Synthesizing Diverse Lung Nodules Wherever Massively: 3D Multi-Conditional GAN-based CT Image Augmentation for Object Detection
Accurate Computer-Assisted Diagnosis, relying on large-scale annotated pathological images, can alleviate the risk of overlooking the diagnosis. Unfortunately, in medical imaging, most available datasets are small/fragmented. To tackle this, as a Dat…
Authors: Changhee Han, Yoshiro Kitamura, Akira Kudo
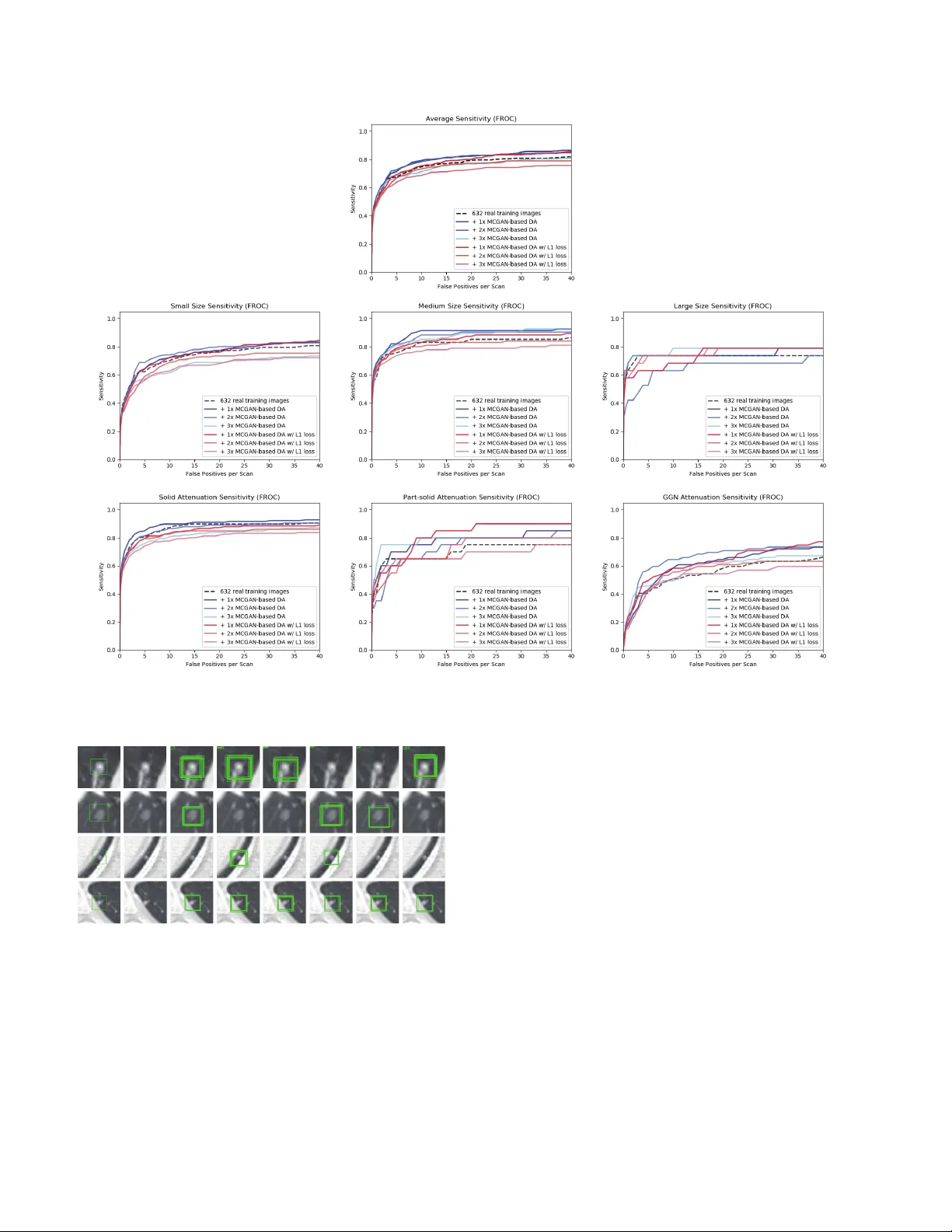
Synthesizing Div erse Lung Nodules Wher ev er Massiv ely: 3D Multi-Conditional GAN-based CT Image A ugmentation f or Object Detection Changhee Han 1 , 3 Y oshiro Kitamura 2 Akira Kudo 2 Akimichi Ichinose 2 Leonardo Rundo 3 Y ujiro Furukaw a 4 Kazuki Umemoto 5 Y uanzhong Li 2 Hideki Nakayama 1 1 The Uni versity of T okyo, T okyo, Japan 2 Fujifilm Corporation, T okyo, Japan 3 Uni versity of Cambridge, Cambridge, UK 4 Jikei Uni v ersity School of Medicine, T okyo, Japan 5 Juntendo Uni versity School of Medicine, T okyo, Japan han@nlab.ci.i.u-tokyo.ac.jp Abstract Accurate Computer-Assisted Diagnosis, r elying on lar ge-scale annotated pathological images, can allevi- ate the risk of overlooking the diagnosis. Unfortu- nately , in medical imaging, most available datasets ar e small/fragmented. T o tackle this, as a Data Augmentation (D A) method, 3D conditional Generative Adversarial Net- works (GANs) can synthesize desir ed r ealistic/diverse 3D images as additional tr aining data. Howe ver , no 3D con- ditional GAN-based D A appr oach exists for gener al bound- ing box-based 3D object detection, while it can locate dis- ease areas with physicians’ minimum annotation cost, un- like rigor ous 3D se gmentation. Moreover , since lesions vary in position/size/attenuation, further GAN-based DA performance requir es multiple conditions. Ther efor e , we pr opose 3D Multi-Conditional GAN (MCGAN) to gener - ate r ealistic/diverse 32 × 32 × 32 nodules placed natu- rally on lung Computed T omography images to boost sen- sitivity in 3D object detection. Our MCGAN adopts two discriminators for conditioning: the context discrimina- tor learns to classify real vs synthetic nodule/surr ounding pairs with noise box-center ed surr oundings; the nodule dis- criminator attempts to classify real vs synthetic nodules with size/attenuation conditions. The results show that 3D Con volutional Neur al Network-based detection can achie ve higher sensitivity under any nodule size/attenuation at fixed F alse P ositive rates and over come the medical data paucity with the MCGAN-gener ated r ealistic nodules—even e xpert physicians fail to distinguish them fr om the r eal ones in V i- sual T uring T est. 1. Introduction Accurate Computer-Assisted Diagnosis (CAD), thanks to recent Con v olutional Neural Netw orks (CNNs), can al- Original Lung Nodule Training Images FLAIR Lung Nodule Detection (Bounding Boxes) (Multi-Conditional GAN) Generate Novel Synthetic 32×32×32 Nodules on CT at Desired Position/Size/Attenuation Additional Lung Nodule Training Images (3D Faster RCNN) Train Figure 1: 3D MCGAN-based DA for better object detection: Our MCGAN generates realistic and di v erse nodules naturally on lung CT scans at desired position/size/attenuation based on bounding boxes, and the CNN-based object detector uses them as additional training data. leviate the risk of ov erlooking the diagnosis in a clinical en vironment. Such great success of CNNs, including di- abetic eye disease diagnosis [12], primarily deri ves from large-scale annotated training data to sufficiently cover the real data distribution. Howe ver , obtaining and annotating such div erse pathological images are laborious tasks; thus, the massiv e generation of proper synthetic training images matters for reliable diagnosis. Researchers usually use clas- sical Data Augmentation (D A) techniques, such as geomet- ric/intensity transformations [29, 23]. Howe ver , those one- to-one translated images ha ve intrinsically similar appear- ance and cannot sufficiently cover the real image distrib u- tion, causing limited performance improv ement; in this re- gard, thanks to its good generalization ability , Generative Adversarial Networks (GANs) [10] can generate realistic but completely new samples using many-to-many mappings for further performance improv ement; GANs showed ex- cellent D A performance in computer vision, including 21% performance improv ement in eye-gaze estimation [33]. This GAN-based D A trend especially applies to medi- cal imaging, where the biggest problem lies in small and fragmented datasets from various scanners. For perfor- mance boost in v arious 2D medical imaging tasks, some researchers used noise-to-image GANs (e.g., random noise samples to div erse pathological images) for classifica- tion [8, 16, 15]; others used image-to-image GANs (e.g., a benign image with a pathology-conditioning image to a malignant one) for object detection [14] and segmen- tation [3]. Howe ver , although 3D imaging is spreading in radiology (e.g., Computed T omography (CT) and Mag- netic Resonance Imaging), such 3D medical GAN-based D A approaches are limited, and mostly focus on segmenta- tion [32, 18]—3D medical image generation is more chal- lenging than 2D one due to expensi ve computational cost and strong anatomical consistency . Accordingly , no 3D conditional GAN-based D A approach exists for general bounding box-based 3D object detection, while it can lo- cate disease areas with physicians’ minimum annotation cost, unlike rigorous 3D segmentation. Moreover , since le- sions v ary in position/size/attenuation, further GAN-based D A performance requires multiple conditions. So, how can GAN generate realistic/div erse 3D nodules placed naturally on lung CT with multiple conditions to boost sensitivity in any 3D object detector? For accurate 3D CNN-based nodule detection (Fig. 1), we propose 3D Multi-Conditional GAN (MCGAN) to gen- erate 32 × 32 × 32 nodules—such nodule detection is clinically valuable for the early diagnosis/treatment of lung cancer , the deadliest cancer [34]. Since nodules vary in position/size/attenuation, to impro ve CNN’ s robustness, we adopt two discriminators with different loss functions for conditioning: the context discriminator learns to classify real vs synthetic nodule/surrounding pairs with noise box- centered surroundings; the nodule discriminator attempts to classify real vs synthetic nodules with size/attenuation conditions. W e also ev aluate the synthetic images’ realism via V isual T uring T est [30] by two expert physicians, and visualize the data distribution via t-Distributed Stochastic Neighbor Embedding (t-SNE) [35]. The 3D MCGAN- generated additional training images can achie ve higher sensitivity under any nodule size/attenuation at fixed False Positiv e (FP) rates. Lastly , this study suggests training GANs without ` 1 loss and using proper augmentation ratio (i.e., 1 : 1 ) for better medical GAN-based D A performance. Research Questions. W e mainly address two questions: • 3D Multiple GAN Conditioning: How can we con- dition 3D GANs to naturally place objects of random shape, unlike rigorous segmentation, at desired posi- tion/size/attenuation based on bounding box masks? • Synthetic Images for DA: Ho w can we set the number of real/synthetic training data and GAN loss functions to achiev e the best detection performance? Contributions. Our main contributions are as follows: • 3D Multi-conditional Image Generation: This first multi-conditional pathological image generation ap- proach shows that 3D MCGAN can generate realis- tic and diverse nodules placed naturally on lung CT at desired position/size/attenuation, which even expert physicians cannot distinguish from real ones. • Misdiagnosis Prevention: This first GAN-based D A method available for any 3D object detector allows to boost sensitivity at fix ed FP rates in CAD with limited medical images/annotation. • Medical GAN-based DA: This study implies that training GANs without ` 1 loss and using proper aug- mentation ratio (i.e., 1 : 1 ) may boost CNN-based detection performance with higher sensitivity and less FPs in medical imaging. 2. Generative Adversarial Networks GANs [10] have rev olutionized image generation [20] via a two-player minimax game. Howe ver , difficult GAN training arises due to its two-player objectiv e function, ac- companying artifacts and mode collapse [11] when gener - ating high-resolution images [27]–especially in 3D or con- ditional image generation; to tackle this, W u et al. pro- posed 3D GAN [37] to generate realistic/div erse 3D ob- jects via a mapping from a low-dimensional probabilistic space; Isola et al. proposed Pix2Pix GAN [17] to produce robust images using paired training samples; Park et al. pro- posed multi-conditional GAN [26] to generate 128 × 128 images from a base image and texts describing desired po- sition. In this way , GANs can usually synthesize more re- alistic/div erse images than other common deep generati ve models, including variational autoencoders [19] suffering from the injected noise and imperfect reconstruction be- cause of a single objective function [22]. Accordingly , as a D A method, most computer vision researchers chose GANs for improving classification [1], object detection [25], and segmentation [39] to o vercome the training data paucity . Also in medical imaging, to facilitate object detec- tion and se gmentation, researchers usually used conditional GANs to generate medical images at desired positions for D A. Han et al. generated 256 × 256 brain MR images with tumors at desired positions/sizes for tumor detection [14]. As 3D GANs for D A, Jin et al. [18] generated 64 × 64 × 64 CT images of both nodules and surrounding tissues—unlike we only generate 32 × 32 × 32 nodules located smoothly on surroundings—for 2D nodule segmentation. Gao et al. [9] generated 40 × 40 × 18 3D subvolumes of nodules for subv olume-based 3D nodule detection via binary classifica- tion; but, the subvolume-based detection accompanies nu- merous FPs, and unlike our work, most other 3D object de- Input volume (w/ surroundings) (w/ surroundings) Synthetic nodule Real nodule 64×64×64×1 64×64×64×1 32×32×32×1 64×64×64×1 32×32×32×1 64×64×64×6 for Generator 32×32×32×6 for Discriminator Input conditions ・ Size (Small/Medium/Large) ・ Attenuation (Solid/Part-solid/GGN) LSGAN loss 32×32×32×64 16×16×16×128 32×32×32×128 32×32×32×64 16×16×16×1 64×64×64×2 8×8×8×256 4×4×4×512 32×32×32×64 16×16×16×128 8×8×8×512 16×16×16×256 3D Nodule Generator Context Discriminator Nodule Discriminator Layers Conv 4×4×4 stride 2 + BN + LeakyReLU Deconv 4×4×4 stride 2 + BN + ReLU Deconv 4×4×4 stride 1 + BN + ReLU Conv 1×1×1 stride 1 + T anh Conv 1×1×1 stride 1 + Sigmoid Concat Dropout 0.5 Fake Pair? Real Pair? WGAN-GP loss 4×4×4×1 32×32×32×7 16×16×16×64 8×8×8×128 (Realism/Size/Attenuation) Fake Nodule? Real Nodule? (Realism/Size/Attenuation) 4×4×4×256 Figure 2: 3D MCGAN architecture for realistic and div erse 32 × 32 × 32 lung nodule generation: The context discriminator learns to classify real vs synthetic nodule/surrounding pairs while the nodule discriminator learns to classify real vs synthetic nodules. tectors cannot use the generated nodules as additional train- ing data since they do not condition nodule positions. T o the best of our knowledge, our work is the first 3D medical GAN-based D A approach using automatic bounding box annotation while 3D bounding boxes re- quire much cheaper annotation cost than rigorous 3D seg- mentation. Moreover , we, for the first time, generate 3D multi-conditional images using GANs. In terms of anno- tation cost, generating realistic and di verse 32 × 32 × 32 lung nodules at desired position/size/attenuation using 3D MCGANs—may become a clinical breakthrough . 3. Methods 3.1. 3D MCGAN-based Image Generation Data Preparation This study exploits the Lung Image Database Consortium image collection (LIDC) dataset [2] containing 1 , 018 chest CT scans with lung nodules. Since the American College of Radiology recommends lung nod- ule ev aluation using thin-slice CT scans [31], we only use scans with the slice thickness ≤ 3 mm and 0 . 5 mm ≤ in- plane pixel spacing ≤ 0 . 9 mm. Then, we interpolate the slice thickness to 1 . 0 mm and exclude scans with slice num- ber > 400 . T o explicitly provide MCGAN with meaningful nodule appearance information and thus boost D A performance, the authors further annotate those nodules by size and at- tenuation for GAN training with multiple conditions: small (slice thickness ≤ 10 mm); medium ( 10 mm ≤ slice thick- ness ≤ 20 mm); large (slice thickness > 20 mm); solid; part-solid; Ground-Glass Nodule (GGN). Afterwards, the remaining dataset ( 745 scans) is di vided into: ( i ) a train- ing set ( 632 scans/ 3 , 727 nodules); ( ii ) a validation set ( 37 scans/ 143 nodules); ( iii ) a test set ( 76 scans/ 265 nod- ules); only the training set is used for MCGAN training to be methodologically sound. The training set contains more average nodules since we exclude patients with too many nodules for the validation/test sets; we arrange a clinical environment-lik e situation, where we could find more healthy patients than highly diseased ones to conduct anomaly detection. 3D MCGAN is a nov el GAN training method for DA, generating realistic but new nodules at desired posi- tion/size/attenuation, naturally blending with surrounding tissues (Fig. 2). W e crop/resize various nodules to 32 × 32 × 32 voxels and replace them with noise box es from a uniform distribution between [ − 0 . 5 , 0 . 5] , while maintain- ing their 64 × 64 × 64 surroundings as V olumes of Inter- est (V OIs)—using those noise boxes, instead of boxes filled with the same voxel values, improves the training robust- ness; then, we concatenate the V OIs with 6 size/attenuation conditions tiled to 64 × 64 × 64 voxels (e.g., if the size is small, each vox el of the small condition is filled with 1 , while the medium/lar ge condition voxels are filled with 0 to consider the effect of scaling factor). So, our generator uses the 64 × 64 × 64 × 7 inputs to generate desired nodules in the noise box regions. The 3D U-Net [5]-like generator adopts 4 con v olutional layers in encoders and 4 decon volu- tional layers in decoders respectiv ely with skip connections to effecti vely capture both nodule/context information. W e adopt two Pix2Pix GAN [17]-like discriminators with different loss functions: the context discriminator learns to classify real vs synthetic nodule/surrounding pairs with noise box-centered surroundings using Least Squares loss (LSGAN) [21]; the nodule discriminator attempts to classify real vs synthetic nodules with size/attenuation conditions using W asserstein loss with Gradient Penalty (WGAN-GP) [11]. The LSGAN in the context discrimina- tor forces the model to learn surrounding tissue background by reacting more sensitively to every pixel in images than regular GANs. The WGAN-GP in the nodule discriminator allows the model to generate realistic/di verse nodules with- out focusing too much on details. Empirically , we confirm that such multiple discriminators with the mutually com- plementary loss functions, along with size/attenuation con- ditioning, help generate realistic/div erse nodules naturally placed at desired positions on CT scans; similar results are also reported by this work [25] for 2D pedestrian detec- tion without label conditioning. W e apply dropout to in- ject randomness and balance the generator/discriminators. Batch normalization is applied to both conv olution (using LeakyReLU) and decon volution (using ReLU). Most GAN-based D A approaches use reconstruction ` 1 loss [9] to generate realistic images, even modifying it for further realism [18]. Howe v er , no one has ev er v alidated whether it really helps D A—it assures synthetic images re- sembling the original ones, sacrificing div ersity; thus, to confirm its influence during classifier training, we compare our MCGAN objectiv e without/with it, respecti vely: G ∗ = arg min G max D 1 ,D 2 L LSGAN ( G, D 1) + L WGAN-GP ( G, D 2) , (1) G ∗ = arg min G max D 1 ,D 2 L LSGAN ( G, D 1) + L WGAN-GP ( G, D 2) + 100 L ` 1 ( G ) . (2) W e set 100 as a weight for the ` 1 loss, since empirically it works well for reducing visual artifacts introduced by the GAN loss and most GAN works adopt the weight [17, 25]. 3D MCGAN Implementation Details T raining lasts for 6 , 000 , 000 steps with a batch size of 16 and 2 . 0 × 10 − 4 learning rate for the Adam optimizer . W e use horizon- tal/vertical flipping as D A and flip real/synthetic labels once in three times for robustness. During testing, we augment nodules with the same size/attenuation condi- tions by applying a random combination to real nodules of width/height/depth shift up to 10% and zooming up to 10% for better D A. As post-processing, we blend bounding boxes’ 3 nearest surfaces from all the boundaries by aver - aging the values of 6 nearest vox els/itself for 5 iterations. W e resample the resulting nodules to their original resolu- tion and map back onto the original CT scans to prepare additional training data. 3.2. Lung Nodule Detection Using 3D Faster RCNN 3D Faster RCNN is a 3D version of Faster RCNN [28] using multi-task loss with a 27 -layer Region Proposal Net- work of 3D conv olutional layers, batch normalization lay- ers, and ReLU layers. T o confirm the ef fect of MCGAN- based D A, we compare the following detection results trained on ( i ) 632 real images without GAN-based D A, ( ii ), ( iii ), ( iv ) with 1 × / 2 × / 3 × MCGAN-based D A (i.e., 632 / 1 , 264 / 1 , 896 additional synthetic training images) , ( v ), ( vi ), ( vii ) with 1 × / 2 × / 3 × MCGAN-based DA trained with ` 1 loss. During training, we shuf fle the real/synthetic image order . W e ev aluate the detection performance as follo ws: ( i ) Free Receiv er Operation Characteristic (FR OC) analysis, sensitivity as a function of FPs per scan; ( ii ) Competition Performance Metric (CPM) score [24], average sensitivity at sev en pre-defined FP rates: 1/8, 1/4, 1/2, 1, 2, 4, and 8 FPs per scan—this quantifies if a CAD system can identify a significant percentage of nodules with both very few FPs and moderate FPs. 3D F aster RCNN Implementation Details During train- ing, we use a batch size of 2 and 1 . 0 × 10 − 3 learning rate ( 1 . 0 × 10 − 4 after 20 , 000 steps) for the SGD optimizer with momentum. The input volume size to the network is set to 160 × 176 × 224 voxels. As classical D A, a random combi- nation of width/height/depth shift up to 15% and zooming up to 15% are also applied to both real/synthetic images to achie v e the best performance. For testing, we pick the model with the highest sensitivity on validation between 30 , 000 - 40 , 000 steps under Intersection ov er Union (IoU) threshold 0 . 25 /detection threshold 0 . 5 to a v oid sev ere FPs. 3.3. Clinical V alidation Using V isual T uring T est T o quantitativ ely ev aluate the realism of MCGAN- generated images, we supply , in a random order , to two expert physicians a random selection of 50 real and 50 synthetic lung nodule images with all of 2D ax- ial/coronal/sagittal views at the center . They take four classification tests in ascending order: T est1, 2: real vs MCGAN-generated 32 × 32 × 32 nodules, trained with- out/with ` 1 loss; T est3, 4: real vs MCGAN-generated 64 × 64 × 64 nodules with surroundings without/with ` 1 loss. Such V isual Turing T est [30] can ev aluate the visual quality of GAN-generated medical images in a clinical en- vironment, where physicians’ specialty is critical [13, 4]. 3.4. V isualization Using t-SNE T o visually analyze the distribution of real/synthetic im- ages, we use t-SNE [35] on a random selection of 500 real, 500 synthetic, and 500 ` 1 loss-added synthetic nodule im- Lung CT (Real nodule w/ surr oundings, 64 × 64 × 64) Lung CT (Noise bo x-replaced nodule w/ sur roundings, 64 × 64 × 64) Lung CT (Synthetic nodule w/ sur roundings, 64 × 64 × 64) Lung CT (L1 loss-added synthetic nodule w/ sur roundings, 64 × 64 × 64) Figure 3: 2D axial view of e xample 64 × 64 × 64 lung nodules with surrounding tissues; 3D MCGANs generate only 32 × 32 × 32 nodules. T able 1: Nodule detection results (CPM) with/without 3D MCGAN-based DA for 3D Faster RCNN under IoU ≥ 0 . 25 . Both results without/with ` 1 loss at dif ferent augmentation ratio are compared. CPM is av erage sensiti vity at 1 / 8 , 1 / 4 , 1 / 2 , 1 , 2 , 4 , and 8 FPs per scan. CPM by Size CPM by Attenuation CPM Small Medium Large Solid Part-solid GGN 632 real images 0.518 0.447 0.618 0.624 0.655 0.464 0.242 + 1 × 3D MCGAN-based D A 0.550 0.452 0.683 0.662 0.699 0.521 0.244 + 2 × 3D MCGAN-based D A 0.527 0.447 0.674 0.429 0.655 0.407 0.289 + 3 × 3D MCGAN-based D A 0.512 0.411 0.644 0.662 0.616 0.579 0.277 + 1 × 3D MCGAN-based D A w/ ` 1 0.508 0.430 0.633 0.556 0.626 0.471 0.271 + 2 × 3D MCGAN-based D A w/ ` 1 0.509 0.406 0.644 0.654 0.649 0.436 0.233 + 3 × 3D MCGAN-based D A w/ ` 1 0.479 0.389 0.594 0.617 0.596 0.507 0.226 ages, with a perplexity of 100 for 1 , 000 iterations to get a 2D representation. W e normalize the input images to [0 , 1] . The t-SNE method represents high-dimensional data into a lower -dimensional space by reducing the dimensionality; it uses perplexity to non-linearly balance between the input data’ s local and global aspects. 4. Results 4.1. Lung Nodules Generated by 3D MCGAN W e generate realistic nodules in noise box regions at v ar - ious position/size/attenuation, naturally blending with sur- rounding tissues including v essels, soft tissues, and thoracic walls (Fig. 3). Especially , when trained without ` 1 loss, those synthetic nodules look much more different from the original real ones, including slight shading difference. 4.2. Lung Nodule Detection Results T able 1 and Fig. 4 sho w that it is easier to detect nodules with larger size and lower attenuation due to their clear ap- pearance. 3D MCGAN-based D A with less augmentation ratio consistently increases sensitivity at fixed FP rates— especially , training with 1 × MCGAN-based D A without ` 1 loss outperforms training only with real images under any Figure 4: FROC curv es of dif ferent D A setups by a verage/size/attenuation. Ground Truth w/o GAN + 1× GAN + 2× GAN + 3× GAN + 1× GAN w/ L1 + 2× GAN w/ L1 + 3× GAN w/ L1 (a) (b) (c) (d) (e) (f) (g) (h) Figure 5: Example detection results with detection threshold 0 . 5 : (a) ground truth; (b) without GAN-based D A; (c), (d), (e) with 1 × / 2 × / 3 × 3D MCGAN-based DA; (f), (g), (h) with 1 × / 2 × / 3 × ` 1 loss-added 3D MCGAN-based D A. size/attenuation in terms of CPM, achieving average CPM improv ement by 0.032. It especially boosts nodule detec- tion performance with larger size and lower attenuation. Fig. 5 visually re veals its ability to alleviate the risk of ov er - looking the nodule diagnosis with clinically acceptable FPs (i.e., the highly-ov erlapping bounding boxes around nod- ules only require a physician’ s single check by switching on/off transparent alpha-blended annotation on CT scans). Surprisingly , adding more synthetic images tends to de- crease sensitivity , probably due to the real/synthetic train- ing image balance. Moreover , further nodule realism in- troduced by ` 1 loss rather decreases sensitivity as ` 1 loss sacrifices div ersity in return for the realism. 4.3. V isual T uring T est Results As T able 2 shows, expert physicians fail to classify real vs MCGAN-generated nodules without surrounding tissues—ev en regarding the synthetic nodules trained with- out ` 1 loss more realistic than the real ones. Contrarily , they relativ ely recognize the synthetic nodules with sur- roundings due to slight shading dif ference between the nod- ules/surroundings, especially when trained without the re- construction ` 1 loss. Considering the synthetic images’ re- T able 2: V isual T uring T est results by two physicians for classifying 50 real vs 50 3D MCGAN-generated images: T est1, 2: 32 × 32 × 32 lung nodules, trained without/with ` 1 loss; T est3, 4: 64 × 64 × 64 nodules with surrounding tissues, trained without/with ` 1 loss. Accuracy denotes the physicians’ successful classification ratio between the real/synthetic images. Accuracy Real Selected as Real Real as Synt Synt as Real Synt as Synt T est1 Physician1 43% 19 31 26 24 Physician2 43% 13 37 20 30 T est2 Physician1 57% 22 28 15 35 Physician2 53% 11 39 8 42 T est3 Physician1 62% 25 25 13 37 Physician2 79% 32 18 3 47 T est4 Physician1 58% 21 29 13 37 Physician2 66% 36 14 20 30 Figure 6: T -SNE plot with 500 32 × 32 × 32 lung nodule images per each category: (a), (b) 3D MCGAN-generated nodules, trained without/with ` 1 loss; (c) real nodules. alism, CPGGANs might perform as a tool to train medi- cal students and radiology trainees when enough medical images are unav ailable, such as abnormalities at rare posi- tion/size/attenuation. Such GAN applications are clinically promising [7]. 4.4. T -SNE Results Implying their ef fective D A performance, synthetic nod- ules have a similar distribution to real ones, but concen- trated in left inner areas with less real ones especially when trained without ` 1 loss (Fig. 6)–using only GAN loss during training can avoid overwhelming influence from the real im- age samples, resulting in a moderately similar distribution; thus, those synthetic images can partially fill the real image distribution unco vered by the original dataset. 5. Conclusion Our bounding box-based 3D MCGAN can gen- erate div erse CT -realistic nodules at desired posi- tion/size/attenuation, naturally blending with surrounding tissues—those synthetic training data boost sensiti vity un- der any size/attenuation at fixed FP rates in 3D CNN-based nodule detection. This attributes to the MCGAN’ s good generalization ability coming from multiple discriminators with mutually complementary loss functions, along with informativ e size/attenuation conditioning; they allow to cov er real image distribution unfilled by the original dataset, improving the training rob ustness. Surprisingly , we find that adding over -suf ficient synthetic images produces worse results due to the real/synthetic image balance; as t-SNE results show , the synthetic images only partially cover the real image distribution, and thus GAN-ov erwhelming training images rather harm training. Moreov er , we notice that GAN training without ` 1 loss obtains better D A performance thanks to increased div ersity providing robustness; also expert physicians confirm their sufficient realism without ` 1 loss. Overall, our 3D MCGAN could help minimize expert physicians’ time-consuming annotation tasks and ov ercome the general medical data paucity , not limited to lung CT nodules. As future work, we will inv estigate the MCGAN- based DA results without size/attenuation conditioning to confirm their influence on DA performance. Moreov er , we will compare our D A results with other non-GAN-based re- cent D A approaches, such as mixup [38] and cutout [6]. For further performance boost, we plan to directly opti- mize the detection results for MCGANs, instead of re- alism, similarly to the three-player GAN for classifica- tion [36]. Lastly , we will inv estigate how our MCGAN can perform as a physician training tool to display random re- alistic medical images with desired abnormalities (i.e., po- sition/size/attenuation conditions) to help train medical stu- dents and radiology trainees despite infrastructural and legal constraints [7]. References [1] A. Antoniou, A. Storkey , and H. Edwards. Data aug- mentation generative adversarial networks. arXiv preprint arXiv:1711.04340 , 2017. [2] S. G. Armato III, G. McLennan, L. Bidaut, M. F . McNitt- Gray , C. R. Meyer , et al. The lung image database consor- tium (LIDC) and image database resource initiati ve (IDRI): a completed reference database of lung nodules on CT scans. Medical physics , 38(2):915–931, 2011. [3] O. Bailo, D. Ham, and Y . M. Shin. Red blood cell image generation for data augmentation using conditional genera- tiv e adversarial netw orks. arXiv preprint , 2019. [4] M. J. Chuquicusma, S. Hussein, J. Burt, and U. Bagci. How to fool radiologists with generativ e adversarial networks? a visual T uring test for lung cancer diagnosis. In Pr oc. IEEE International Symposium on Biomedical Imaging (ISBI 2018) , pages 240–244, 2018. [5] ¨ O. C ¸ ic ¸ ek, A. Abdulkadir , S. S. Lienkamp, T . Brox, and O. Ronneberger . 3D U-Net: learning dense volumetric se g- mentation from sparse annotation. In Proc. International Confer ence on Medical Image Computing and Computer- Assisted Intervention (MICCAI) , pages 424–432. Springer, 2016. [6] T . DeVries and G. W . T aylor . Improved regularization of con v olutional neural networks with cutout. arXiv preprint arXiv:1708.04552 , 2017. [7] S. G. Finlayson, H. Lee, I. S. Kohane, and L. Oakden- Rayner . T ow ards generativ e adversarial networks as a new paradigm for radiology education. In Pr oc. Machine Learn- ing for Health (ML4H) W orkshop arXiv:1812.01547 , 2018. [8] M. Frid-Adar , I. Diamant, E. Klang, M. Amitai, J. Gold- berger , and H. Greenspan. GAN-based synthetic medical image augmentation for increased CNN performance in li ver lesion classification. Neur ocomputing , 321:321–331, 2018. [9] C. Gao, S. Clark, J. Furst, and D. Raicu. Augmenting LIDC dataset using 3D generati ve adv ersarial networks to impro ve lung nodule detection. In Medical Imaging 2019: Computer- Aided Diagnosis , v olume 10950, page 109501K, 2019. [10] I. Goodfellow , J. Pouget-Abadie, M. Mirza, B. Xu, D. W arde-Farley , et al. Generati ve adversarial nets. In Ad- vances in Neural Information Pr ocessing Systems (NIPS) , pages 2672–2680, 2014. [11] I. Gulrajani, F . Ahmed, M. Arjo vsky , V . Dumoulin, and A. C. Courville. Improv ed training of W asserstein GANs. arXiv pr eprint arXiv:1704.00028 , 2017. [12] V . Gulshan, L. Peng, M. Coram, M. C. Stumpe, D. W u, et al. Dev elopment and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus pho- tographs. J AMA , 316(22):2402–2410, 2016. [13] C. Han, H. Hayashi, L. Rundo, R. Araki, W . Shimoda, et al. GAN-based synthetic brain MR image generation. In IEEE International Symposium on Biomedical Imaging (ISBI) , pages 734–738, 2018. [14] C. Han, K. Murao, T . Noguchi, Y . Kaw ata, F . Uchiyama, et al. Learning more with less: Conditional PGGAN- based data augmentation for brain metastases detection using highly-rough annotation on MR images. In Proc. A CM In- ternational Confer ence on Information and Knowledge Man- agement (CIKM) , 2019. (In press). [15] C. Han, L. Rundo, R. Araki, Y . Furukawa, G. Mauri, H. Nakayama, and H. Hayashi. Infinite brain MR images: PGGAN-based data augmentation for tumor detection. In Neural Approac hes to Dynamics of Signal Exchanges , vol- ume 151 of Smart Innovation, Systems and T echnologies . Springer , 2018. (In press). [16] C. Han, L. Rundo, R. Araki, Y . Nagano, Y . Furukawa, et al. Combining noise-to-image and image-to-image GANs: Brain MR image augmentation for tumor detection. arXiv pr eprint arXiv:1905.13456 , 2019. [17] P . Isola, J.-Y . Zhu, T . Zhou, and A. A. Efros. Image-to-image translation with conditional adversarial networks. In Pr oc. IEEE Conference on Computer V ision and P attern Recogni- tion (CVPR) , pages 1125–1134, 2017. [18] D. Jin, Z. Xu, Y . T ang, A. P . Harrison, and D. J. Mollura. CT- realistic lung nodule simulation from 3D conditional gen- erativ e adversarial networks for robust lung segmentation. In Pr oc. International Conference on Medical Image Com- puting and Computer-Assisted Intervention (MICCAI) , pages 732–740, 2018. [19] D. P . Kingma and M. W elling. Auto-encoding variational bayes. In Pr oc. International Confer ence on Learning Rep- r esentations (ICLR) arXiv:1312.6114 , 2013. [20] C. Ledig, L. Theis, F . Husz ´ ar , J. Caballero, A. Cunningham, et al. Photo-realistic single image super-resolution using a generativ e adversarial network. In Pr oc. IEEE confer ence on Computer V ision and P attern Recognition (CVPR) , pages 4681–4690, 2017. [21] X. Mao, Q. Li, H. Xie, R. Y . Lau, Z. W ang, and S. Paul Smol- ley . Least squares generative adversarial networks. In Proc. IEEE International Confer ence on Computer V ision (ICCV) , pages 2794–2802, 2017. [22] L. Mescheder , S. No wozin, and A. Geiger . Adv ersarial vari- ational bayes: unifying v ariational autoencoders and genera- tiv e adv ersarial networks. In Pr oc. International Conference on Machine Learning (ICML) , pages 2391–2400, 2017. [23] F . Milletari, N. Navab, and S.-A. Ahmadi. V -Net: Fully con v olutional neural networks for volumetric medical image segmentation. In Pr oc. IEEE International Conference on 3D V ision (3D V) , pages 565–571, 2016. [24] M. Niemeijer , M. Loog, M. D. Abramoff, M. A. V ierge v er, M. Prokop, and B. van Ginneken. On combining computer - aided detection systems. IEEE Tr ansactions on Medical Imaging , 30(2):215–223, 2011. [25] X. Ouyang, Y . Cheng, Y . Jiang, C.-L. Li, and P . Zhou. Pedestrian-Synthesis-GAN: Generating pedestrian data in real scene and beyond. arXiv pr eprint arXiv:1804.02047 , 2018. [26] H. P ark, Y . Y oo, and N. Kw ak. MC-GAN: Multi- conditional generativ e adversarial network for image synthe- sis. In Pr oc. British Machine V ision Conference (BMVC) arXiv:1805.01123 , 2018. [27] A. Radford, L. Metz, and S. Chintala. Unsupervised repre- sentation learning with deep con v olutional generati ve adv er- sarial networks. In Pr oc. International Conference on Learn- ing Repr esentations (ICLR) arXiv:1511.06434 , 2016. [28] S. Ren, K. He, R. Girshick, and J. Sun. Faster R-CNN: T o- wards real-time object detection with region proposal net- works. In Advances in Neural Information Processing Sys- tems (NIPS) , pages 91–99, 2015. [29] O. Ronneberger , P . Fischer, and T . Brox. U-Net: Con volu- tional networks for biomedical image segmentation. In Pr oc. International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI) , pages 234–241, 2015. [30] T . Salimans, I. Goodfello w , W . Zaremba, V . Cheung, A. Rad- ford, and X. Chen. Improved techniques for training GANs. In Advances in Neural Information Pr ocessing Sys- tems (NIPS) , pages 2234–2242, 2016. [31] A. A. A. Setio, A. Tra verso, T . De Bel, M. S. Berens, C. v an den Bogaard, et al. V alidation, comparison, and com- bination of algorithms for automatic detection of pulmonary nodules in computed tomography images: the LUNA16 challenge. Medical image analysis , 42:1–13, 2017. [32] H.-C. Shin, N. A. T enenholtz, J. K. Rogers, C. G. Schwarz, M. L. Senjem, et al. Medical image synthesis for data aug- mentation and anonymization using generative adversarial networks. In International W orkshop on Simulation and Syn- thesis in Medical Imaging (SASHIMI) , pages 1–11, 2018. [33] A. Shriv asta va, T . Pfister , O. T uzel, J. Susskind, W . W ang, and R. W ebb. Learning from simulated and unsupervised images through adv ersarial training. In Pr oc. IEEE Confer- ence on Computer V ision and P attern Recognition (CVPR) , pages 2107–2116, 2017. [34] R. L. Siegel, K. D. Miller , and A. Jemal. Cancer statistics, 2019. CA: a cancer journal for clinicians , 69(1):7–34, 2019. [35] L. van der Maaten and G. Hinton. V isualizing data using t-SNE. J. Mac h. Learn. Res. , 9:2579–2605, 2008. [36] S. V andenhende, B. De Brabandere, D. Nev en, and L. V an Gool. A three-player GAN: generating hard sam- ples to improve classification networks. In Pr oc. Interna- tional Conference on Machine V ision Applications (MV A) arXiv:1903.03496 , 2019. [37] J. W u, C. Zhang, T . Xue, B. Freeman, and J. T enenbaum. Learning a probabilistic latent space of object shapes via 3D generativ e-adversarial modeling. In Advances in neural in- formation pr ocessing systems , pages 82–90, 2016. [38] H. Zhang, M. Cisse, Y . N. Dauphin, and D. Lopez-Paz. mixup: Beyond empirical risk minimization. arXiv preprint arXiv:1710.09412 , 2017. [39] Y . Zhu, M. Aoun, M. Krijn, J. V anschoren, and H. T . Cam- pus. Data augmentation using conditional generative ad- versarial networks for leaf counting in arabidopsis plants. In Pr oc. Computer V ision Pr oblems in Plant Phenotyping (CVPPP) , 2018.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment