Detecting muscle activation using ultrasound speed of sound inversion with deep learning
Functional muscle imaging is essential for diagnostics of a multitude of musculoskeletal afflictions such as degenerative muscle diseases, muscle injuries, muscle atrophy, and neurological related issues such as spasticity. However, there is currentl…
Authors: Micha Feigin, Manuel Zwecker, Daniel Freedman
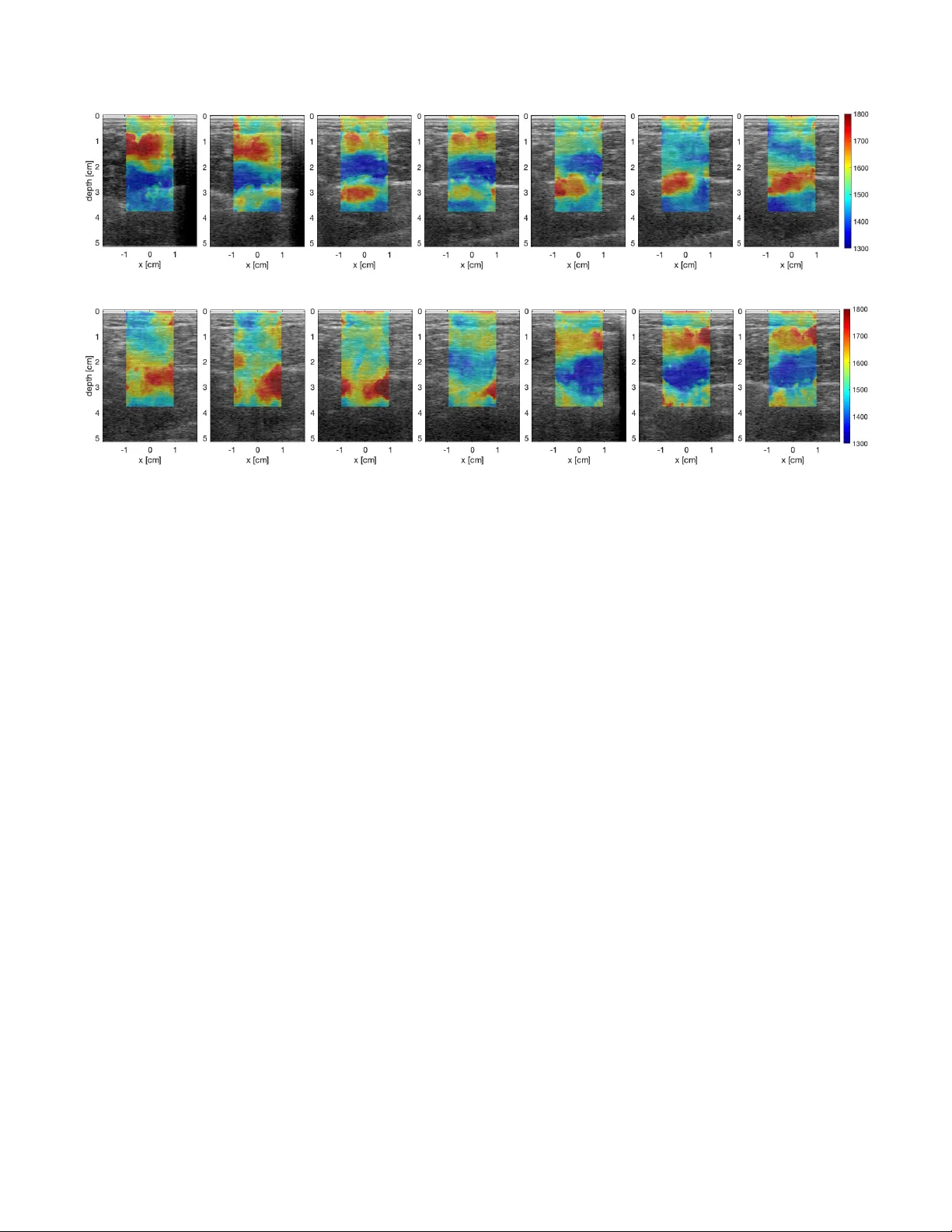
DETECTING MUSCLE A CTIV A TION USING UL TRASOUND SPEED OF SOUND INVERSION WITH DEEP LEARNING Micha F eigin ? Manuel Zweck er †‡ Daniel F r eedman ?? Brian W . Anthony ? ? Department of Mechanical Engineering, Massachusetts Institute of T echnology , USA † Department of Neurological Rehabilitation, The Chaim Sheba Medical Center , T el Hashomer , Israel ‡ Sackler Faculty of Medicine, T el A vi v Uni versity , Israel ?? Google Research, Israel ABSTRA CT Functional muscle imaging is essential for diagnostics of a multitude of musculoskeletal af flictions such as degenerati ve muscle diseases, muscle injuries, muscle atrophy , and neu- rological related issues such as spasticity . Howe ver , there is currently no solution, imaging or otherwise, capable of pro- viding a map of activ e muscles over a large field of view in dynamic scenarios. In this work, we look at the feasibility of longitudinal sound speed measurements to the task of dynamic mus- cle imaging of contraction or activ ation. W e perform the assessment using a deep learning network applied to pre- beamformed ultrasound channel data for sound speed inv er- sion. Preliminary results show that dynamic muscle contraction can be detected in the calf and that this contraction can be positiv ely assigned to the operating muscles. Potential frame rates in the hundreds to thousands of frames per second are necessary to accomplish this. Index T erms — Ultrasound, In verse methods, Muscle 1. INTR ODUCTION Muscle contraction and concomitant muscle force may be reduced in various musculoskeletal and neuromuscular dis- orders such as traumatic brain injury (TBI), cerebral palsy (CP) and multiple sclerosis (MS). For a long time, indi- rect quantification of dynamic muscle contraction has been performed on the joint level (range of motion and joint kine- matics). Intrinsic parameters such as individual muscle force and moment, as well as joint load, have been estimated from in viv o measurements using biomechanical analysis of musculoskeletal models. In recent years, there have been increasing research efforts in developing dynamic measures of musculo-tendon mechanical properties during mo vement using ultrasound (US) based methods, visible in real-time sonography . Elastograph y methods can be used to objec- tiv ely assess muscle viscoelastic properties. Further , assess- ment of muscle contraction velocity enables us to precisely determine the muscle contraction characteristics (eccentric versus concentric) at different time points during movement so that an individual muscle’ s function as an actuator, de- celerator , or a stabilizer for that particular mov ement can be better understood. Muscle contraction velocity may provide evidence about muscle heterogeneity , which has not been extensi vely examined in viv o. In particular , muscle deforma- tion across dif ferent dynamic behaviors, reflecting uniform or non-uniform muscle characteristics may be useful to quantify disparities between healthy and pathological conditions. The architectural changes in muscle that have been studied during static and dynamic contractions include muscle thickness, fiber pennation, and fascicle length. The nature of the rela- tionships between changes in force and muscle architecture varies between muscles and cannot be assumed. US has the potential to measure contractile ability if the nature of the relationship between changes in contraction le vel and muscle dimensions is elucidated. W e look at the problem through an elastic perspectiv e. When muscles contract they stif fen. The Y oung’ s modulus is one of the physical property most closely related to our intuitiv e notion of stiffness. Ultrasound shear wa ve elastog- raphy (SWE) is a method in medical ultrasound that aims to estimate this value. Consequently , the medical ultrasound re- search community has used SWE to assess muscle contrac- tion and muscle force [1, 2, 3, 4, 5]. Ho wev er , reliable results require imaging longitudinally to the muscles (i.e. with the probe oriented along the muscle fibers), thus not pro viding a cross-sectional image of muscle activ ation. The bulk of this work has also been limited to isomet- ric (static) muscle imaging. SWE generally suffers from e x- tremely low frame rates and high sensiti vity to both sonogra- pher and subject mov ement. This makes the method unsuit- able for assessing muscle activ ation during isotonic or other motion, even more so, with large fields of view of muscle cross-sections. Due to power limitations, SWE is also limited to high-end devices. The closest result is the application of SWE to myocardial imaging [6]. These results howe ver also present the physical (a) b-mode (b) Channel data Fig. 1 : B-Mode image of the calf muscles (a), the soleus and gastorcnemius. W e opt to work with raw US channel data (b - cropped) instead as most cues required for SoS in version are discarded as part of the image formation step. limit of the technology , where due to limitations on tissue and probe heating, and the time to activ ate each source, frame rate reached for a 1.4 cm field is 10Hz. Related work has looked into using ultrasound strain imaging [7, 8]. This requires capturing multiple frames at the same point of vie w using different external loads on the ultrasound probe, making it unsuitable for dynamic imaging as well. Preliminary results using electro impedance tomog- raphy [9, 10] do not extend to dynamic imaging either . W e opt to look at longitudinal speed of sound (SoS). SoS depends on Y oung’ s modulus in a similar way to the shear speed of sound. It also depends on the b ulk modulus, which is expected to increase during muscle contraction as well. In this work, we explore the feasibility of SoS in version to the application of assessing muscle activ ation and func- tional muscle imaging based on modifications to a pre viously presented deep learning framework for sound speed inv er- sion [11]. The SoS in version network is applied to the pre- beamformed raw RF channel data generated from a single acoustic plane wa ve (Fig. 1). Cues regarding the sound speed of the domain through which the US pulse has propagated are hidden in the measured signal. Howe ver , most of this infor- mation is discarded as part of the classic delay and sum and en velope detection steps of the b-mode imaging process. Deep learning has seen increasing popularity and success in the fields of vision and medical imaging for such tasks as classification, detection, and segmentation. Howe ver , only recently , deep learning has started being applied to physics and inv erse problems [12] and the imaging pipeline [13, 14]. A neural network can be viewed as a black box performing non-linear regression with a set of parameters that control the relationship of the output on the input. The structure of the network af fects both training properties as well as regulariza- tion, through the expressibility of the network. In this case, 1x64x2048 32x64x1024 32x64x512 32x64x256 32x64x128 64x32x64 128x16x32 512x8x16 (128+128)x16x32 (64+64)x32x64 (32+32)x64x128 32x128x256 1x128x256 Strided 3x15 conv Strided 3x13 conv Strided 3x11 conv Strided 3x9 conv 3x7 conv + maxpool 3x5 conv + maxpool 3x3 conv + maxpool 3x3 conv + interpolate 3x5 conv + interpolate 3x7 conv + interpolate 3x9 conv + interpolate 1x1 conv Fig. 2 : T opology of the deep neural network used in this word. Blue denotes the do wn sampling steps, orange the up sam- pling steps and green the output stage. Arrows show skip connections used. we wish to train the network to take raw channel data as in- put and produce sound speed maps as output. This is done by training it using simulation data. 2. METHODS For this work, we modified the network presented in [11]. Our modifications are presented in Fig. 2. For sound speed in version, we use a fully con volutional neural network in an encoder-decoder topology , taking as input the ra w RF ultra- sound channel data, 64 channels by 2048 samples resulting from a single plane w av e transmit. The network compresses the signal down to 8 by 16 samples with 512 features and expands it back to produce a 128 by 256 sound speed map. Due to the 2:1 decimation f actor at each step, a filter size that complies with Nyquist sampling requirements is used. At the top lev el, the filter size is 3 channels by 15 samples, gradually decreasing in size to 3 channels by 3 samples, and increasing again in the upsampling steps to 3 by 9. This improved pre- vious results while also removing block artifacts presenting with smaller filter sizes. Skip connections were added to the three innermost layers. Due to a lack of an existing expert system capable of generating ground truth speed of sound maps, it is practi- cally impossible to generate a large (or in fact, an y) classified dataset on real data for training purposes. The network was thus trained on simulation data. W e generated 6000 training samples and 800 testing samples by randomly generating be- tween 1 and 5 spheres over the domain. Spheres were used (a) Anatomy (b) Squat Fig. 3 : Experimental setup. Image (a) sho ws the cross-section of the imaging anatomy , the calf muscles. W e image the gas - trocnemius and soleus muscles from a posterior view . The subject performs a squat maneuv er , transitioning from stand- ing to kneeling and back while standing on tip-toes. W e ex- pect the outer gastrocnemius to be acti ve (red) in the standing position and the soleus to be relax ed (blue), and vice versa in the kneeling position. due to simplicity , while still being close enough in shape to most org ans. Attenuation and density were kept constant at 2.5 dB/cm and 0.9 gram/cm 3 . Sound speed v aried between 1300 m/s and 1800 m/s. Random speckle noise was added to the data as well as quantization and random Gaussian noise. 3. EXPERIMENT AL RESUL TS For this work, we wish to sho w that we can dif ferentiate be- tween activ e and non-active or relaxed muscles. T o this end we chose to look at the soleus and gastrocnemius (GC) mus- cles locates on the backside of the calf from a posterior view , as sho wn in Fig. 3. This was done for two reasons (1) we can see both muscles next to each other in the same ultrasound frame (2) the two muscles are active at different joint posi- tions. This allo ws us to see the activ ation pattern shift from one muscle to the other . The two muscles are responsible for performing plantar flexion of the foot (with the GC also flexing the knee). Both muscles are connected at the Achilles tendon on the lower end. On the upper end, the GC connects to the femur while the soleus connects just below the knee. As a result, the GC performs most of the work with a straight leg while the soleus activ ates with a bent knee, as the GC deactiv ates due to the shortening of the muscle. T o image the work transitioning between the two muscles, we asked a healthy human subject to perform a squat maneu- ver , i.e. transition from a standing position to a crouching position, and back to a standing position, while standing on tiptoes (Fig. 3). Experiments were carried using a protocol approv ed by the MIT Committee on the Use of Humans as Experimental Subjects (COUHES). Data was collected with a 128 element linear probe with a center frequency of 5 MHz using a Cephasonics Cicada ultrasound system. Our experimental results are presented in Fig. 3, with Fig. 3(a) sho wing the response for the transition from stand- ing to crouching (down phase) and Fig. 3(b) for the transition from crouching to standing. Red color denotes higher sound speeds that correlate with contracted muscle, while blue color denotes lower sound speeds, that correlate with relaxed mus- cle. Results in both cases match the expected action. In the down phase, we see the task start with the GC (outside of the le g / top part of the image) with the soleus starting to ac- tiv ate on the third frame, and slowly transition to the soleus, where we see that the GS slo wly relaxes and the soleus slowly contracts. On the up phase, we see the complementary beha v- ior , where the work is initially carried out by the soleus, and slowly transitions to the GC. While the sound speeds we are currently recov ering are out of the expected range, due to limitations on domain transfer of 2D simulations based training to real work mea- surements, the functional results show that the method is extremely capable at functional muscle imaging with strong potential for improving the results farther . 4. CONCLUSION In this work, we ha ve presented a longitudinal speed of sound based technique for dynamic functional muscle imaging. Re- sults are based on a simulation-based deep learning method for sound speed in version in ultrasound as applied to plane wa ve channel data. Highly encouraging results sho w dif ferentiation of muscle activ ation with potential frame rates of hundreds to thousands of frames per second. This novel technique has the potential to assess dynami- cal muscle contraction with high resolution and capability of muscle delineation. 5. REFERENCES [1] M. Shinohara, K. Sabra, J. L. Gennisson, M. Fink, and M. L. T anter , “Real-time visualization of muscle stiff- ness distribution with ultrasound shear wave imaging during muscle contraction, ” Muscle and Nerve , v ol. 42, no. 3, pp. 438–441, 2010. [2] F . Hug, K. T ucker, J. L. Gennisson, M. T anter , and A. Nordez, “Elastography for muscle biomechanics: T o- ward the estimation of individual muscle force, ” Exer c. Sport Sci. Rev . , vol. 43, no. 3, pp. 125–133, jul 2015. [3] J. E. Brandenb urg, S. F . Eby , P . Song, S. Kingsley-Ber g, W . Bamlet, G. C. Sieck, and K. N. An, “Quantifying passiv e muscle stiffness in children with and without cerebral palsy using ultrasound shear wav e elastogra- phy , ” Dev . Med. Child Neur ol. , vol. 58, no. 12, pp. 1288–1294, 2016. (a) Down phase (b) Up phase Fig. 4 : Results for a squat maneuv er . (a) shows the results for the transition from standing (left) to crouching (right) and (b) for the transition from crouching to standing position. Fig. 1(a) sho ws the location of the muscles in the frame. Blue denotes lo wer sound speeds, correlated with relaxed muscles, red denotes higher sound speeds, correlated with contracted muscles. Both cases show that the soleus (inside of the leg - bottom of the image) is doing most of the work in the crouching position while the gastrocnemius (outside of the leg - top of the image) is doing most of the w ork in the standing position. [4] J. Ryu and W . K. Jeong, “Current status of muscu- loskeletal application of shear wa ve elastography , ” Ul- trasonogr aphy , vol. 36, no. 3, pp. 185–197, jan 2017. [5] K. M. M. Lima, J. F . S. Costa Jr ., W . C. A. Pereira, and L. F . de Oliveira, “ Assessment of the mechanical proper- ties of the muscle-tendon unit by supersonic shear wa ve imaging elastography: a re view , ” Ultrasono graphy , v ol. 37, no. 1, pp. 3–15, jan 2018. [6] S. J. Hsu, R. R. Bouchard, D. M. Dumont, P . D. W olf, and G. E. T rahey , “In V i vo Assessment of Myocardial Stiffness with Acoustic Radiation F orce Impulse Imag- ing, ” Ultrasound Med. Biol. , vol. 33, no. 11, pp. 1706– 1719, nov 2007. [7] A. J. Pigula, J. S. W u, M. W . Gilbertson, B. T . Darras, S. B. Rutkov e, and B. W . Anthony , “Force-controlled ultrasound to measure passi ve mechanical properties of muscle in duchenne muscular dystrophy , ” in EMBC , aug 2016, pp. 2865–2868. [8] R. S. Witte, K. Kim, B. J. Martin, and M. O’Donnell, “Effect of fatigue on muscle elasticity in the human fore- arm using ultrasound strain imaging, ” in EMBC , aug 2006, pp. 4490–4493. [9] O. L. Silva, Muscle Contraction Detection Using Elec- trical Impedance T omo graphy , Phd, Escola Polit ´ ecnica da Univ ersidade de S ˜ ao Paulo, 2012. [10] E. K. Murphy , J. Skinner , M. Martucci, S. B. Rutkov e, and R. J. Halter , “T ow ard electrical impedance tomog- raphy coupled ultrasound imaging for assessing muscle health, ” IEEE T rans. Med. Imaging , v ol. 38, no. 6, pp. 1409–1419, jun 2019. [11] M. Feigin, D. Freedman, and B. W . Anthon y , “ A deep learning frame work for single-sided sound speed in ver - sion in medical ultrasound, ” IEEE T rans. Biomed. Eng. , jul 2019. [12] M. Raissi, P . Perdikaris, and G. E. Karniadakis, “Physics informed deep learning (part II): Data-driv en discov ery of nonlinear partial differential equations, ” arXiv1711.10566 , Mar 2017. [13] D. Perdios, A. Besson, F . Martinez, M. V onlanthen, M. Arditi, and J.-P . Thiran, “On problem formulation, efficient modeling and deep neural networks for high- quality ultrasound imaging : In vited presentation, ” in CISS , mar 2019, number 20, pp. 1–4. [14] A. A. Nair , M. R. Gubbi, T . D. T ran, A. Reiter, and M. A. L. Bell, “ A fully con volutional neural network for beamforming ultrasound images, ” in IUS , oct 2018, pp. 1–4.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment