Projection-Based 2.5D U-net Architecture for Fast Volumetric Segmentation
Convolutional neural networks are state-of-the-art for various segmentation tasks. While for 2D images these networks are also computationally efficient, 3D convolutions have huge storage requirements and require long training time. To overcome this …
Authors: Christoph Angermann, Markus Haltmeier, Ruth Steiger
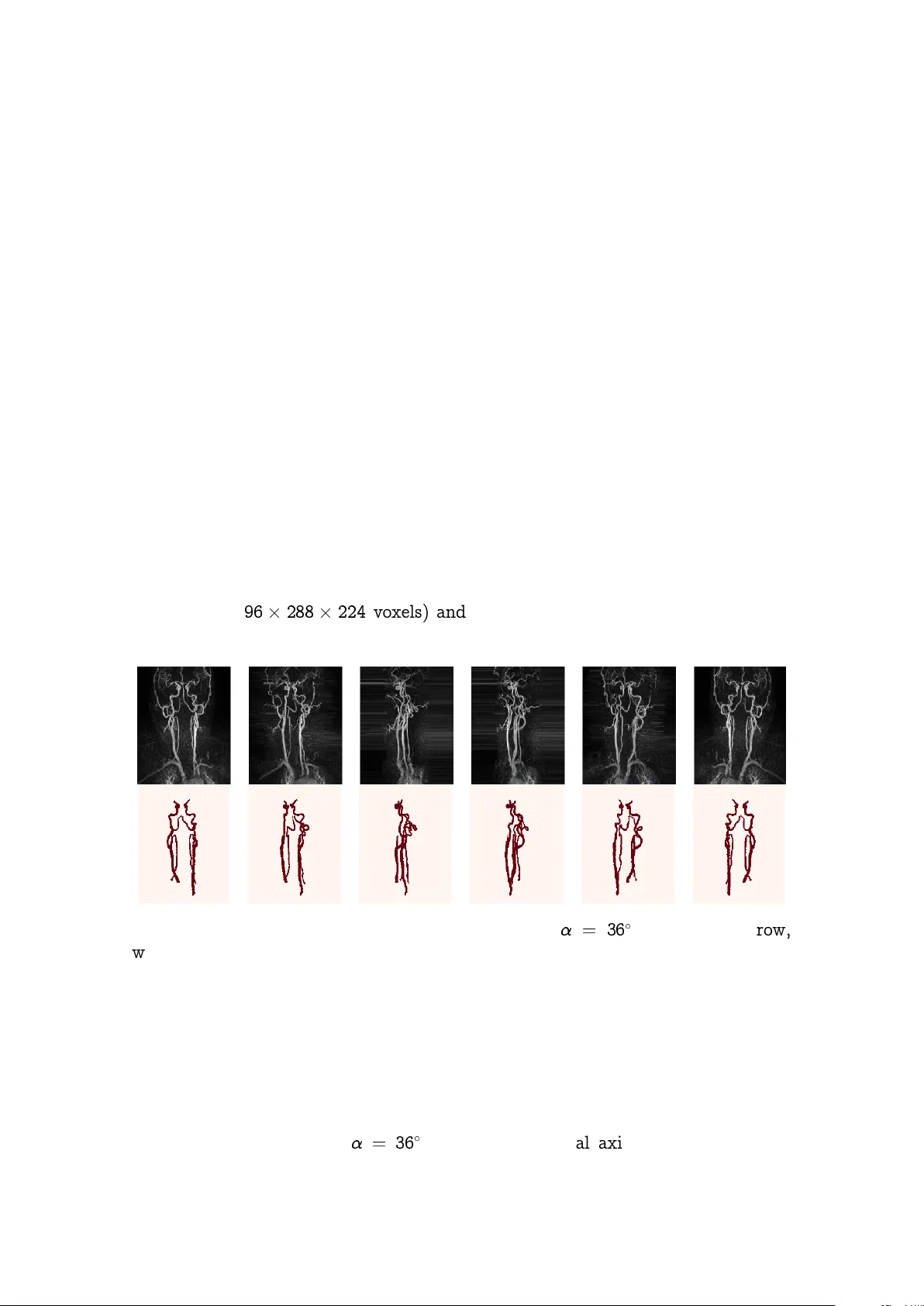
Pro jection-Based 2.5D U-net Arc hitecture for F ast V olumetric Segmen tation Christoph Angermann and Markus Haltmeier Departmen t of Mathematics Univ ersity of Innsbruc k T ec hnikerstrasse 13, 6020 Innsbruc k, Austria {christoph.angermann,markus.haltmeier}@uibk.ac.at Ruth Steiger, Sergiy P ereverzy ev Jr., and Elk e Gizewski Univ ersitätsklinik für Neuroradiologie Medizinisc he Universität Innsbruc k Anic hststraße 35, 6020 Innsbruck neuroradiologie@i-med.ac.at August 5, 2019 Abstract Con v olutional neural net works are state-of-the-art for v arious segmen ta- tion tasks. While for 2D images these net w orks are also computationally efficien t, 3D conv olutions hav e huge storage requirements and require long training time. T o o v ercome this issue, we in tro duce a net w ork structure for v olumetric data without 3D con volutional la yers. The main idea is to include maxim um in tensity pro jections from differen t directions to transform the vol- umetric data to a sequence of images, where each image contains information of the full data. W e then apply 2D con volutions to these pro jection images and lift them again to volumetric data using a trainable reconstruction algo- rithm. The prop osed netw ork arc hitecture has less storage requirements than net w ork structures using 3D conv olutions. F or a tested binary segmentation task, it even shows b etter p erformance than the 3D U-net and can b e trained m uc h faster. 1 1 In tro duction Deep con volutional neural net works ha v e b ecome a p o werful method for image recognition ([1, 2]) . In the last few y ears they also exceeded the state-of-the-art in providing segmentation masks for images. In [3], the idea of transforming V GG- nets [2] to deep con v olutional filters to obtain seman tic segmen tations of 2D images came up. Based on these deep con v olutional filters, the authors of [4] introduced a nov el net work architecture, the so-called U-net. With this arc hitecture they redefined the state-of-the-art in 2D image segmen tation till today . The U-net pro vides a p o w erful 2D segmentation to ol for biomedical applications, since it has b een demonstrated to learn highly accurate ground truth masks from only very few training samples. Among others, the fully automated generation of v olumetric segmentation masks b ecomes increasingly important for biomedical applications. This task still is c hallenging. One idea is to extend the U-net structure to v olumetric data b y using 3D conv olutions, as has b een prop osed in [5, 6]. Essen tial dra wbac ks are the h uge memory requiremen ts and long training time. Deep learning segmen tation metho ds therefore are often applied to 2D slice images (compare [5]). Ho w ev er, these slice images do not contain information of the full 3D data whic h makes the segmen tation task m uch more challenging. T o address the dra wbac ks of existing approaches, we in tro duce a netw ork structure whic h is able to generate accurate volumetric segmen tation masks of v ery large 3D volumes. The main idea is to integrate maxim um intensit y pro jection (MIP) la yers from differen t directions which transform the data to 2D images con taining information of the full 3D image. As an example, we test the netw ork for segmen t- ing blo o d v essels (arteries and veins) in magnetic resonance angiography (MRA) scans (Figure 1.1). (a) T ransversal. (b) Sagittal. (c) Coronal. (d) 3D segmen tation. Figure 1.1: In every plane (a)-(c) the blo o d vessels of in terest are mark ed in red. In (d) we see the corresp onding 3D segmentation mask. The segmentation was conducted with the freeware ITK-SNAP [7]. The prop osed netw ork can b e trained 15 faster and requires order of magnitude less memory than net w orks with 3D con v olutions, and still pro duces more accurate results. 2 2 Bac kground 2.1 V olumetric segmen tation of blo o d v essels W e aim at generating v olumetric binary segmen tation masks. In particular, as one targeted application, w e aim at segmenting blo o d vessels (arteries and veins) whic h assists the do ctor to detect abnormalities like stenosis or aneurysms. F ur- thermore, the medical sector is lo oking for a fully automated metho d to ev aluate large cohorts in the future. The Departmen t of Neuroradiology Innsbruc k has pro vided volumetric MRA scans of 119 different patients. The images face the arteries and veins b et ween the brain and the chest. F ortunately , also the volumet- ric segmentation masks (ground truths) of these 119 patien ts hav e b een provided. These segmen tation masks hav e b een generated b y hand which is long hard work (Figure 1.1). Our goal is the fully automated generation of the 3D segmen tation masks of the blo o d vessels. F or that purp ose we use deep learning and neural net works. At the first glance, this problem ma y seem to b e quite easy because w e only hav e t wo lab els (0: bac kground, 1: bloo d vessel). Ho w ev er, there are also arteries and v eins whic h ha v e lab el 0 whic h migh t confuse the netw ork since w e only w ant to segmen t those vessels of interest. Other challenges are caused b y the big size of the volumes ( 96 288 224 v oxels) and by the very un balanced distribution of the tw o lab els (in av erage, 99.76 % of all vo xels indicate background). Figure 2.1: MIP images of a 3D MRA scan with = 36 . In the first ro w, w e see the pro jections of the original scan, in the second ro w the corresp onding pro jections of the ground truth. 2.2 Segmen tation of MIP images W e first solv e a 2D v ersion of our problem. This can b e done b y applying maximum in tensity pro jections to the 3D data and the corresp onding 3D ground truths. Using a rotation angle of = 36 around the vertical axis w e obtain 10 MIP 3 images out of each patient, which results in a data set to 1190 pairs of 2D images and corresp onding 2D segmen tation masks. Data corresp onding for one patient are shown in Figure 2.1. The U-net used for binary segmentation is a mapping U : R a b ! [0 ; 1] a b whic h tak es an image as input and outputs for eac h pixel the probabilit y of b eing a foreground pixel. It is formed b y the following ingredien ts [4]: The c ontr acting p art : It includes stacking ov er conv olutional blo c ks (con- sisting of 2 con volutional la yers) and max-p o oling la y ers considering follow- ing prop erties: (1) W e only use 3 3 filters to hold down complexity and zero-padding to guaran tee that all la yer outputs hav e ev en spatial dimen- sion. (2) Eac h max-p o oling la y er has stride (2 ; 2) to half the spatial sizes. W e m ust b e a ware that the spatial dimensions of the input can get divided b y 2 often enough without pro ducing any rest. This can b e done b y slight cropping. (3) After eac h p o oling lay er we use t wice as man y filters as in the previous conv olutional blo ck. The upsampling p art : T o obtain similarit y to the con tracting part, we mak e use of transp osed con v olutions to double spatial dimension and to halve the n umber of filters. They are follo w ed b y con volutional blo c ks consisting of t wo con volutional la yers with k ernel size 3 3 after each upsampling lay er (compare Figure 2.2). Figure 2.2: Visualization of the ground architecture of a 2D U-net. Ev ery con volutional la yer in this structure gets follo wed b y a ReLu-activ ation- function. T o link the con tracting and the upsampling part, concatenation la y- ers are used, where t wo images with same spatial dimension get concatenated 4 o ver their c hannel dimension (see Figure 2.2). This ensures a com bination of eac h pixel’s information with its lo calization. At the end, the sigmoid-activ ation- function is applied, whic h outputs for eac h pixel the probabilit y for b eing a fore- ground pixel. T o get the final segmen tation mask, a threshold (usually 0.5) is applied p oint-wise to the output of the U-net. All netw orks in this pap er are build with the Ker as library [8] using T ensorflow bac kend [9]. Our implemented 2D U-net has filter size 32 at the b eginning and filter size 512 at the end of the con tracting part. The v alues of the start weigh ts are normally distributed with ex pectation 0 and deviation 1 f l , where f l denotes the size of the l -th conv olutional lay er. The netw ork is trained with the Dice-loss function [6] ` ( y ; ^ y ) = 1 2 P k ( y ^ y ) k P k ^ y k + P k y k ; where denotes pixelwise m ultiplication, the sums are taken ov er all pixel lo- cations, ^ y = ^ f ( x ) are the probabilities predicted b y the U-net, and y is the re- lated ground truth. The Dice-loss function measures similarit y by comparing all correctly predicted v essels pixels with the total num b er of v essels pixels in the prediction. F or i; j 2 f 0 ; 1 g let us denote b y p ij the set of all pixels of class i predicted to class j , and by t i the n um b er of all pixels b elonging to class i . With this notation, we ev aluate the following metrics during training: Me an A c cur acy : MA , 1 2 p 00 t 0 + p 11 t 1 ; Me an Interse ction over Union [10]: IU , 1 2 p 00 t 0 + p 10 + p 11 t 1 + p 01 ! ; Dic e-c o efficient ([6, 11]): DC , 2 p 11 2 p 11 + p 01 + p 10 : T o guaran tee that all samples ha ve satisfying spatial dimensions, the images get symmetrically cropp ed a little bit. W e also mak e use of batc h normalization la y ers [12] b efore each conv olutional blo c k to sp eed up con vergence. T o handle o verfit- ting [13], w e also recommend the in tegration of drop out lay ers [14] with drop out rate 0.5 in the deep est con v olutional blo ck and dropout rate 0.2 in the second deep est blo cks. F or training, A dam-optimizer [15] is used with learning rate 0.001 in combination with learning-rate-sc heduling, i.e. if the v alidation loss do es not decrease within 3 ep o c hs the learning rate gets reduced b y the factor 0.5. F urther- more, if the netw ork shows no improv emen t for 5 ep o c hs the training pro cess gets 5 stopp ed (early stopping) and the w eigh ts of the b est ep o c h in terms of v alidation loss get restored. W e use a (70, 15, 15) split in training, v alidation, and ev alua- tion data and a threshold of 0.5 for the construction of the segmen tation masks. T raining the U-net on NVIDIA GeF or c e R TX 2080 GPU with a minibatch size of 6 yields the follo wing results: Dice-loss of 0.088, mean accuracy of 95.7%, mean IU of 91.6%, and Dice-co efficient of 91.3%. In av erage, training the 2D U-net lasts 809 seconds, the application only 0.013 seconds. During training, the 2D U-net allo cates a memory space of 1.7 gigabytes. 2.3 Segmen tation with the 3D U-net No w w e aim at generating binary segmen tation masks of sparse volumetric data using a 3D version of the prior in tro duced U-net. The resulting 3D U-net follo ws the same structure as in 2.2, the only difference is the usage of 3D con volutions and 3D p o oling la y ers. F or the 3D U-net we hav e to take sp ecial care ab out o v erfitting [13] and ab out memory space. Therefore, for the 3D U-net we ha ve chosen filter size 4 at the b eginning and filter size 16 at the end of the contracting part. Also the use of high drop out rates [14] (0.5 in the deep est conv olutional blo ck and 0.4 in the second deep est blo c ks) is necessary to ensure an efficien t training pro cess. Due to the huge size of our training samples ( 96 288 224 v oxels), we train the netw ork on batc h, i.e. with minibatc h size 1. During training the 3D U-net allocates more than 8 gigabyte memory space and therefore is not manageable an y more by our GPU. Therefore, the training pro cess on a AMD Ryzen 7 1700X eigh t-core pro cessor takes in av erage 969 min utes. Since the n umber of 3D samples is only 119, w e conducted 5 training-runs with random c hoice of training, v alidation and ev aluation data. Using the 3D U-net w e obtained in a verage follo wing results: Dice-loss of 0.254, mean accuracy of 87.3%, mean IU of 80.5% and Dice-co efficien t of 74.8%. Although the 3D U-net demonstrates high precision in our application (see Figures 3.4,3.5), w e are not satisfied with the long training time. In addition to it, we are v ery limited in the choice of conv olutional la y ers and the corresp onding n umber of filters due to the h uge size of the input data. So it is hardly p ossible to conduct v olumetric segmentation for even larger biomedical scans without using cropping or sliding-window tec hniques. 3 Pro jection-Based 2.5D U-net As men tioned in the in tro duction, the naiv e approach for accelerating volumet- ric segmentation and reducing memory requiremen ts is to pro cess each of the 96 slice images indep endently through a 2D-net work (compare [5]). How ev er, this causes the loss of connection b et ween the slice images. F or our targeted applica- tion, applying the 2D U-net out of 2.2 to eac h slice image of the 3D MRA scans 6 indep enden tly yields following disapp oin ting results: Dice-loss of 0.849, mean ac- curacy of 54.5%, mean IU of 54.3%, and Dice-co efficient of 15.1%. Therefore we are lo oking for an alternativ e approach. Figure 3.1: Reconstruction operator R 2 : V oxel v alue is defined as the sum o ver the corresp onding 2D v alues, here illustrated for 2 MIP images with directions f 0 ; 90 g . 3.1 Prop osed 2.5D U-net architecture As w e hav e seen in 2.2, the 2D U-net do es v ery well on the MIP images. Recall that a netw ork for binary volumetric segmen tation is a function N : R a b c ! [0 ; 1] a b c that maps the 3D scan to the probabilities that a vo xel corresp onds to the desired class. F or a 3D input x , the prop osed 2.5D U-net takes the form N ( x ) = T R p F p 2 6 6 4 U M 1 ( x ) . . . U M p ( x ) 3 7 7 5 ; (3.1) where M i : R a b c ! R b c are MIP images for different pro jection directions 1 ; : : : ; p , U : R b c ! [0 ; 1] b c is the same 2D U-net as in 2.2 pro ducing probabilities, F p : ([0 ; 1] b c ) p ! ( R b c ) p is a learnable filtration, 7 R p : ( R b c ) p ! R a b c is a reconstruction op erator using p line ar b ackpr o- je c tions as sho wn in Figure 3.1, T : R a b c ! [0 ; 1] a b c is a fine-tuning op erator (av erage p o oling follow ed b y a learnable shift -op erator follo w ed by the sigmoid-activ ation-function). The bac kpro jection op erator R p causes a kind of shroud (Figure 3.2a), so we hav e to think ab out a filtrated backpro jection. Therefore, w e apply a conv olutional la yer F p b efore bac kpro jection. Using 1 3 filters, whic h get adapted during training for each pro jection direction 1 ; : : : ; p individually , leads to a more sat- isfying result (Figure 3.2b). Figure 3.2a: Netw ork’s output (b efore threshold) without filtration. Figure 3.2b: Netw ork’s output (before threshold) with filtration. F or the fine-tuning operator T w e use a verage p o oling with po ol-size (2, 2, 2). This is follow ed b y a learnable shift-op erator, which shifts the p o oled data by an adjusted parameter since the decision b oundaries ha ve b een c hanged b y R p . This ensures, that the application of the sigmoid function deliv ers accurate probabili- ties. F or our targeted application, again we only pro cess one 3D sample through the net work p er iteration (minibatch size 1). The start w eights of the con v olutional part U in 3.1 are initialized in the same w ay as in Section2.2. The parameters of F p and T are initialized empirically . F or the amount of the pro jection directions we c ho ose equidistan t angles = f k 180 p j k = 0 ; : : : ; p 1 g to ensure w e obtain at most different information of the 3D data for different pro jection directions. This causes the task of finding the b est v alue for p in 3.1. Therefore w e trained the prop osed netw ork N for differen t v alues for p and compared p erformance in terms of the ev aluation metrics (Figure 3.3). Lo oking at Figure 3.3, we observ e that = f k 180 12 j k = 0 ; : : : ; 11 g seems to b e a go o d choice for the amount of pro jection directions. W e hav e conducted 5 8 Figure 3.3: P erformance of the 2.5D U-net for differen t num b er p of pro jection directions . training runs with random c hoice of training, v alidation and ev aluation data and obtained in av erage follo wing results: Dice-loss of 0.201, mean accuracy of 91.6 %, mean IU of 86.1 % and Dice-co efficient of 83.7 %. Figure 3.4: Comparison b et ween ground truth (first ro w), segmentation generated b y 3D U-net (second row) and segmentation generated by 2.5D U-net (third row). As we can see, for the volumetric segmentation of MRA scans the prop osed 2.5D U-net clearly outp erforms 3D U-net in terms of ev aluation metrics. F urthermore, 9 Figure 3.5: 3D segmentation mask generated by hand (left), by 3D U-net (middle) and by 2.5D U-net (righ t). adjusting the w eigh ts of 2.5D U-net only tak es in av erage 3914 seconds. With 11.37 seconds, the application time increased due to the construction of the MIP images. During training, the 2.5D U-net allo cates a memory space of 3.7 gigab ytes. F urther tasks w ould b e to inv estigate if applying data augmen tation techniques to the 3D samples inc reases accuracy of the 3D U-net. Considering our application, data deformation could cause problems due to the fact, that the orien tation of the v essels has huge impact to the netw ork’s prediction. W e will inv estigate that in the future. T able 1: Summarization of the ev aluation results for the naive 2D U-net slice-p er- slice approach, the 3D U-net and the prop osed 2.5D U-net. Net work loss MA in % IU in % DC in % 2D U-net 0.849 54.5 54.3 15.1 3D U-net 0.254 87.3 80.5 74.8 2.5D U-net 0.201 91.6 86.1 83.7 T able 2: Summarization of ti me and storage observ ations for the naiv e 2D U-net slice-p er-slice approach, the 3D U-net and the prop osed 2.5D U-net.. Net work W eigh ts T rain Appl. Mem. 2D U-net 8 : 6 10 6 809 sec. 1.83 sec. 1.7 Gb 3D U-net 2 : 5 10 4 58140 sec. 5.28 sec. > 8 Gb 2.5D U-net 8 : 6 10 6 3914 sec. 11.37 sec. 3.7 Gb 4 Conclusion In this pap er w e prop osed a new pro jection-based 2.5D U-net structure for fast v ol- umetric segmentation. The construction of v olumetric segmentation masks with the help of a 3D U-net deliv ers very satisfying results, but the long training time 10 and the big need of memory space are hardly sustainable. The 2.5D U-net using 12 deterministic pro jection directions is able to conduct 3D segmentation of very big biomedical 3D scans as reliable as the 3D U-net and can b e trained muc h faster without any conc ern ab out memory space. F or our targeted application, the 2.5D U-net enables the generation of 3D segmentations in a storage efficient wa y more accurate than other approaches using 3D con v olutions and can b e trained almost 15 faster. All n umerical results considering the ev aluation metrics are display ed in T able 1. A v erage training time, application time and storage requiremen ts for eac h netw ork are summarized in T able 2. In the curren t implemen tation, w e only use MIP images for deterministic pro jection directions. In future work, we will in vestigate the use of random pro jection directions for net w ork training. This could pro vide the p ossibility to use all a v ailable information from each pro jection direction for the construction of 3D segmen tation masks. Also the conduction of comparativ e studies will b e a future task with the aim to researc h, if the 2.5D U-net also increases accuracy in other applications compared to 3D con volutions. References [1] K. He, X. Zhang, S. Ren, and J. Sun, “Deep residual learning for image recognition,” in Pr o c e e dings of the IEEE c onfer enc e on c omputer vision and p attern r e c o gnition , 2016, pp. 770–778. [2] K. Simony an and A. Zisserman, “V ery deep conv olutional net works for large- scale image recognition,” arXiv pr eprint arXiv:1409.1556 , 2014. [3] J. Long, E. Shelhamer, and T. Darrell, “F ully con v olutional net w orks for se- man tic segmentation,” in Pr o c e e dings of the IEEE c onfer enc e on c omputer vision and p attern r e c o gnition , 2015, pp. 3431–3440. [4] O. Ronneb erger, P . Fischer, and T. Brox, “U-net: Con v olutional netw orks for biomedical image segmen tation,” in International Confer enc e on Me dic al image c omputing and c omputer-assiste d intervention . Springer, 2015, pp. 234–241. [5] Ö. Çiçek, A. Ab dulk adir, S. S. Lienk amp, T. Bro x, and O. Ronneb erger, “3D U-net: learning dense v olumetric segmentation from sparse annotation,” in International c onfer enc e on me dic al image c omputing and c omputer- assiste d intervention . Springer, 2016, pp. 424–432. [6] B. Erden, N. Gam b oa, and S. W o o d, “3D con v olutional neural net w ork for brain tumor segmen tation,” 2018. [7] P . A. Y ushk evich, J. Piven, H. Co dy Hazlett, R. Gimp el Smith, S. Ho, J. C. Gee, and G. Gerig, “User-guided 3D activ e con tour segmentation of anatomi- cal structures: Significantly impro ved efficiency and reliabilit y ,” Neur oimage , v ol. 31, no. 3, pp. 1116–1128, 2006. 11 [8] F. Chollet et al. , “Keras,” h ttps://keras.io, 2015. [9] M. Abadi, A. Agarwal, P . Barham, E. Brevdo, Z. Chen, C. Citro, G. S. Corrado, A. Da vis, J. Dean, M. Devin, S. Ghemaw at, I. Go o dfello w, A. Harp, G. Irving, M. Isard, Y. Jia, R. Jozefowicz, L. Kaiser, M. Kudlur, J. Lev enberg, D. Mané, R. Monga, S. Mo ore, D. Murra y , C. Olah, M. Sch uster, J. Shlens, B. Steiner, I. Sutsk ever, K. T alw ar, P . T uck er, V. V anhouck e, V. V asudev an, F. Viégas, O. Vin y als, P . W arden, M. W attenberg, M. Wic k e, Y. Y u, and X. Zheng, “ T ensorFlo w: Large-scale mac hine learning on heterogeneous systems,” 2015, softw are a v ailable from tensorflo w.org. [Online]. A v ailable: h ttp://tensorflow.org/ [10] A. Rosebro c k, “In tersection o ver union (IOU) for ob- ject detection,” h ttps://www.pyimagesearc h.com/2016/11/07/ in tersection- o ver- union- iou- for- ob ject- detection/, 2016, last accessed: 2019-04-29. [11] L. R. Dice, “Measures of the amount of ecologic asso ciation b etw een sp ecies,” Ec olo gy , v ol. 26, no. 3, pp. 297–302, 1945. [12] S. Ioffe and C. Szegedy , “Batc h normalization: A ccelerating deep net- w ork training by reducing in ternal co v ariate shift,” arXiv pr eprint arXiv:1502.03167 , 2015. [13] I. Go o dfello w, Y. Bengio, and A. Courville, De ep le arning . MIT press, 2016. [14] N. Sriv asta v a, G. Hin ton, A. Krizhevsky , I. Sutskev er, and R. Salakhutdino v, “Drop out: a simple wa y to preven t neural net w orks from o v erfitting,” The Journal of Machine L e arning R ese ar ch , vol. 15, no. 1, pp. 1929–1958, 2014. [15] J. Bro wnlee, “Gentle in tro duction to the adam optimization al- gorithm for deep learning,” https://mac hinelearningmastery .com/ adam- optimization- algorithm- for- deep- learning/, 2017, last accessed: 2019-01-29. 12
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment