Adaptive template systems: Data-driven feature selection for learning with persistence diagrams
Feature extraction from persistence diagrams, as a tool to enrich machine learning techniques, has received increasing attention in recent years. In this paper we explore an adaptive methodology to localize features in persistent diagrams, which are …
Authors: Luis Polanco, Jose A. Perea
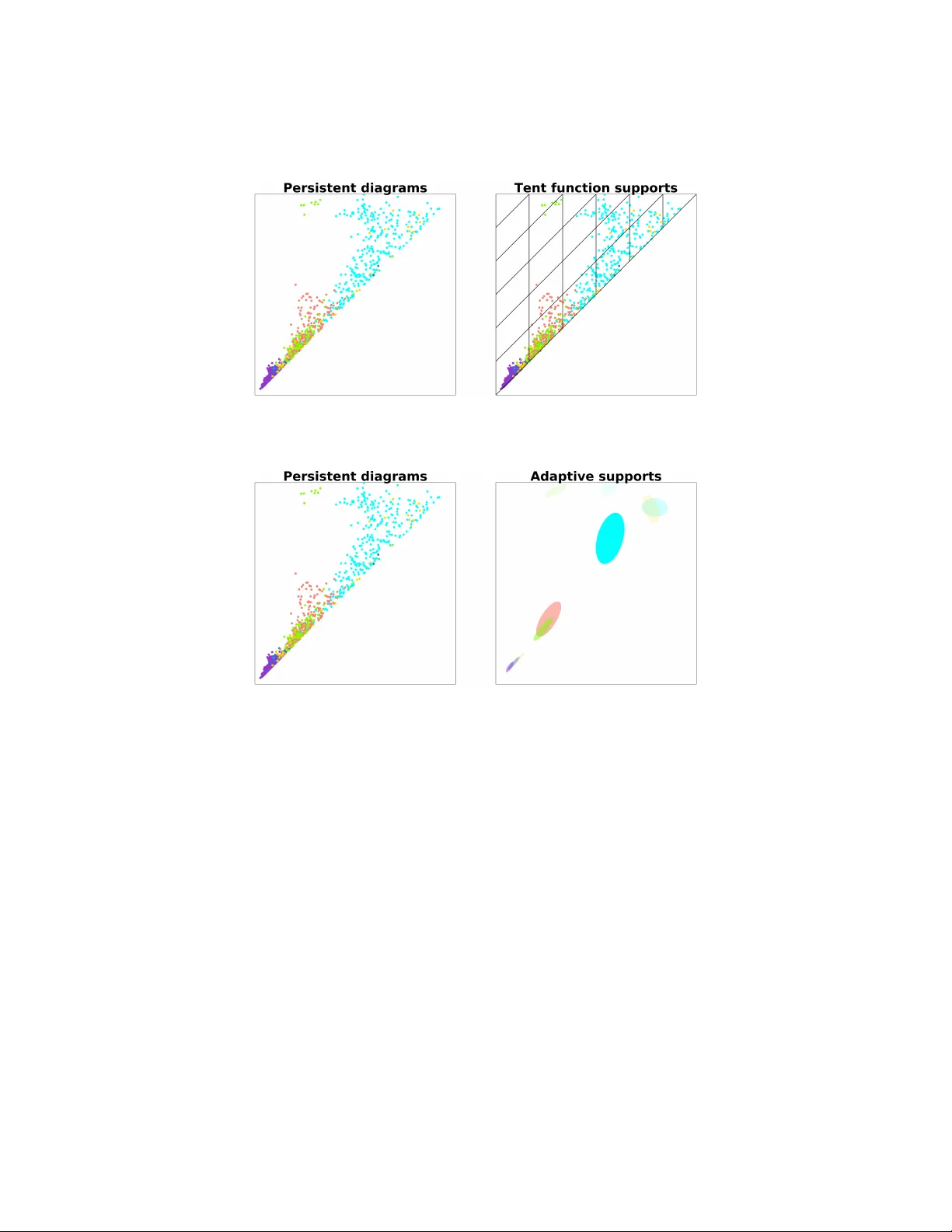
AD APTIVE TEMPLA TE SYSTEMS: D A T A-DRIVEN FEA TURE SELECTION F OR LEARNING WITH PERSISTENCE DIA GRAMS LUIS POLANCO AND JOSE A. PEREA Abstract. F eature extraction from persistence diagrams, as a to ol to en- rich machine learning techniques, has received increasing attention in recen t years. In this pap er w e explore an adaptive methodology to lo calize features in persistent diagrams, whic h are then used in learning tasks. Sp ecifically , w e i n- vestigate three algorithms, CDER, GMM and HDBSCAN, to obtain adaptiv e template functions/features. Said features are ev aluated in three classification experiments with p ersistence diagrams. Namely , manifold, human shap es and protein classification. The main conclusion of our analysis is that adaptiv e template systems, as a feature extraction tec hnique, yield competitive and often sup erior results in the studied examples. Moreov er, from the adaptive algorithms here studied, CDER consistently provides the most reliable and robust adaptive featurization. 1. Introduction One of the central questions in T op ological Data Analysis (TD A) is ho w to lev erage top ological information, like persistence diagrams [7], for machine learning purp oses. This idea has b een explored, for instance, in [2, 4, 8, 10] and [1]. In particular, [8] establishes theoretical and computational to ols to translate su- p ervised mac hine learning tasks (e.g., classification and regression) with topological features, in to the problem of appro ximating con tinuous real-v alued functions on the space of p ersistence diagrams, D , endo w ed with the Bottleneck distance. The main concept is that of templates. These are con tinuous real-v alued compactly supported functions on W := ( x 1 , x 2 ) ∈ R 2 | 0 ≤ x 1 < x 2 , which (b y in tegration against p er- sistence measures, whereb y a diagram is replaced b y a sum of Dirac deltas) yield con tinuous functions on D . The same work sho ws that one can construct countable families of template functions (a template system), which in turn giv e rise to dense subsets of C ( D , R ) with resp ect to the compact-op en top ology . Theorem 2.8 b elow indicates ho w a template system can b e utilized to generate (p olynomial) features for sup ervised mac hine learning problems on p ersistence diagrams. In this pap er we address the question of pro ducing template systems that are attuned (adaptive) to the input data set and the sup ervised classification problem at hand. W e explore and compare differen t strategies to assem ble adaptiv e template systems; namely , Cov er-T ree Entrop y Reduction (CDER) [12], Gaussian Mixture Mo dels (GMM) [11] and Hierarchical densit y-based spatial clustering of applications 2010 Mathematics Subje ct Classific ation. Primary 55N99, 68W05; Secondary 55U99. Key wor ds and phr ases. T op ological Data Analysis, Persisten t Homology , Machine Learning, F e aturization. This work was partially supported by the NSF (DMS-1622301). 1 2 LUIS POLANCO AND JOSE PEREA with noise (HDBSCAN) [3]. The conclusion is that CDER is the most consisten tly successful strategy out of the ones explored. W e presen t three differen t examples where w e use adaptive template functions to extract features from persistence diagrams for supervised classification tasks. First, w e explore a 6 class classification problem presented in [8]. In this problem, sev eral random samples are taken form each of 6 manifolds, and p ersistence diagrams are computed as descriptors in eac h case. W e then use our adaptive template functions and compare to the results provided in [8]. The av erage classification accuracy of adaptiv e templates for b oth the training and testing sets is comparable to that of [8]. On the other hand, the standard deviation of our results is muc h smaller, making our metho dology more stable compared to the template systems prop osed in [8]. W e then rep ort results on the SHREC 2014 synthetic data set [9], whic h in v olves a 15 class sup ervised learning problem. Each class in this data set corresp onds to a human b o dy in five different p oses and three different shap es: male, female and c hild. The data p oin ts in this data set are 3D meshes. In [10] a heat kernel signature is computed for each mesh and for 10 different k ernel parameter v alues. This defines 10 different classification tasks, each of which uses the corresp onding heat sub-level set p ersistence diagrams as inputs. The results we obtain using adaptiv e template functions are con trasted with [8]. The results from this exp eriment highligh t how CDER pro vides a more reliable method for obtaining adaptive templates when compared to GMM and HDBSCAN. F urthermore, when w e select the heat k ernel signature corresp onding to the 6-th frequency , CDER adaptive templates generate a classification mo del with accuracy on par with the best results in [8] and [10]. When compared to the non-adaptive (tent) templates of [8], our classification results are sup erior. Finally , w e presen t results for a protein classification problem on the publicly a v ailable Protein Classification Benchmark data set PCB00019 [13]. This data set con tains spatial information for 1 , 357 proteins as w ell as 55 distinct sup ervised classification tasks. The results on [4] are used as a b enc hmark, since they also use persistence diagrams, but the extracted features are hand-crafted to reflect c hemical/physical prop erties of interest. In this exp eriment, adaptive template functions impro ve the av erage classification accuracy reported in [4] from 82% to around 98%. 2. Appro xima ting Functions on Persistence Dia grams The goal of this section is to provide the theoretical framew ork in whic h template functions are used as a means to approximating contin uous functions on the space of p ersistence diagrams. A p ersistence diagram is a pair S µ = ( S, µ ) where (1) S ⊂ W := ( x 1 , x 2 ) ∈ R 2 | 0 ≤ x 1 < x 2 is such that for any > 0, the set u ( S ) = { ( x 1 , x 2 ) ∈ S | per s ( x 1 , x 2 ) := x 2 − x 1 > } is finite. (2) µ : S → N is an arbitrary (multiplicit y) function. W e will denote the elemen ts of S µ as ( x, m ), where x ∈ S and 1 ≤ m ≤ µ ( x ) is an in teger. Let D denote the set of p ersistence diagrams ; this set comes equipp ed with a metric, the b ottlenec k distance, which w e describe next. ADAPTIVE TEMPLA TE SYSTEMS 3 Definition 2.1. A partial matching M betw een p ersistence diagrams S µ and T α is a bijection b et ween a subset of S µ and a subset of T α ; i.e., M : S 0 µ ⊂ S µ → T 0 α ⊂ T α . If ( y , n ) = M ( x, m ) we say that ( x, m ) is matc hed with ( y , n ). If ( z , k ) / ∈ S 0 µ or ( z , k ) / ∈ T 0 α w e call it unmatc hed . Definition 2.2. Giv en δ > 0, a partial matching M is called a δ -matching if (1) If ( x, m ) ∈ S 0 µ is matched with ( y , n ), then k x − y k ∞ ≤ δ (2) If ( z , m ) ∈ S µ ∪ T α is unmatched, then pers ( x ) ≤ 2 δ . Definition 2.3. The b ottleneck distance d B : D × D → R + is defined as d B ( D 1 , D 2 ) = inf { δ > 0 : M : D 1 → D 2 is a δ -matching } . In [5] it is shown that d B defines a metric on D and that D is the metric com- pletion of D 0 := { ( S, µ ) ∈ D | S is finite } with resp ect to d B . In [8] the authors presen t a complete characterization of (relativ ely) compact subsets of ( D , d B ). In particular, one can prov e that compact subsets of ( D , d B ) hav e empty interiors (see Theorem 13 in [8]). This implies that ( D , d B ) is not lo cally compact and there- fore the compact-op en top ology on C ( D , R ), the space of real-v alued contin uous functions on D , is not metrizable. 2.1. T emplate functions. Lemma 2.4. L et C c ( W ) denote the set of r e al-value d c ontinuous functions on W with c omp act supp ort. F or e ach f ∈ C c ( W ) , the function ν f : D → R define d as ν f ( S µ ) = X x ∈ S µ ( x ) f ( x ) is c ontinuous. Pr o of. See Lemma 23 in [8]. Definition 2.5. A co ordinate system for D is a collection F ⊂ C ( D , R ) with the following property: for an y tw o distinct D , D 0 ∈ D , there exists F ∈ F such that F ( D ) 6 = F ( D 0 ). Remark 2.6. F = C ( D , R ) is itself a co ordinate system, but at least for compu- tational purp oses, it is to o large to b e of algorithmic use. Definition 2.7. A template system for D is a set T ⊂ C c ( W ) such that { ν f : f ∈ T } is a co ordinate system for D . The elemen ts of T are called template functions. The main utility of template systems is that they can b e used to construct dense subsets of C ( D , R ) with resp ect to the compact-op en top ology . In this top ology , whic h is not metrizable as w e men tioned abov e, tw o functions are deemed to b e nearb y if their v alues on compact sets are similar. Since the space of p ersistence diagrams is rather large and complicated, suc h comparisons (weak er than L 2 or k · k ∞ ) are desirable. Theorem 2.8. L et T b e a template system for D , let C ⊂ D b e c omp act, and let F : C → R b e c ontinuous. Then, for any > 0 ther e exist N ∈ N , a p olynomial p ∈ R [ t 1 , . . . , t N ] , and template functions f 1 , . . . , f N ∈ T so that (1) | p ( ν f 1 ( D ) , . . . , ν f N ( D )) − F ( D ) | < for al l D ∈ C . 4 LUIS POLANCO AND JOSE PEREA Pr o of. See Theorem 29 in [8]. In other words, Corollary 2.9. L et T ⊂ C c ( W ) b e a template system for D . Then, the c ol le ction of functions of the form D − → R D 7→ p ( ν f 1 ( D ) , . . . , ν f N ( D )) wher e N ∈ N , p ∈ R [ t 1 , . . . , t N ] , and f n ∈ T , is dense in C ( D , R ) with r esp e ct to the c omp act-op en top olo gy. The problem of constructing template systems of reasonable size (e.g., coun table) is addressed by the following theorem. Theorem 2.10. L et f ∈ C c ( W ) , n ∈ N , m ∈ Z 2 and define f n, m ( x ) = f nx + m n If f is nonzer o, then T = { f n, m | n ∈ N , m ∈ Z 2 } ∩ C c ( W ) is a template system for D . Pr o of. See Theorem 30 in [8]. The goal of this paper is to in vestigate and iden tify data-driv en methodologies for selecting the supp ort of an initial template function f , as well as its most relev ant re-scaled translates f n, m , so that the scalar features ν f n, m can b e used successfully in learning problems on p ersistence diagrams. 3. Adaptive templa te systems In [8] tw o different template systems are suggested: tent functions and interpo- lating p olynomials. Sp ecifically , let f W = { ( x, y ) | x ∈ R , y ∈ R > 0 } b e the conv ersion of W to the birth-lifetime plane. This conv ersion is defined b y ( a, b ) ∈ W 7→ ( a, b − a ) ∈ f W . The template system of tent functions in the birth-lifetime plane is defined as follo ws. Let a = ( a, b ) ∈ f W and 0 < δ < b , then g a ,δ ( x, y ) = max 1 − 1 δ max {| x − a | , | y − b |} , 0 In a similar manner one can define a template system of in terp olating p oly- nomials . Given { ( a i , b j ) } i,j ⊂ f W and { c i,j } i,j ⊂ R , one can use Lagrange in terp o- lating p olynomials to construct a function f such that f ( a i , b j ) = c i,j . In general these tw o approaches require the user to input the meshes used in defining the template systems. By construction, such meshes define the supp ort of the template functions. One shortcoming of this pro cedure, when applied to Theorem 2.8, is that without prior kno wledge ab out the compact set C ⊂ D the n umber of template functions that carry no information relev an t to the problem can b e high. This drawbac k is illustrated in Figure 1. The main goal of this pap er is to present a metho dology to define the template system used in Theorem 2.8 that incorp orates the prior information we hav e ab out the particular learning task. Suc h metho dology is what we refer to as adaptiv e template functions . ADAPTIVE TEMPLA TE SYSTEMS 5 Figure 1. Left: Collection of persistent diagrams colored by class. Right: Mesh co v ering the collection on the left. Figure 2. Left: Collection of persistent diagrams colored by class. Right: collection of open balls as supp orts for template functions. Our approac h to defining adaptiv e template functions is to first identify a col- lection of op en ellipses in f W or W as in Figure 2. Eac h ellipse in this collection will b e the supp ort for a template function defined in the following manner. Let A ∈ M 2 × 2 ( R ) represent the quadratic form in tw o v ariables corresp onding to an ellipse in the collection, and let x ∈ W b e its center. Then, the asso ciated template function is f A ( z ) = ( 1 − ( z − x ) ∗ A ( z − x ) , ( z − x ) ∗ A ( z − x ) < 1 0 , ( z − x ) ∗ A ( z − x ) ≥ 1 T o obtain the collection of ellipses ( A ) men tioned ab o v e we will use and compare three different approaches; namely , Cov er-T ree Entrop y Reduction (CDER, [12]), Gaussian Mixture Mo dels (GMM) and Hierarchical Densit y-Based Spatial Cluster- ing of Applications with Noise (HDBSCAN). W e now pro vide a brief description of eac h metho d. 3.1. Co v er-T ree Entrop y Reduction - CDER. The main ob jective of this al- gorithm is to find a partial co v er tree for a collection of lab eled p oint clouds in R n . 6 LUIS POLANCO AND JOSE PEREA CDER searc hes for the co v er tree in con vex regions that are likely to ha v e minim um lo cal entrop y . In this section we will explain the notion of en trop y used by CDER, as it is relev an t to explaining some of the results in Section 4. Let χ = { X 1 , . . . , X N } b e a collection of p oint clouds X i ⊂ R d — which in our case will be p ersistence diagrams — and define χ := N G i =1 X i . W e also ha v e a labeling map at the level of p oin t clouds λ : χ → { 1 , . . . , L } := L . Notice that w e hav e a natural map ind : χ → χ giv en by ind ( x ) = X i if x ∈ X i . This allow us to define χ l := ( λ ◦ ind ) − 1 ( l ) , the set of all p oin t in a point cloud labele d l ∈ L . No w we will assign weigh ts to eac h p oint x ∈ χ in the following manner: (1) Eac h lab el is equally likely among the data and χ has a total w eight of 1. Th us each lab el l ∈ L has an allocated weigh t of 1 /L . (2) No w consider λ − 1 ( l ) = { All p oin t clouds with label l } , w e assume again that eac h point cloud in λ − 1 ( l ) is equally lik ely , so eac h X i ∈ λ − 1 ( l ) has an allo cated weigh t of 1 / ( N l L ), where N l = λ − 1 ( l ) . (3) Finally each p oint in x ∈ X i is equally likely , and th us w ( x ) = 1 k X i k N l L Definition 3.1. Let χ = X 1 , . . . , X N | X i ⊂ R d b e a collection of p oint clouds together with a lab el map λ : χ → { 1 , . . . , L } := L and a weigh t function w : χ → R . F or an y conv ex and compact set Ω ⊂ R d w e define the follo wing quantities: (1) The total weight of l ∈ L in Ω w l (Ω) = X n w ( x ) | x ∈ Ω ∩ χ l o . (2) the total weigh t of Ω W (Ω) = X l ∈L w l (Ω) Using the weigh ts assigned ab ov e we can in terpret w l (Ω) W (Ω) as the probability of a p oint in Ω to ha v e lab el l . This leads to the following definition. Definition 3.2. Let χ , λ , and w b e as b efore. F or an y conv ex and compact set Ω ⊂ R d its entr opy is defined by S (Ω) = − X l ∈L w l (Ω) W (Ω) log L w l (Ω) W (Ω) . This definition is b orro w ed from information theory . Notice that if all w l (Ω) are roughly the same, then S (Ω) ≈ 1. But, if for example, Ω only con tains p oints with a single lab el then w e m ust hav e S (Ω) = 0. ADAPTIVE TEMPLA TE SYSTEMS 7 Figure 3. Ω 0 ⊂ Ω but S (Ω) < S (Ω 0 ). Remark 3.3. Generally , w e would exp ect the entrop y to decrease as we select smaller subsets of Ω. How ev er, this is not alwa ys the case. F or instance, consider point clouds X 1 and X 2 suc h that | X 1 | = 10 and | X 2 | = 20, eac h p oint cloud with a differen t lab el { 0 , 1 } , and compact sets Ω , Ω 0 as show in Figure 3. W e can easily see that ω 0 (Ω 0 ) = ω 1 (Ω 0 ) = 7 / (20 · 20 · 2), so W (Ω 0 ) = 7 / (20 · 20 · 2) + 7 / (20 · 20 · 2) = 7 / 400. Finally S (Ω 0 ) = 1. A similar calculation will sho w that S (Ω) ≈ 0 . 9182, and thus Ω 0 ⊂ Ω but S (Ω) < S (Ω 0 ). 3.2. Gaussian Mixture Mo dels - GMM. This algorithm is an implementa- tion of the Exp ectation-Maximization (EM) algorithm to fit Gaussian mo dels to a collection of p oin ts. Recall that an EM algorithm is an iterative metho d to solv e maxim um lik eliho o d estimation of parameters for a given mo del; in our case Gaussian mo dels. The EM algorithm iterates o v er tw o steps: an exp ectation step and a maximiza- tion step. The first step defines the expected v alue of the log likelihoo d function using a given set of parameters for the mo del. The maximization step finds a new set of parameters that maximizes the previously described exp ected v alue. 3.3. Hierarc hical Densit y-Based Spatial Clustering of Applications with Noise - HDBSCAN. HDBSCAN [3] is a hierarichal clustering algorithm extend- ing BDSCAN. The latter, finds clusters as the connected comp onen ts of a graph. The vertices of this graph are the elements in the data set after remo ving “noise p oin ts” and the adjacency is defined by a user-provided parameter . HDBSCAN constructs a hierarchical collection of DBSCAN solutions by c hang- ing the parameter used to compute the adjacency in the DBSCAN algorithm. Once this hierarch y of solutions is obtained, the algorithm extracts from its hierarc hy dendrogram a sumarized collection of significan t clusters. 4. Experiment al Resul ts Th us far we hav e developed a metho dology for deriving adaptiv e template sys- tems from lab eled persistence diagrams. The main idea is to use algorithms such as CDER, GMM and HDBSCAN to identify the supp orts of the functions in the 8 LUIS POLANCO AND JOSE PEREA template system. W e will use these adaptive templates to pro duce feature vectors in order to solve supervised classification problems. In this section we presen t three examples of supervised learning, where adaptiv e template functions yield featur- izations that impro v e the classification accuracy or robustness of the classification mo del. Our implementation of adaptive template systems with CDER, GMM and HDBSCAN as well as the scripts to replicate all the results in this section can b e found in the asso ciated GitHub rep ository 1 . Figure 4. Example of the 6 manifolds, from top left to b ottom righ t we hav e: annulus, cub e, 3 clusters, 3 cluster of 3 clusters, S 2 (pro jected on the xy -plane) and torus (pro jected on the xy -plane). 4.1. Manifolds. F or this example we revisit an exp erimen t presen ted in [8] and [1]. W e generated p oin t clouds sampled from different manifolds in R 2 or R 3 . Each p oin t cloud has 200 points and the manifolds considered are the following: an ann ulus with inner radius 1 and outer radius 2 cen tered at (0 , 0), 3 clusters of p oin ts drawn from normal distributions with means (0 , 0), (0 , 2) and (2 , 0) all with standard deviation 0 . 05, 3 cluster of 3 clusters of p oints drawn from normal distributions with standard deviation 0 . 05 and means (0 , 0), (0 , 1 . 5), (1 . 5 , 0), (0 , 4), (1 , 3), (1 , 5), (3 , 4), (3 , 55) and (4 . 5 , 4), cube defined as [0 , 1] 2 ⊂ R 2 , torus obtained from rotating a circle of radius 1 cen tered at (2 , 0) on the xz -plane around the z -axis and sphere S 2 ⊂ R 3 with uniform noise in [ − 0 . 05 , 0 . 05] on the normal direction. W e used CDER, GMM and HDBSCAN to generate the supports of the functions that form our template systems. W e reserved 33% of the data for testing, and trained a kernel ridge regression mo del on the remaining data (%67). In addition, w e inv estigate the effect of increasing the num b er of p oin t clouds sampled from eac h manifold. In T able 1 (see page 9) we present the mean accuracy of the mo del on b oth the training and testing data after a veraging the results o v er 10 experiments for eac h sampling size. F urthermore, T able 1 contains the results obtained from using 1 https://gith ub.com/lucho8908/adaptiv e template systems.git ADAPTIVE TEMPLA TE SYSTEMS 9 T able 1. Manifold classification: Each row corresp onds to the n umber of samples tak en from each manifold. The first tw o columns show the best results reported in [8]. CDER GMM HDBSCAN T rain T est T rain T est T rain T est T rain T est 10 0.99 ± 0.9 0.96 ± 3.2 0.99 ± 0.001 0.98 ± 0.034 0.99 ± 0.001 0.91 ± 0.075 1.00 ± 0.000 0.90 ± 0.092 25 0.99 ± 0.3 0.99 ± 1.0 0.99 ± 0.001 0.99 ± 0.001 0.99 ± 0.004 0.99 ± 0.009 1.00 ± 0.000 0.89 ± 0.031 50 1.00 ± 0.0 0.99 ± 0.9 0.99 ± 0.001 0.99 ± 0.001 0.99 ± 0.002 0.99 ± 0.008 1.00 ± 0.000 0.95 ± 0.003 100 0.99 ± 0.1 0.99 ± 0.4 0.99 ± 0.001 0.99 ± 0.001 0.99 ± 0.004 0.99 ± 0.005 1.00 ± 0.000 0.97 ± 0.011 200 0.99 ± 0.1 0.99 ± 0.3 0.99 ± 0.002 0.99 ± 0.005 0.99 ± 0.002 0.99 ± 0.003 1.00 ± 0.000 0.98 ± 0.005 CDER, GMM and HDBSCAN to find the adaptive templates as well as the best results presented by [8] on the same classification problems. The first important feature to highligh t is that for all the differen t sampling sizes the adaptiv e templates accuracy results show a smaller standard deviation than the results rep orted in [8]. A t the same time, the mean accuracy of our metho dology is comparable with the state of the art results (in [8]). It is w orth men tioning that across all the differen t methods used in this work to obtain adaptive template systems, CDER provides the most stable results. This is sp ecially significant for the smaller sample size (the first ro w in T able 1). 4.2. Shap e data. In this example we consider the syn thetic SHREC 2014 data set [9], of which some instances are shown in Figure 5. W e compare our result to the metho ds rep orted in [8] by extracting features using adaptive template systems for the same data from [10] and [8]. In [10] the authors defined a function on each mesh using a heat k ernel signature, [14], for 10 parameters and computed p ersistent diagrams for dimensions 0 and 1. Figure 5. Examples of shap es and p oses in the SHREC synthetic data set. F or eac h one of the 10 parameter v alues w e ha v e 300 pairs of persistence diagrams and the goal of the problem is to predict the human model. The mo dels corresp ond to 5 different p oses for people labeled as male, female and child; giving us a total 10 LUIS POLANCO AND JOSE PEREA of 15 lab els. Lets us remark that each one of the 10 parameters yields a different classification problem. T able 2. Shap e classification: Portion out of the 10 problems, for whic h each adaptive template system yields the b est classification results. The cells with a dash (-) indicate that no computations w ere carried out. P olynomial RBF Sigmoid Kernel CDER = 0.6 CDER = 0.4 CDER = 0.2 Ridge GMM = 0.3 GMM = 0.4 GMM = 0.4 HDBSCAN = 0.1 HDBSCAN = 0.1 HDBSCAN = 0.4 CDER = 0.7 CDER = 0.7 CDER = 0.8 SVM GMM = 0.3 GMM = 0.2 GMM = 0.2 HDBSCAN = 0.0 HDBSCAN = 0.0 HDBSCAN = 0.0 Random CDER = 0.6 - - F orest GMM = 0.3 - - HDBSCAN = 0.1 - - W e considered CDER, GMM and HDBSCAN as metho ds to obtain adaptive template systems. F or each one of these template systems w e used three differen t k ernel metho ds to solve the classification problems, namely , kernel ridge regression, k ernel supp ort vector mac hines (SVM) and random forest. Finally , three differ- en t k ernels were examined, p olynomial, radial basis function (RBF) and sigmoid k ernels. T able 2 shows the p ortion, out of the 10 problems, for which a given adaptiv e template system yields the b est classification results. F or instance, the entry corre- sp onding to ridge regression with a p olynomial k ernel (first ro w and first column). sho ws that the CDER template system yields the b est classification accuracy in 6 our of 10 problems, GMM is the b est for 3 out of 10 and HDBSCAN is sup erior in 1 out of the 10 problems. With this in terpretation of T able 2 in mind, we can see that CDER adaptive template systems yield more accurate classification results than GMM or HDB- SCAN templates. This holds true for all but one, of the kernel and k ernel metho d com binations explored in this exp eriment. Ha ving stablished how CDER, GMM and HDBSCAN compare across differen t k ernel metho ds and kernels, next we compare our classification results with the state of the art. T able 3 contains our classification accuracy on the training and testing shap e data, as well as the results obtained using the tent functions and in terp olating p olynomials from [8]. The first important feature to remark is that for 7 out of the 10 problems, the mo del obtained from adaptiv e co ordinates has less o verfitting than interpolating p olynomials. Meaning that for most of the classification problems studied in this example, adaptiv e template systems pro vide a more robust classification model. An additional argument in fa vor of the robustness of our approac h is that the standard deviation of all the mo dels using adaptiv e templates is smaller than those presen ted in [8]. Moreov er, when comparing adaptiv e template systems with ten t functions (see T able 3) we attain b etter classification results on the testing set across all the ADAPTIVE TEMPLA TE SYSTEMS 11 T able 3. Shape classification: Classification accuracy for adap- tiv e templates (ours), tent functions and interpolating p olynomi- als [8] Global features Lo cal features In terp olating P olynomials T ent functions Adaptiv e templates F req. T rain T est T rain T est T rain T est 1 0.99 ± 0.3 0.90 ± 5.3 0.08 ± 0.30 0.03 ± 0.5 0.79 ± 0.01 0.73 ± 0.02 GMM Kernel Ridge 2 1.00 ± 0.0 0.95 ± 2.4 0.08 ± 0.40 0.03 ± 1.00 0.84 ± 0.00 0.8 ± 0.03 GMM 3 0.99 ± 0.5 0.90 ± 2.0 0.80 ± 1.3 0.44 ± 4.3 0.7 ± 0.01 0.66 ± 0.03 HDBSCAN 4 0.98 ± 0.9 0.84 ± 3.9 0.89 ± 1.5 0.69 ± 4.9 0.7 ± 0.01 0.67 ± 0.03 GMM 5 0.99 ± 0.4 0.93 ± 2.2 0.76 ± 2.7 0.58 ± 7.9 0.78 ± 0.04 0.76 ± 0.08 CDER 6 0.98 ± 0.5 0.92 ± 1.8 0.96 ± 0.67 0.89 ± 1.7 0.97 ± 0.01 0.92 ± 0.03 CDER SVM 7 0.99 ± 0.4 0.95 ± 1.4 0.98 ± 0.60 0.94 ± 2.5 0.96 ± 0.02 0.94 ± 0.05 CDER 8 0.99 ± 0.4 0.94 ± 2.2 0.91 ± 1.2 0.89 ± 3.3 1 ± 0.00 0.88 ± 0.05 CDER Random F orest 9 0.98 ± 1.3 0.92 ± 2.1 0.64 ± 2.3 0.53 ± 3.8 1 ± 0.00 0.88 ± 0.03 CDER 10 0.97 ± 1.1 0.89 ± 4.6 0.27 ± 3.4 0.18 ± 5.6 1 ± 0.00 0.9 ± 0.08 CDER problems presented. This highlights the b enefits of adaptive v ersus non-adaptive lo cal template systems. It is p ertinent to mention that the results in T able 3 corresp ond to a specific selec- tion of parameters for eac h k ernel and regularization in eac h classification metho d. In fact, from our metho dology we can find a combination of kernel parameters and regularization that yields higher classification accuracy in the training set. But, for suc h mo dels the ov erfitting issues are more noticeable. Finally , since the end goal of this problem is to solve the multiclass classification problem in the syn thetic SHREC 2014 data set. W e can select the problem corre- sp onding to the frequency 6 in the heat kernel signature (row 6 in table 3) as the one that gives us the b est classification accuracy while minimizing the ov erfiting concern. Such result is obtained using a CDER template system and a regularized SVM metho d. 4.3. Protein classification. W e consider next the data set PCB00019 from the Protein Classification Benchmark Collection [13]. This problem set contains 1 , 357 proteins and 55 classification tasks. Our results will b e compared to those rep orted in [4]. T o compare our results with the ones in [4], w e rep ort the av erage classification accuracy ov er the 55 classification tasks in the data set PCB00019. T able 4. Protein classification: a v erage classification accuracy for the 55 tasks in the data set PCB00019. T rain T est P olynomial 0.90 ± 0.07 0.98 ± 0.02 CDER RBF 0.91 ± 0.06 0.97 ± 0.02 Sigmoid 0.90 ± 0.07 0.98 ± 0.02 T op ological features in [4] - 0.82 ± —- T able 4 sho ws the a v erage classification accuracy and standard deviation for eac h of the classification tasks from PCB00019. Here we used a regularized kernel ridge regression with p olynomial, RBF and sigmoid k ernels. The last row in T able 4 displa ys the a verage classification accuracy for the testing set rep orted in [4]. W e 12 LUIS POLANCO AND JOSE PEREA Figure 6. Examples of data p oin ts in PCB00019 and their cor- resp onding p ersistent diagrams. T op row: Protein domains from [6]. Bottom row: Persisten t diagrams in dimensions 0and 1. note that the authors do not rep ort standard deviation or av erage classification accuracy for the testing set. It is meaningful to remark that in [4] top ological features are used to solve the classification problems. Those features w ere constructed using persistence diagrams as to reflect relev an t prop erties of the proteins in the giv en data set. 5. Discussion This pap er inv estigates the viability of utilizing data-driven methodologies to lo calize features in p ersistence diagrams. These features are used in subsequent sup ervised learning tasks for classification problems where shap e is an imp ortant feature. W e examined three different algorithms, CDER, GMM and HBDSCAN to produce adaptive template functions. Through extensive testing with real and syn thetic data sets, w e demonstrate that CDER pro vides a more robust collection of adaptiv e features while maintaining classification accuracy on par with the state of the art. In terms of time complexity , CDER also outperforms GMM and HDBSCAN for the problems here considered. A cknowledgment The authors gratefully thank Elizabeth Munch for pro viding the data set used in section 4.2, Kelin Xia for providing the data set used in section 4.3, and Lida Kanari for providing a data set whic h did not make it into the final v ersion of the pap er. This w ork w as partially supp orted by the NSF under gran t DMS-1622301. References [1] H. Adams, T. Emerson, M. Kirby , R. Neville, C. Peterson, P . Shipman, S. Chepushtano v a, E. Hanson, F. Motta, and L. Ziegelmeier. P ersistence images: A stable v ector representation of persistent homology . Journal of Machine L e arning R ese arch , 18(8):1–35, 2017. [2] P . Bub enik and P . Dlotko. A p ersistence landscap es toolb ox for top ological statistics. arXiv e-prints , page arXiv:1501.00179, Dec 2014. ADAPTIVE TEMPLA TE SYSTEMS 13 [3] R. J. G. B. Camp ello, D. Moula vi, and J. Sander. Density-based clustering based on hier- archical density estimates. In J. P ei, V. S. Tseng, L. Cao, H. Moto da, and G. Xu, editors, A dvances in Know ledge Disc overy and Data Mining , pages 160–172, Berlin, Heidelb erg, 2013. Springer Berlin Heidelberg. [4] Z. Cang, L Mu, K. W u, K. Opron, K. Xia, and W. W ei. A top ological approac h for protein classification. Computational and Mathematic al Biophysics , 3, 2015. [5] D. Cohen-Steiner, H. Edelsbrunner, and J. Harer. Stability of p ersistence diagrams. Discr ete & Computational Ge ometry , 37(1):103–120, Jan 2007. [6] N. F ox, S. E Brenner, and J. Chandonia. Scope: Structural classification of proteins - ex- tended, in tegrating scop and astral data and classification of new structures. Nucleic acids r esear ch , 42, 12 2013. [7] J. A. Perea. A brief history of persistence. pr eprint arXiv:1809.03624 , 2018. https://arxiv. org/abs/1809.03624 . [8] J. A. Perea, A. Munch, and F. A. Khasawneh. Approximating contin uous functions on p er- sistence diagrams using template functions. CoRR , abs/1902.07190, 2019. [9] D. Pickup, X. Sun, P . L. Rosin, R. R. Martin, Z. Cheng, Z. Lian, M. Aono, A. Ben Hamza, A. Bronstein, M. Bronstein, S. Bu, U. Castellani, S. Cheng, V. Garro, A. Giachetti, A. Godil, J. Han, H. Johan, L. Lai, B. Li, C. Li, H. Li, R. Litman, X. Liu, Z. Liu, Y. Lu, A. T atsuma, and J. Y e. Shap e retriev al of non-rigid 3d human mo dels. In Pr oc e e dings of the 7th Eur ogr aphics Workshop on 3D Obje ct R etrieval , 3DOR ’14, pages 101–110, Goslar Germany , German y , 2014. Eurographics Association. [10] J. Reininghaus, S. Hub er, U. Bauer, and R. Kwitt. A Stable Multi-Scale Kernel for T op olog- ical Machine Learning. arXiv e-prints , page arXiv:1412.6821, Dec 2014. [11] D. Reynolds. Gaussian Mixtur e Mo dels , pages 659–663. Springer US, Boston, MA, 2009. [12] A. Smith, P . Bendich, J. Harer, A. Pieloch, and J. Hineman. Sup ervised Learning of Labeled P oin tcloud Differences via Co v er-T ree Entrop y Reduction. arXiv e-prints , page arXiv:1702.07959, F eb 2017. [13] P . Sonego, M. Pacurar, S. Dhir, A. Kertesz-F ark as, A. Ko csor, Z. Gspri, J. A M Leunissen, and S. Pongor. A protein classification b enchmark collection for machine learning. Nucleic acids rese ar ch , 35:D232–6, 02 2007. [14] J. Sun, M. Ovsjanik ov, and L. Guibas. A Concise and Prov ably Informative Multi-Scale Signature Based on Heat Diffusion. Computer Graphics F orum , 2009. Dep ar tment of Comput a tional Ma thema tics, Science & Engineering Dep ar tment of Ma thema tics, Michigan St a te University East Lansing, MI, USA. E-mail address : polanco2@msu.edu Dep ar tment of Comput a tional Ma thema tics, Science & Engineering Dep ar tment of Ma thema tics, Michigan St a te University East Lansing, MI, USA. E-mail address : joperea@msu.edu
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment