-contact organic t ransistors with voids in t he organic semiconductor (OSC) fi lm believed to be responsible for def ects sites (left). Example of threshold voltage extracti on from transfer charact eristics of top-gat e bottom-conta ct IDT-BT semiconductor tran sistor with and wi thout TCNQ, demonstrating TCNQ is capable of improving dev ice characte ristics in OFETs (ri ght). Figure 6: A ll molec ular structure s in the test set wi th their corresponding identificati on number. Mole cules highli ghted by a green square improved device c haracteristics. Molecules mar ked with the blue charac ters F, R, E and C were inaccu rately classified by FraGV A E, random FraGVAE, E CFP and ChemVAE r espectively. arXiv Fra gment Gra phical Vari ational Aut oEn c oding f or Screening M olecules wi th S mall Data J . A r m i t a g e , 2 0 1 9 | 9 74, 76, 7 9 and 80 have the 1,1 -dicyan oethene fragme nt observed in alm ost every functi oning add itive i n the t rainin g data. In ad dition, we te sted ad ditives # 70, 73 a nd 75, w hic h have el ectro n-a ccepting gro ups attached t o quino nes, whic h are s imilar struct ural mot ifs to molecu lar additiv es classif ied a s working i n the t raining set . We also synt hesize d add itive # 8 0, which a dds a s olub le side cha in o nto F2 -T CNQ (# 58). Th e modificat ion o f the transfer char acterist ics wit h the addit ives can be seen in the supp lement ary inform ation *. The cr oss validation and test metrics can be s een in Table 1 for variou s fingerprint ing methods. Discussion These results demo nstrat e tha t Chem VAE, w hich ap pear s promisi ng f or B ayesian opt imizati on o f m olecular st ructures i n big dat a co ntext s, does not se em t o be e ffectiv e for sm a ll dataset s. This is believed t o be ca used by the inh erent ly discrete rando m j umps betw een near id entica l molecu l ar struc tures due t he t ext encoding algorit hm nec essity of converting an ar bitrary topology of a mol ecul ar struct ure to a one-dimens ion al obj ect. For exampl e, the c anonic al S MILE S represent ations o f dimeth yl-T CNQ (# 65) are cons idera bl y different t han t he SMILES of both T CNQ (# 69) a nd F 2-TCN Q (#58). Th is w ould r esult in mo lecules wit h near ide ntic al struc tures (an d assu med pro perties) bein g locat ed in completel y di f ferent locations of the latent space and more experiment al dat a is neede d t o fu lfil the Nyqui st criterion . I n big data B ayesian optim ization applic ations, w here t here are appropriat e theoretic al m odels of the s ys tem, N yquist crit eria can be overcome com putat ionall y throug h big data. ECFP w orked e xtre mely well on the t raining s et, as a ll molecular stru ctures in the t est s et can be correct ly classifie d by ident ifyi ng t he pres ence of fragm ent (#72 ) in a r ing or multiple methyl 2-cy ano-3 -methy lcroton ate fragment s (#42 ). This appro ach is over fitting so bre aks dow n in the test s et where not a ll addit ives c lassified as work ing c ontain t he s ame fragme nt, such as additives # 77, 78, 79 a nd 80 , whic h hav e different fr agment s or sm aller sim ilar s ub -fr agments. Fr aGVA E was ab le to detect simi larities between funct ionin g addit ive s in the training and te st set even though there was no obv ious fragme nt correl ation, whi ch is t he major b enefit of us ing graphical enco ded fin gerpri nts. By sorting t he gra ph t hroug h MPNN, the alg orithm c an rec ognise s imil arities betw ee n fragme nts which w ould be ign ored by stand a rd appro ache s which c ount discr et e fragment s. Experim entall y the authors were surpr ised t hat additive # 71 did not wor k as it is a know n orga nic dopa nt. Int eresting ly FraGVAE also p redicted ad ditive #71 as an appr opri ate ad ditiv e based on the tra ining set . This suggest s t hat FraGV AE i ntuitio n was reason abl e to pre dict ad ditive #71 as f unctio ning eve n though it does not contai n e xactly the s ame func tional gr oups that are exclusi vely pres ent in positiv e train ing set exam ples. Conclusions In this work, we addre ss the fu ndament al pr oble ms in appl ying artificial int elligen ce t o t he majorit y of molec ula r optimiz atio n problems . T he obs tacl e in ap plying ML is th at t here is insufficient experime nt al dat a or the oretica l k nowle dge t o build a robust st atist ical mod el to screen c andid ate s truct ures. We propose an approa ch whic h uses graph ica l aut oenc oders to sort molecules base d on their st ructures. As t he graphic al decoders are curre ntly an are a of interes t, we pro pose the f irs t fragme nt based dec ode r w hich reconstru cts a mo lecular gr aph first t hrough t he direct deco ding of sma ll grap h f ragment s, followed by t he recom bin ation o f the fragment s. F inally, w e demonst rate that sorting mole cular g raphs w ith a gr aphic al autoenco der is a valid a pproach to i mpro ve the pre dictiv e accuracy of quant itative struct ural models in the smal l data regime co mpare d t o standard m olecular fi ngerpr ints. We have demonst rated that t his m ethod appe ars us ual ly for org ani c electronic a pplications w ith novel mat erials syst ems w hic h d o not ha ve an es tab lished t heor y and e xperime nt al practice s. This appr oac h ap pears promising for ot her f ields su ch as drug discover y, che mistry and mat erial sc ience. Experimental IDT-BT OFET s where Top-Gate B ott om Co ntact de vices with a W/L of 50. All device s w ere prepared o n C orning 1 737 F subst rates supplied by Pre cision Glass & Optics . IDT -BT wa s supplie d by I.M. an d dissol ved at 10g/L in D CB w ith the molecular add itives ‡ . The bot tom c ont acts w ere go ld. IDT -B T was spun onto the subs trate, bak ed at 90 ° C f or 1hr. A C ytop M layer o f 500n m was depos ited as a dielect ric. Alumi nium g ate s were eva porated via s hadow mask. Conflicts of interest There are no c onflict s to decl are. Acknowledgements J.W.A. ac know ledges d oct oral s upport from t he Canad ian Centenn ial Schol arship F und , Christ's College Cambridg e and FlexEnab le. L.J.S. is grat eful t o Mar cin Ab ram for insig ht ful disc ussion s. I.E.J ac know ledges funding fr om a R oya l Societ y N ewto n Internat ional Fel lowshi p. G.S. acknowledg es postdoct oral fellowship support from the Wiener- Ansp ach Foun dation and T he Leve rhulm e Trust (Earl y Career F ellowsh ip sup ported by the Is aac N ewt on Trust ). I.D. ac knowled ges NanoDT C E RC and Cambridg e Philosophi c al society. Fragment Grap hical Var iational AutoEnc oding for Scr eening Molecules with Sm all Data arXiv 10 | J . A r m i t a g e , 2 0 1 9 Notes and references ‡ For comple te t abl e of all additive s na mes and supplier s along with the corr espondi ng OFET transf er charac teristic s please see the suppl em entary i nforma tion*. * Supplemen tary material will be released with manuscri pt publica tion. 1 G. Mall iaras a nd R. Fri end, Phys. To day , 2005, 58 , 53 – 58. 2 R. Geng, H. M. Luong, M . T . Pham, R. Das, K. S . Repa, J. Robles-garc i a, T. A. Duo ng, H. T. Pham , T. H . Au, N. D. La i, G. K. Lar sen, M . Pha n and T . D. Ng uyen, , DOI:10.103 9 /c9mh0026 5k . 3 G. R. Bick erton, G. V. Paolini, J. Be snard, S . M uresan an d A. L. Hopk ins, Na t. Chem . , 2012, 4 , 90 – 98. 4 R. G m ez-Bom barelli, D . Sheber l a, J. Agui lera-iparrag uirre, T. D. Hirze l a nd R. P. Adam s, ACS Cen t. Sci. , 2018, 268−276 . 5 W. Jin, R. Barzi lay and T. Ja akkola, ar Xiv , , DOI:180 2.04364v 2. 6 R. Góm ez-Bom barelli, J . Agu ilera-Ipar ragu irre, T. D. Hirze l, D. Duv enaud, D. Macla urin, M . A. B lood-Forsy the, H. S . Chae, M. Einzing er, D. -G. Ha, T. Wu, G. Mark opou los, S. Jeon, H. Kang, H. Miyaz aki, M. Numa t a, S. Kim, W. Hua ng, S. I. Ho ng, M . Baldo, R. P. Ada ms and A. Aspuru-Guz ik, Nat. Mater. , , DO I :10.103 8 /nmat47 17. 7 J. Gilm er, S. S. Schoenhol z, P. F . Riley, O. V inyals a nd G. E. Dahl, arXiv , , DO I:10.1002 / nm e.2457 . 8 D. E. Rum elhart and J. L. McC lellan d, in Par allel Di stribu ted Proces sing: Explora tions i n the Mi cros tructure of C ognitio n: Foun dations , M IT Pres s, 1985 . 9 P. Baldi, 2012 , 37 – 50 . 10 B. Cao, L . A. Ad utwum , A. O . Oliyny k, E. J. Luber, B. C. Olsen, A. Mar and J. M. Buriak , ACS Nan o , , DOI:10.102 1 /acsnano .8b0472 6. 11 C. Daska lakis, C. Tzam os and M. Za mpe takis, . 12 L. G. May , Co nstrai ned Gra ph Vari ational Autoe ncoders for Molecu le Desi gn , . 13 R. C. G lem, A . Bender, C. H . Arnby, L. Car lsson, S . Boyer and J. Smith, IDr ugs , 2006, 9 , 199 – 204. 14 G. R. U . Sing a nd V. A. A. U toencoder s, 20 18, 1 – 12 . 15 D. Duv enaud, D. Macla urin, J . Agu ilera-Iparra guirre, R. G´omez -Bombare lli, T. Hirzel, A. As puru-Gu zik and R. P . Adams, D ig. Asi a-Pac ific Ma gn. Rec . Con f. 2010, APMR C 2010 , 2015 , 1 – 9. 16 D. Rog ers and M. Hah n, J. Che m. In f. Model . , 2010, 50 , 742 – 754. 17 T. Ster ling and J . J. I rw in, J. Che m. In f. Mode l. , 2015, 55 , 2324 – 2337. 18 RDKit: O pen-Sourc e C heminform ati c s Soft ware, http://w ww .rdkit.or g. 19 M. J. Kusner, B. Paige and J . Migu el, . 20 H. Dai, Y. Tian, B. Da i, S. Sk iena, L. Song an d A. Financial, 2018 , 1 – 17. 21 J. S. De laney, 2004 , 1000 – 1005. 22 Z. Wu, B. Rams undar, E. N. Fe inber g, J. Gomes, C. Genie sse, A. S. Papp u, K . Leswin g and V. Pand e, Chem. Sci. , 2018 , 9 , 513 – 530. 23 J. Liu, H. Zhang, H. Do ng, L. M eng, L. Jia ng, L. Jian g, Y. Wang, J. Yu, Y. Sun, W. Hu a nd A. J. Heeger , Nat. Comm un. , 2015 , 6 , 1 – 8. 24 X. Gao and Z. Zhao, Sci. China C hem. , 2015 , 58 , 947 – 968. 25 M. Niko lka, I. Nasrall ah, B. Rose, M . K. Ravva, K . Broc h, D. Harkin, J . Charm et, M. Hurhan gee, A. Brown, S . Illig, P. Too, J. Jongman, I. McC ull och, J.-L. Bredas and H. Sirrin ghaus, Nat. M ater. , 2016 , 1 , 356 – 362. 26 M. Niko lka, K. Broch, J. Arm itage, D. Hani fi, P . J. Now ack, D. Venkates hvaran, A. Sadhana la, J . Sask a, M. Masca l, S . H. Jung, J . K. Lee, I. McC ulloch, A. Sal leo and H. S irringhau s, Nat. Co mm un. , 2019, 10 , 1 – 9. 27 D. M. d e Leeu w, M. M . J. Sime non, a. R. Brown and R. E . F. Einerhan d, Sy nth. Met . , 1997, 87 , 53 – 59. 28 M. Niko lka, G . Schwei cher, J . Armi tage, I. Nasra llah, C. Jellett, Z. Guo, M. Hurhange e, A . Sadhanala, I. McC ulloch, C. B. Nielsen a nd H . Sirringha us, Ad v. Mater . , 2018 , 1801874 , 1801 874.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment