Automatic Calcium Scoring in Cardiac and Chest CT Using DenseRAUnet
Cardiovascular disease (CVD) is a common and strong threat to human beings, featuring high prevalence, disability and mortality. The amount of coronary artery calcification (CAC) is an effective factor for CVD risk evaluation. Conventionally, CAC is …
Authors: Jiechao Ma, Rongguo Zhang
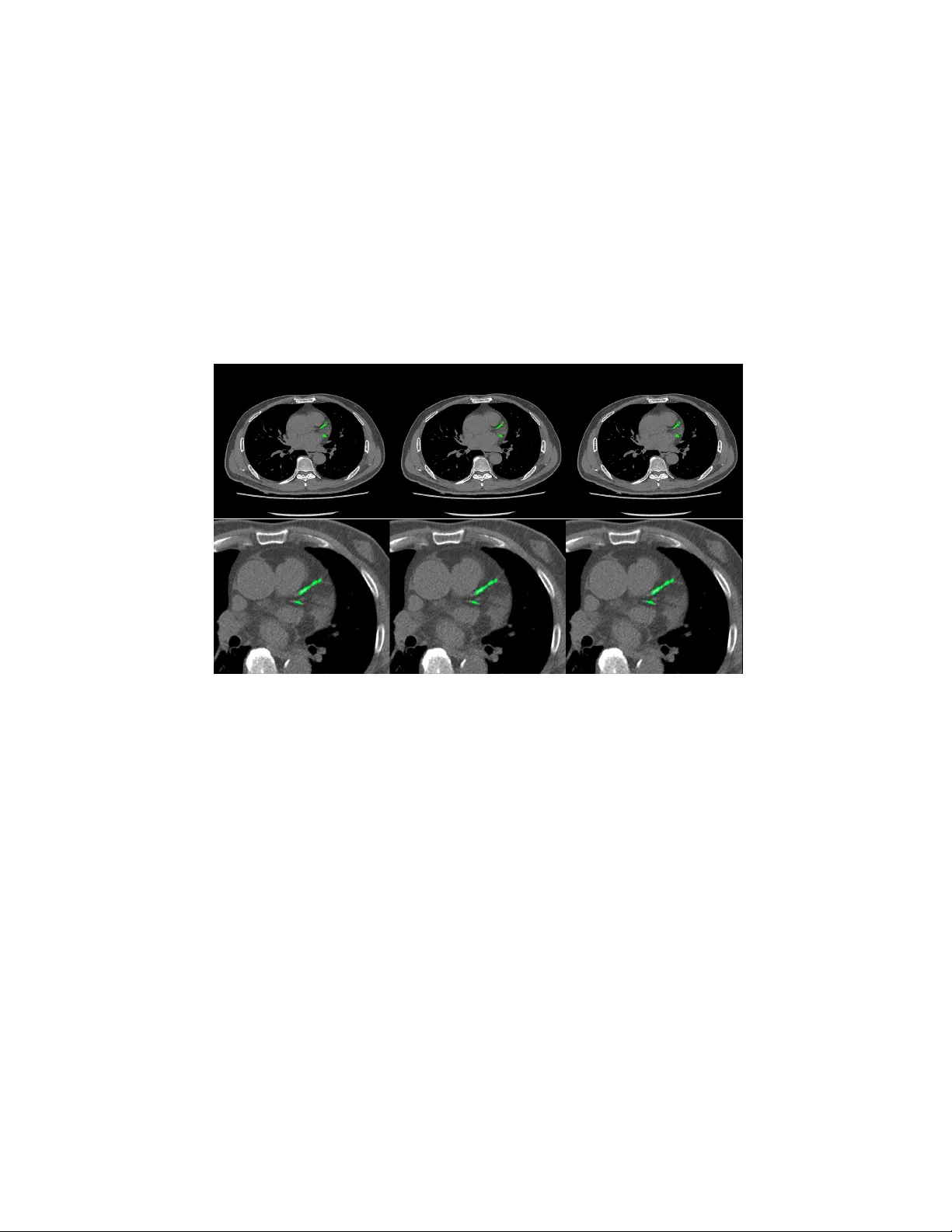
Automatic Calcium Scoring in Cardiac and Chest CT Using DenseRA Unet Jiec hao Ma and Rongguo Zhang Infervision Inc. Abstract. Cardio v ascular disease (CVD) is a common and strong threat to h uman b eings, featuring high prev alence, disabilit y and mortality . The amoun t of coronary artery calcification (CAC) is an effectiv e factor for CVD risk ev aluation. Conv entionally , CAC is quantified using ECG- sync hronized cardiac CT but rarely from general c hest CT scans. How- ev er, compared with ECG-synchronized cardiac CT, chest CT is more prev alen t and economical in clinical practice. T o address this, w e propose an automatic metho d based on Dense U-Net to segmen t coronary calcium pixels on b oth types of CT scans. Our contribution is tw o-fold. First, we prop ose a no vel netw ork called DenseRAUnet, which tak es adv an tage of Dense U-net, ResNet and atrous conv olutions. W e prov e the robustness and generalizabilit y of our mo del by training it exclusively on chest CT while test on both types of CT scans. Second, we design a loss function com bining b ootstrap with IoU function to balance foreground and bac k- ground classes. DenseRAUnet is trained in a 2.5D fashion and tested on a priv ate dataset consisting of 144 scans. Results sho w an F1-score of 0.75, with 0.83 accuracy of predicting cardiov ascular disease risk. Keyw ords: Calcium scoring · deep learning · Cardiac CT · Chest CT · Agatston score · Conv olutional neural net work. 1 In tro duction Cardio v ascular disease (CVD) has b ecome one of the most high-mortalit y dis- eases, for whic h the amount of coronary artery calcification acts as a strong indicator of CVD risk [1]. In clinical practice, CA C is quan tified b y the Agat- ston score, using dedicated cardiac CT scans, follo wed b y a expert who man ually iden tify CA C lesions. T o assist medical professionals, previous w ork based on classical machine learning ha ve attempted to design CAD methods for computation of CAC score. Durlak et al. [2] applied an atlas-based feature approac h in combination with a random forest classifier whic h is used to incorporate fuzzy spatial kno wledge from offline data. Isgum et al. [3] emplo yed a nearest neighbor classifier directly and a tw o-stage classification with nearest neigh b or as well as supp ort v ector mac hine classifiers. There are plent y of other researc h can b e explored [4,5,6]. In recen t years, conv olutional neural net works (CNNs) hav e exhibited great success in Computer Vision by data-driven, esp ecially in image classification 2 Jiec hao Ma and Rongguo Zhang tasks. Mean while, fully con volutional netw orks (FCNs) , as the extension of CNNs, also obtained state-of-the-art p erformance for segmentation problems. In the context of medical image segmentation, sp ecifically cardiac calcification seg- men tation, algorithms based on deep learning ha ve shown promise. W olterink et al. [7] first attempted to apply CNNs to CAC scoring in con trast-enhanced cardiac CT, with a tw o-stage netw ork structure but only one stage using deep learning. Recen tly , some w orks used t wo-stage deep learning structure [8,9], with the first stage identifying CAC-suspected vo xels and the second stage more pre- cisely identifying CAC. Shadmi et al. [10] emplo yed Dense-FCN, a design differ- en t from the tw o-stage methods, to segmen t the lesion directly in cardiac CT. But all the automatic CAC scoring approaches ab o ve are designed for either cardiac or c hest CT only . More recen tly , m ultiple screening in one CT session has b ecome a trend in clinical practice. Huang et al. [11] presented an automatic metho d with t wo CNNs that p erforms direct computation of CAC score in b oth cardiac and chest CT scans. On the other hand, according to the work of W olterink et al. [9], 2.5D input has a great adv antage compared with 3D input in CAC scoring, as the n umber of parameters are greatly reduced while retaining spatial information. Both Lessmann et al. and wolterink et al. [8,9] used the 2.5D Con vNets com- bining features from three identical 2D ConvStac ks with shared w eights, eac h pro cessing an input patc h from a differen t orthogonal viewing direction (axial, sagittal and coronal). T o our kno wledge, none hav e applied the efficien t 2.5D F CN arc hitecture on multiple types of non-enhanced CT. In this w ork, we prop ose an automatic method for CAC scoring on b oth ECG-sync hronized cardiac and chest CT. Unlike the the metho ds required tw o cascaded netw orks to calculate CA C scoring, our net work directly segment the calcified vo xels and obtain CAC scoring. Meanwhile, we adopt a 2.5D patch input to reduce the computational ov erhead of 3D input. Instead of previous 2.5D metho ds [8,9]which input patc h from axial, sagittal and coronal direction, our netw ork tak es 9-channel stacks of images with corresp onding 2D labels for segmen tation of the corresp onding center slice. W e applied our metho d on a priv ate dataset comp osed of 44 Cardiac CT scans and 805 chest CT scans. In comparison to exp erts’ manual annotations, our algorithm achiev ed comp etitive results. 2 Materials and Metho ds 2.1 Data A dataset of 849 CT scans w as collected from several medical cen ters in China, whic h consists of 805 chest CT scans and 44 cardiac CT scans. The CT scans w ere acquired b y different CT scanners with Philips, GE and Siemens. Each CT scan contains a sequence of slices at the thin-section slice spacing (range from 1.0 to 3.0 mm). CAC lesions were manually labe led b y three exp erienced radiologists from differen t cen ters. Automatic Calcium Scoring in Cardiac and Chest CT Using DenseRAUnet 3 W e newly connected 144 CT scans as a test set, incorporating c hest CT scans and cardiac CT scans from medical centers in China. And to ev aluate our net work p erformance, lesions w ere delineated by exp erienced radiologists. 2.2 Data preprocessing Since the connected dataset con tains v arious sizes of chest CT scans and cardiac CT scans, w e process all images as follows. First, w e resize all CT images to 512 × 512 pixel resolution. Second, we randomly crop and then resize images to 512 × 512, where maximum size of the cropped image is 256 × 256. W e con tinuously select nine pro cessed slices as the input of our net work, and for such an input, its lab el is the ground-truth lab el of its middle slice. W e also pro cess all ground- truth lab els to alter the pixel lab el when its corresp onding CT v alue low er than 130HU. 2.3 DenseRA Unet for segmen tation W e prop osed a no vel F CN architecture based on dense U-Net for calcification segmen tation, called DenseRA Unet. The netw ork consists of tw o main comp o- nen ts: (1) a basic netw ork for feature extraction, and (2) three task-specific sub-net work structures, incorporating R esidual A tr ous Unit (RAU) , scSE blo ck and Extr a Dense Blo ck (EDB) . Fig. 1 depicts our prop osed DenseRAUnet. Dense Block 1 Dense Block 2 T Dense Block 3 T Dense Block 4 T upsample block2 upsample block3 !" sample block 1 scSE scSE scSE RAU RAU RAU EDB upsample block Atrous Conv Dilation=2 Atrous Conv Dilation=4 Atrous Conv Dilation=8 BatchNorm BatchNorm BatchNorm ReLU ReLU ReLU Concact + X Identity F(X)+X Residual Atrous Unit Conv Deconv , 3 # 3 Conv , Relu T T ransition Layer Input Output Concat Avg-pooling 2 ×2 Fig. 1. The ov erall structure of DenseUnet and details in Residual Atrous Unit The basic net w ork is an enco der-deco der architecture, similar to dense U-Net. W e adopt a bac kb one netw ork (DenseNet-121) as the enco der sub-netw ork. The deco der sub-netw ork consists of three deco der modules. Each deco der mo dule is 4 Jiec hao Ma and Rongguo Zhang an upsampling blo c k follow ed by a scSE blo c k, where upsampling block contains a deconv olution la yer and tw o conv olution la yers, which follow ed by a Batch Normalization (BN) la yer and an activ ation function called ReLU. Residual A trous Unit. Accurately segmenting v arious sizes of calcified areas ma y require differen t com binations of local and global information. So w e con- sider that a simple skip connection is not enough for the complex segmen taiton task. Inspired by ASPP [12] and embed the idea of Inception [13], we further design a lateral connection called Residual Atrous Unit (RAU). Suc h a mo dule is a residual blo c k, and is used to capture multi-scale information b y combining sev eral conv olutional lay ers with differen t dilation rates in parallel. As shown in Fig.1, we use a concatenation of three 3 × 3 dilated conv olution la yers with dilation rates are 2, 4, and 8 in each RA U. scSE blo ck. T o take full adv an tage of lo cal and global information, w e added scSE blo ck in the deco der sub-netw ork, whic h is in tro duced in [14] for recali- brating the feature maps separately along c hannel and space. Extra Dense Blo c k. In order not to waste the image features extracted from input images, we insert an Extra Dense Blo ck (EDB) in the first skip connection. Suc h a blo c k could make more accurate use of shallo w information, which do not represen t input image in a high dimensional space, via adding more nonlinear in to the first long connection. 2.4 Loss F unction In ter-class imbalances are common problems when using deep learning metho ds for image segmentation, and even more in me dical image segmentation. T o solve it, w e prop ose a new loss function, the com bination of Bo otstrap Loss and IoU Loss: Loss = B ootstr ap Loss + I oU Loss (1) Bo otstrap Loss. When we train a FCN, though images were cropped, there ma y b e thousands of lab eled pixels to predict. How ev er, many of them may b e easily distinguishable, and con tinuing to learn from these pixels do es not impro v e mo del p erformance. In the context of medical image segmentation, most of such pixels are mark ed as bac kground. F or this reason, w e design a weigh ted b o otstrap loss, whic h not only forces net work to fo cus on hard pixels but also balances p ositiv e and negativ e pixels during training. Supp ose there is only one processed image p er mini-batc h and there are a total N pixels to predict. There are only tw o categories c j in the lab el space. Let y i denotes the ground-truth lab el of pixel x i , and p i,j denotes the predicted probabilit y that pixel x i b elongs to the category c j . Then, the loss function could b e defined as: l = − α P i ∈ N ,j =0 log p i,j 1 { p i,j < t and y i = j } P i ∈ N ,j =0 1 { p i,j < t and y i = j } + β P i ∈ N ,j =1 log p i,j 1 { y i = j } P i ∈ N ,j =1 1 { y i = j } (2) Automatic Calcium Scoring in Cardiac and Chest CT Using DenseRAUnet 5 where t is a threshold. Here 1 {·} is equal to one when the condition in parenthe- ses, and otherwise is zero. In other words, we fo cus all p ositive pixels and drop negativ e pixels when they are too easy for the curren t mo del, i.e. their predicted probabilit y greater than t . In practice, we hope that positive and negative pixels are balanced, hence we add α and β as trade-off co efficients. IoU Loss. Bootstrap loss is similar to cross en tropy loss, focusing more on its o wn predictions of pixels and ignoring the relationship b et ween adjacent ones. T o b etter obtain the boundary of lesion, w e add IoU in the loss function using suc h a relationship. Supp ose there are N pixels to predict. T o ensure that losses are on the same magnitude, we use the following exp onen tial form of IoU: iou l oss = − ln P i ∈ N p i g i P i ∈ N p i + P i ∈ N g i − P i ∈ N p i g i (3) where p i is the predicted probabilit y of pixel x i , g i is the ground-truth lab el of pixel x i . 2.5 P ost-pro cessing The final segmentation result of the netw ork is obtained b y a predefined thresh- old (here set to 0.5), and each lesion segmented b y the netw ork is considered a calcification candidate. Then each candidate is classified as CA C b y thresholding with 130 HU and p erforming connected-components analysis. Since the CT slice thic kness is mostly 1mm, calculation of the final Agatston score for the whole v olume is done b y the following corrected form ula: Ag atston S cor e = X i X n f i,n A i,n ∆S 3 (4) where i is the i th CT slice of a CT volume, n is the n th selected lesion, f is the weigh ted intensit y , A is the lesion area, and ∆S is the slice spacing (mm). 3 Exp erimen ts and Results Ev aluation Metric. W e ev aluate the pixel-level segmentation pe rformance of the netw ork by F1 score: F 1 = 2 · P r ecision · Recal l P r ecision + Recal l (5) W e also define CAC r ate denotes the prop ortion of patien ts who w as correctly predicting the CVD risk level without p ost-processing, and CA C filter R ate rep- resen ts the prop ortion of patien ts with p ost-pro cessing. Implemen tation details. The exp erimen ts conducted were all trained from scratc h and initialized b y the Gauss metho d. During training, w e collected one pro cessed image as a mini-batch for each iteration and trained for 25 ep ochs. T o 6 Jiec hao Ma and Rongguo Zhang optimize these experiments with fast conv ergence, w e employ ed the SGD opti- mizer with momentum of 0.9. The initial learning rate is 0.001 and is reduced b y 0.99 times per 2000 iterations. The parameters in the loss function are exp er- imen tally set as t = 0 . 9, α = 8 and β = 1. W e implemen ted all the exp eriments via the deep learning toolki MXNet and trained on a GTX 1080 (NVIDIA) GPU. Ablation exp erimen ts. W e use “Dense U-net & Bo otstrap Loss” as the base- line for all exp eriments. T o ev aluate the effectiv eness of v arious structures in our metho d, we conducted ablation exp erimen ts. First, using the b o otstrap loss, w e compare the role of three mo dules in the netw ork. Second, we studied the effect of tw o loss functions through trained our netw ork. T able 1. Comprasion of p erformance of the basic netw ork using different tricks Basic network Bo otstrap RA U EDB scSE IoU F1-Score Dense U-Net √ 0.65 √ √ 0.68 √ √ √ 0.69 √ √ √ √ 0.71 √ √ √ √ √ 0.75 Ours T able 2. Qualitativ e results of CAC rate and CA C filter rate for patients, Patien ts represen ts the total num b er of patients, CA C No. and CA C fliter No. represents the n umber of patients w ere predicted correctly by model and p ost-processing resp ectively . T ric ks CAC No. CAC filter No. Patien ts CAC Rate CAC filter Rate Dense U-net 101 99 144 0.70 0.69 Dense U-net+RAU 104 111 144 0.72 0.77 Dense U-net+RAU+EDB 109 115 144 0.76 0.80 Dense U-net+RAU+EDB+scSE 109 117 144 0.76 0.81 Our prop osed metho d 113 120 144 0.78 0.83 T able 1 lists the F1 scores of Dense U-Net using different tricks, T able 2 indicates the performance of corresp onding netw ork arc hitectures in T able 1 on CA C. It is sho wn that all the tric ks provide increase in F1 score and CAC in comparison to the baseline. W e further observ e that adding RA U in the net work ac hieves more significant improv emen t for CAC segmen tation. Comparing the results across T able 1, our metho d yields the best p erformance. F rom T able 2 w e can also conclude that p ost-processing b y the definition of CA C score is essential. 4 Conclusion This paper prop osed an algorithm based on deep learning. Our method consists of t wo core elements: (1) a no v el fully con volutional netw ork, DenseRA Unet, and Automatic Calcium Scoring in Cardiac and Chest CT Using DenseRAUnet 7 (2) a loss function combined b ootstrap loss and IoU. W e trained our netw ork in a 2.5D-patc h fashion to reduce input parameters while preserving spatial information. While trained solely on chest CT, our mo del achiev ed comp etitiv e and robust p erformance on b oth chest CT and cardiac CT which has significant higher resolution and lo wer spacing compared to training data, thanks to the p o w er of residual atrous unit that enlarges the receptiv e field with down ward compatibilit y . W e aim to futher explore and extend our method to other medical image analysis c hallenges in future w ork. Fig. 2. Segmentation results of chest CT (top) and cardiac CT (b ottom). F rom left to righ t: the segmen tation result of our mo del without p ost-processing, the result of with p ost-processing, ground truth. References 1. John A Rum b erger, Bruce H Brundage, Daniel J Rader, and George Kondos. Electron beam computed tomographic coronary calcium scanning: a review and guidelines for use in asymptomatic persons. In Mayo Clinic Pr o c e e dings , volume 74, pages 243–252. Elsevier, 1999. 2. F elix Durlak, Michael W els, Chris Sch wemmer, Michael S ¨ uhling, Stefan Steidl, and Andreas Maier. Growing a random forest with fuzzy spatial features for fully automatic artery-sp ecific coronary calcium scoring. In International Workshop on Machine L e arning in Me dic al Imaging , pages 27–35. Springer, 2017. 3. Iv ana Isgum, Mathias Prokop, Meindert Niemeijer, Max A Viergev er, and Bram V an Ginneken. Automatic coronary calcium scoring in lo w-dose chest computed tomograph y . IEEE tr ansactions on me dic al imaging , 31(12):2322–2334, 2012. 8 Jiec hao Ma and Rongguo Zhang 4. Uda y Kurkure, Deepak R Chitta jallu, Gerd Brunner, Y en H Le, and Ioannis A Kak adiaris. A sup ervised classification-based metho d for coronary calcium de- tection in non-con trast ct. The international journal of c ar diovascular imaging , 26(7):817–828, 2010. 5. Rahil Shahzad, Theo v an W alsum, Michiel Schaap, Alexia Rossi, Stefan Klein, Annic k C W eustink, Pim J de F eyter, Lucas J v an Vliet, and Wiro J Niessen. V essel specific coronary artery calcium scoring: an automatic system. A c ademic r adiolo gy , 20(1):1–9, 2013. 6. Jelmer M W olterink, Tim Leiner, Ric hard AP T akx, Max A Viergev er, and Iv ana I ˇ sgum. An automatic machine learning system for coronary calcium scoring in clinical non-contrast enhanced, ecg-triggered cardiac ct. In Me dic al Imaging 2014: Computer-Aide d Diagnosis , v olume 9035, page 90350E. In ternational Society for Optics and Photonics, 2014. 7. Jelmer M W olterink, Tim Leiner, Max A Viergever, and Iv ana I ˇ sgum. Automatic coronary calcium scoring in cardiac ct angiography using conv olutional neural net- w orks. In International Confer enc e on Medic al Image Computing and Computer- Assiste d Intervention , pages 589–596. Springer, 2015. 8. Nik olas Lessmann, Bram v an Ginneken, Ma jd Zreik, Pim A de Jong, Bob D de V os, Max A Viergever, and Iv ana I ˇ sgum. Automatic calcium scoring in lo w-dose chest ct using deep neural net works with dilated conv olutions. IEEE tr ansactions on me dic al imaging , 37(2):615–625, 2018. 9. Jelmer M W olterink, Tim Leiner, Bob D de V os, Robb ert W v an Hamersvelt, Max A Viergever, and Iv ana I ˇ sgum. Automatic coronary artery calcium scoring in cardiac ct angiography using paired conv olutional neural netw orks. Medic al image analysis , 34:123–136, 2016. 10. Ran Shadmi, Victoria Mazo, Orna Bregman-Amitai, and Eldad Elnek a ve. F ully- con volutional deep-learning based system for coronary calcium score prediction from non-con trast chest ct. In 2018 IEEE 15th International Symp osium on Biome dic al Imaging (ISBI 2018) , pages 24–28. IEEE, 2018. 11. Gao Huang, Zhuang Liu, Laurens V an Der Maaten, and Kilian Q W einberger. Densely connected con volutional netw orks. In Pr o c e e dings of the IEEE confer enc e on c omputer vision and pattern re c o gnition , pages 4700–4708, 2017. 12. Liang-Chieh Chen, George Papandreou, Iasonas Kokkinos, Kevin Murphy , and Alan L Y uille. Deeplab: Semantic image segmentation with deep conv olutional nets, atrous conv olution, and fully connected crfs. IEEE transactions on p attern analysis and machine intel ligenc e , 40(4):834–848, 2018. 13. Christian Szegedy , W ei Liu, Y angqing Jia, Pierre Sermanet, Scott Reed, Dragomir Anguelo v, Dumitru Erhan, Vincen t V anhouc ke, and Andrew Rabinovic h. Going deep er with con volutions. In Pr o c e e dings of the IEEE c onfer enc e on c omputer vision and p attern r e c o gnition , pages 1–9, 2015. 14. Abhijit Guha Ro y , Nassir Nav ab, and Christian W achinger. Concurrent spatial and c hannel squeeze & excitationin fully con volutional net works. In International Con- fer enc e on Me dical Image Computing and Computer-Assiste d Intervention , pages 421–429. Springer, 2018.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment