Graph-Based Method for Anomaly Prediction in Brain Network
Resting-state functional MRI (rs-fMRI) in functional neuroimaging techniques have improved in brain disorders, dysfunction studies via mapping the topology of the brain connections, i.e. connectopic mapping. Since, there are the slight differences be…
Authors: Jalal Mirakhorli, Hamidreza Amindavar, Mojgan Mirakhorli
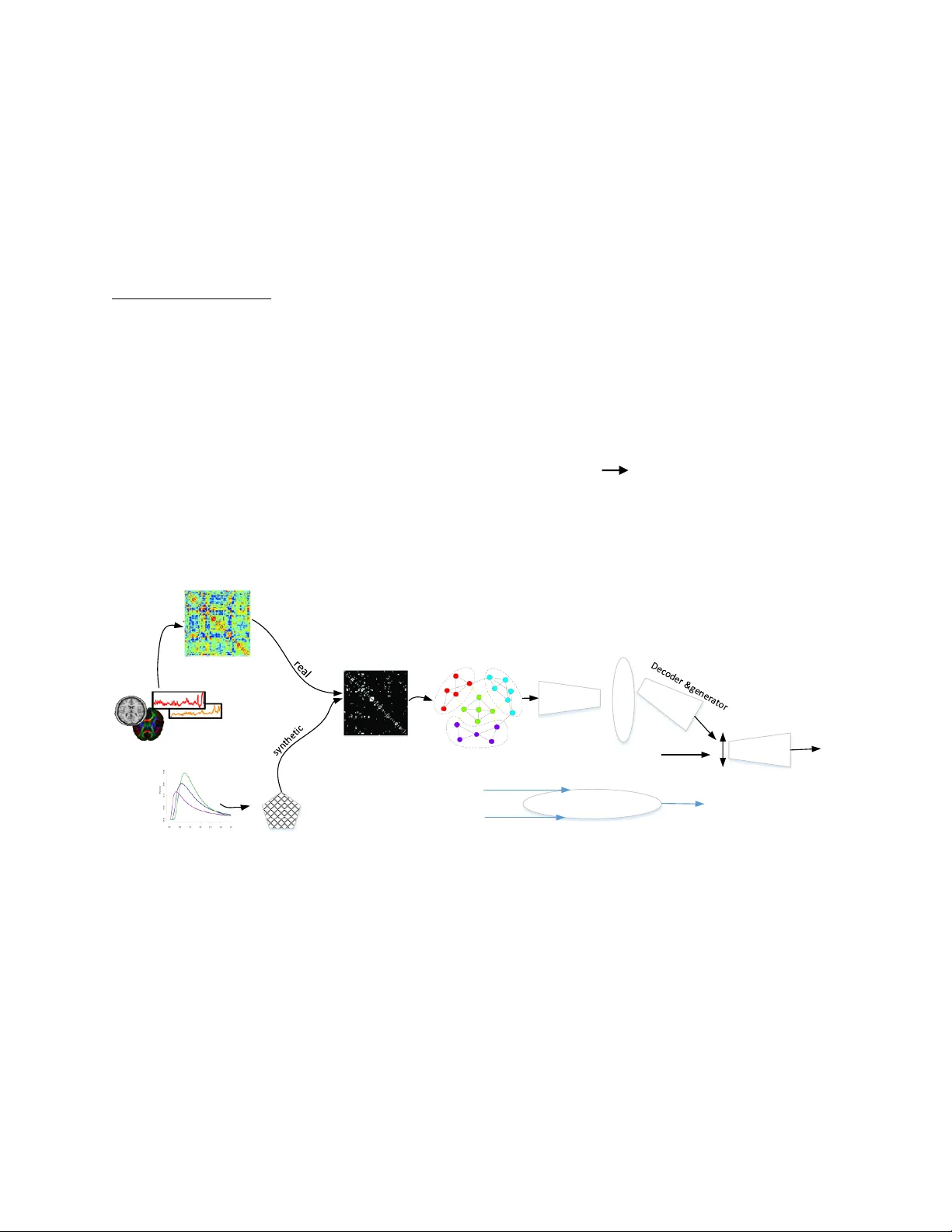
1 Graph-Based Me thod for Ano m aly Prediction In Brain Netw orks Jalal Mirakhorli 1 jalalmiry@aut.ac.ir Hamidreza Amindavar 1 hamidami@aut.ac.ir Mojgan Mirakhorli 2 genomic66@gmail.com 1 Department of El ectrical Eng ineering, Am irkabir Univ ersity of Technolog y. 2 Medical Genetic Lab, I ranian Com prehensive Hem ophilia Care Cen ter (ICHCC). Abstract : Functional magnetic resonance imaging (fMRI) in neuroimag ing techniques have improved in brain disorders, dysfunction studies via mapping the topology of the brain connections, i.e. connectopic mapping. Si nce, there are the slight differences between healthy and unhealthy brain regions and functions, in vestigation in to th e complex topology of functional and structural brain networks in human is a complicated task with the growth of evaluation criteria. Irregular graph d eep learning applications have widel y spread to understanding human cognitive functions that are linked to gene expression and related distributed spatial patterns, because the neuronal networks of the brain can hold dynamically a variety of brain solutions with different activit y patterns and functional c onnectivity, these applications might also be involved with both node- centric and graph-centric tasks. In this paper, we performed a novel approach of individual generative model and high order graph analysis for the region of interest recognition areas of the brain which do not h ave a normal connection during appl y ing certain tasks and resting-state or decompose ir regular obs ervations. Here, we proposed a high order fra mework of Graph Auto- Encoder (GAE) with a hypersphere distributer for f unctional data anal y sis in brain imaging studies that is underlying non-Euclidean structure in the learning of strong non-rigid graphs among large scale data. I n addition, we distinguish ed the possible modes of correlati ons in abnormal brain connections. Our finding will show the de gree of correlation between the affected r egions and their simultaneous occurrence over time that ca n be u sed to dia gnose brain dis ease s or revealing the ability of the nervous s ystem to modify in brain topolog y at all angles, brain plasticity, according to input stimuli. Keywords : Semantic Brain Ne tworks, Graph Theory, Ge nerative Mode l, Neural Plastici ty, Posterior Contrac tion. Introduction : 2 The human brain has a c omplex connection of various parts which d y namically shift durin g their operation. Therefore , the model and cost of each part is able to change according to the type of its operation in carried out or rest state. The fMR I dat a exhibits non-stationary properties in the context of task-based studies (Hutchison et al . 2013; Calhoun et al. 2014), the analysis of these sections is known to predict the conne ction factors for each indepe ndent profile. Here, we present a theor etical model based on high order VAE and graph theor y to learn the probability distribution of the graph that is known to extract the data model of tasks from brain regions with a semi- unknown prior knowledge method. We used each functional connectivit y matrix tha t is collected in a resting-state fMR I (rs-fMRI) experiment, u sing rs-fMRI data f rom Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset. Functional connectome anal y sis is recognized to reveal biomarkers of individual psy chological or clinical trait s and describes t he pairwise statistical dependencies between brain regions. In this article, we present the brain as a graph b y m eans of functional connectome s tructures. This allows us to probing and infer, how dynamic changes progress of im provement degree in brain disorder or predict the disease as well as identif y the term brain abnormalities. This paper proposes to introduce a framework fo r feature extraction of the brain graphs which provide across many subjects, for prediction of ambiguous parts of the brain. In t his method a Variational Autoencoder ( VAE ) is established to make the graph and experiment a Ba yesian Von Mises – Fisher (VMF ) (Mardia et al. 1976) mixture model as a latent distribution that can place mass on the surface of the unit hypersphere (Banerjee et al. 2005) and stable the VAE. Our ex periments demonstrate that this method significantly outperforms other methods and is a large step forward to infer brain structure. It is capable to handle both homogeneous and heterogeneous graphs. According to the recent studi es, the geometric deep learning methods hav e been successfull y applied to data residing on g raphs and manifolds in terms of various tasks (Bronstein et al. 2017; Mirakhorli et al .2017). For example, function of the brain in predicting and its graph expression analysis a ddress the multifaceted challenges arising in diagnosis of brain dis eases. Here, we present a n ovel method using a hi gh order gra ph model in revealing the relationship between the p arts of brain and recover missing p arts or malfunctioning parts. The method can also be predicted the effects of long-term deep brain stimulation on brain structural and functional connectivity. Related work s : As our approach focuses on completing the graph and predictive defective parts of the graph via obtained feature of netw ork embedding, we review some of th e state- of -the-art research that are close to our work. Xu et al. 2017 construct ed a graph from a set of obje ct proposals to provide initial embedding to each node and edge while using message passing to obtain a consistent prediction. Simonovsky et al. 2018 used a generative model to produce a probabilistic graph from a single opaque vector without specifying the number of nodes or the structure explicitl y . Pan et al . 2018 proposed an adversarial training s cheme to regularize a nd enforce the latent code to mat ch a prior distribution with a graph convolutional Autoencode r. Makhzani et al. 2015 showed an 3 adversarial Autoencoder to learn the latent embedding by m erging the adversarial mec hanism into Autoencoder for general data. Howe ver Dai et al. 2017 applied the adversarial procedure for the graph embedding . Also an encoder with edge condition convolution (ECC) (Johnson et al. 2017) was used for conditioning both encoder and decoder which was associated with each of the input graph s (Sim onovsky at al. 2018), this m ethod is useful only for generation small graph s. In addition, we used a combination of graph convolution VAE to address both re covery and learning problems which can perform in spectral (Defferrard et al. 2016 ; Levie et al. 2019) or spatial domains (Monti et al. 2017). Materials and Method In spite of individual alteration, the human brains perform common patterns among different subjects. Therefore, algorithms based on graph s are essential tools to capture and model the complicated relationship betwe en functional connectivity. In this work, we used a model of graph embedding to convert graph data into a low dimens ional and continuous compac tion fe ature space that is able to detect abnormal parts of input graphs which is involved with graph matchin g and partial graph completion problems ( Verma et al. 2018). To develop this algorithms, we ne ed to present a generative model that is construct ed from a high order Graph Variation al Autoencoder with hypersphere distribution (Da vidson et al. 2018 ; King ma et al. 2014; Kipf et al. 2016). Pa rtial abnormality is able to be demonstrated b y features train in latent space, considering both first- order proximit y , the local pairwise proxim ity between the vertices in the network, and second - order proximity. This refers to vertices sharing many connections to other vertices that are similar to each other. The work flow of the algorithm is shown in figure 1. Brain n etwork as a graph: As shown in figure 1 , using rs-fMRI data of subjects acquired b y preprocessing ADNI dataset to provide an adjacency matrix that encodes similarities betwee n nodes and a feature matrix that repre sents a node’s connectivity profile, to de fine the input data as an undirected graph. In t his paper, we define the connected graph G = (V, E, W), which consists of a finite set of vertices V with |V| = n, a set of edges E, and a weighted a djacency matrix W . I f there is an edge e = ( i, j) connecting vertices i an d j, the entr y W ij or a ij repre sent the weight of the edge a ij>0 , otherwise a ij = 0. For each of n subjects make a data matrix X n ε R d n*dy , where d y is the dimension of the node’s feature v ector. This structure of fMRI d ata will be merge int o the graph de fined. W e will show that applying the graph bas ed algorithm on brain connectivity is useful to analyze brain information processing. Graph Convolutional Neura l Network: For applying convolution-like operators over irregular local supports, as graphs where nodes can have a v arying number of neighbors which c an be used as layers in deep networks, for node classification or recommendation, link prediction and etc. This process is involved with three challenges, a) defining translation structur e on graphs to allow parameters sharing, b) designing compactl y supported filters on graphs, c) agg regating mul ti-scale information, the propose d strategies broadl y fall into two domains, there i s one spatial operation which directly performs the convolution by aggregating the neighbor nodes’ information in a certain batch of the graph, where weights can be easily shared across different structures (Niep ert 4 et al. 2016; Gao et al.2018) and the second is a spectral operation which relies on the Eigen- decomposition of the Laplacian matrix that is app lied to the whole graph at the same time (Henaff et al. 2015; L evie e t al. 2019; Bruna et al. 2014; Kipf at al. 2017), spectral-based decomposition is often unstable makin g th e generalization across different graphs difficult (Pan et al. 2018), that cannot prese rve both the local and g lobal netwo rk structures which require large memor y and computation. On the other hand, local filtering approaches (Boscaini at al. 2016) rel y on possibly suboptimal hard-coded local pseudo- coordinates over graph to defin e filters. The third approa ch relies on point-cloud representation (Klokov et al. 2017) that cannot leverage surface information encoded in meshes or ne ed ad-hoc transformation of mesh data to map it to the unit sphere (Sinha et al. 2016). Overa ll, the spectral approach has the limitation of gra ph structure being the same for all samples i.e. homogeneous structure, thi s is a hard constraint, as most of sample graphs in the learning phase have the same structure and siz e for diffe rent samples i.e. he terogeneous stru ctures. Therefore, we applied the spatial app roach that is not oblig atory homo geneous graph structu re, but in turn requires preprocessing of graph to enable learning on it and used a method that proposes a graph embed pooli ng. Graph convolution transforms only the vertex values (Such et al. 2017) whereas graph pooling transforms both the vertex values a nd the adj acency matrix. Convolution of vertices V with filter H onl y requires matrix multiplication of the form, υ out =Hυ in whe re υ in , υ out ε R N*N . the filter H is defined as the k-th degree polynomial of the graph adjacency matrix A; H=h 0 I+h 1 A+h 2 A 2 +…+ h n A k , H ε R N*N . (1) We used the first three taps of H for any given filter. Graph Autoenc oder (GAE): GAE is inherently an unsupervised generative model, our model is almost based on the framework of VAE that wa s produced in (Kipf et al. 2016; Xu et al. 2018). I n follow, we briefly describe GAE and introduce our method with obj ectives. For learning both encoder, decoder in the figure 1 to map between the space of graph and their continuous embedding Z ε R C , stochastic graph encoder q Φ (Z|G) embed th e graph into c ontinuous representation Z. Given a point in the latent space Z, the graph decoder p θ (G|Z) outputs a probabil istic full y-connected graph Ğ on pr edefined nodes, where Φ,θ are learned parameters. Reconstruction ability of GAE is facilitated b y approximate gra ph matchin g for aligning G with Ğ, as well as a prior distribution P(Z) imposed on the late nt code representation a s a reg ularization and tra in GAE via optimization of the marginal likelihood, P(G) = , then the marginal log likelihood can be written; log p θ (G) =ƘĹ(q Φ (Z|G) || p θ (Z|G)) + ₤(θ,Φ;G) . (2) ₤ ( θ,Φ;G) = - ƘĹ[q Φ (Z|G) || p θ (Z) + log p θ (G|Z)]. (3) Where Kullback – Leibler ( Ƙ Ĺ) and q Φ (Z|G) a re a divergence ter m in loss function that encourages the variational posterior and a v ariational approximation to the posterior distribution, p (Z|G), 5 respectively. Here, we used a h ype rspherical latent structure for parameterization of both prior and posterior, because one of important limitation in using Gaussian mixture is that ƘĹ term may encourage the posterior distribution of the latent variable to collapse in prior or tends to pull the model toward the prior, during approximation the prior, whereas in the VMF (Fish er et al.1953; Kanti et al. 1976) case there is not such pressure toward a single distribution convergence. Therefore a VMF dist ribution is more suitable for capturing da ta (Kipf et al. 2016), VMF distribution defines a probabilit y densit y over p oints on a unit -sphere also the consequences of ignoring the underl y ing spherical manifold are rarel y an alyzed in parts due to computational challenges imposed by directional statistics. Geometric d eep le arning: For graph generation, we employed the G AE to graph G ε R n*m under an unsupervise d learning method, our goa l is to learn an implicit ge nerative mode that can predict abnormal sections in the graph, indeed, we are not confident that close li nks have similar features to detect invisible deformable and hidden angle of graphs. Our method almost inspired in previous studies (Larsen et al. 2015; Wu et al. 2016), in combination from the GAE and generative adversarial network (GAN) that decoder of GAE and generator of GAN h ave been a supportive role. Followin g the above mentioned items, we used the uniform dist ribution VMF(0,Ҡ =0) for our prior and approximate p θ (Z | Ğ) with variational posterior q Φ (Z|G) = VMF(Z; μ, Ҡ), where μ is mean parameter and Ҡ is a constant, the variational distribution is associated with a prior distribution over the latent va riables, our GAE loss combines the g raph reconstruction Ĺ r =|| Ğ – G || 2 encouraging concatenation both the encoder-decoder to be a nearly identical transformation, a regularization prior loss measured b y the ƘĹ divergence, Ĺ p =D ƘĹ (q(z|G)|| P(Z)) and a cross entropy loss Ĺ 2D -GAN for GAN, Ĺ G- GAN =log D(G)+ log (1-D(G(z))), where D is discriminator as a confidence D(G) of the whether a input graph G is real or s ynthetic (Levie et al. 2019). The total GAE+GAN loss is computed as Ĺ= Ĺ r +λ 1 Ĺ p +λ 2 Ĺ G-GAN , where λ 1 and λ 2 are weig hts of ƘĹ divergence loss and re construction loss. As discussed in above, ou r desire to fo cus on graph completion for deformable object classes in brain connectome. Therefore, we used d y namic weight of filtering in each convolutional la y er. Partial graph completion: Once our model GAE&GAN has been tr ained, the encoder and the element of GAN are discarded away, so that the r ole of the d ecoder is onl y as a graph generator where the probabilistic latent space z a cts as a base for finding the tar get graphs for the same g raph prior. At inference, for e ach space o f the latent vector z * may represent a few complete graph correspondence a latent v ector, then partial graph or deformation graph in t he input of the system makes a few complete graphs in the output, the higher deformation rate in input, the more of graph is gene rated. Each partial graph represents a partial adjacency matrix δ that can be applied to any graph Ğ generated b y our mo del and to explore similarity between them, for finding best compatibility or a latent vector z* which can minimiz e differences between input and output graph, to provide more geometric insight on the problem. Process to measure sim ilarities among elements 6 of graphs with probing combination of dependences similar unary, pairwise or high-order ( Le -Huu et al. 2017; Yan et al. 2018; Yu et al. 2016) as well as there are potentials between reference graph and their counterparts si milar to previous stud y (Wang et al.2019; W ang F et al. 2018 ), that follow ed a function used for finding high or der dissimilarity or d eformation into convex optimization problem over a set of doubly stochastic matrices. Graph recover y plan: As mentioned above, our goal is the opti mal choice of a latent vector z* so that minimal dissimilarities exists of between the partial graph related to a disease brain, G , and the generated graph Ğ = dec (z), or m in (Ğ, ζG) . Where ζ denote a non-rigid transformation, this procedure is performed over z * and ζ, alternatively. Minimizing the following function is our goal as an objective function; m in j(p, ζ) = + γ (ζ) . (4) Where γ is a regularization term of geometric transformation ζ: Ğ G, p is a map for measuring the difference of graph attributes in a similar transformation domain. In each step of optimization a weight matrix measures the degree of deformation on the radial basis function method . Graph recovery i s an ill -posed problem that has multiple plausible solution s while in this paper we limit the prediction space to only several structures of the graphs. DI SC RE MI NA TO R 1 Anatomical or fun c t ional imaging Connection matrix Adjacency ma t r ix sampler Wis hart distr ibut ion Graph pool D i s c ri minato r encoder Ž G Ğ loss Gr aph mat ching G Mu ltiple Ğ Be st gra ph Fig. 1. The data flow of the propo sed network architecture. Con clu si on Thr ou gh thi s pa per , we hav e p res en ted a n ovel joi nt pr edi cti on app roa ch for the s ele cti on o f mea ni ng ful p art ial cor re lat ion an d ext rac t rol e of f un cti on al co nn ecti v i ty st at es fr om rs -f MR I to des c ribe t he d ynam ic co nn ecti on s. This met hod hel ps to im pr ove bi oma rk er dis co ve ry esp eci al l y i n hi gh -d im en sio nal s ett in gs wit h a lar ge n um b er o f v ari ab les co nn ecti o ns . Thi s 7 app ro ach al so all ow s u s to re cov er da ta c or rupt e d b y n o is e an d e xp lor e in d o mai n of un kno wn fun ct ion . W e s how ed t hat th e m od eli ng l ate nt s pa ce w ith t he hyp ers ph er e d ist ri but e r im p rov e acc ur ac y in pr edi ct io n c on nec tiv it y st at es of th e b rai n. Ba sed on th e abo ve a nal ysi s an d f ocu s on th es e t opo lo gic al att ri but es t o ex te nd this wo rk , we will ex tr act imp or ta nt inf orm ati on in th e fut ur e o n th e h i ghe r -o rd er fu nct io n of th e b rai n net wo rk vi a se ma nti c fu nct io nal ri ch -clu b or gani za ti on. Ack now le dg e men t The a uth or s gra te ful l y a ckn ow led ges t he assis t anc e p rov i ded b y t he M ed ica l Ge net ic Lab , Ira ni an Com pr eh ens iv e Hemo ph ili a Ca re C ente r (IC HC C). References: Hutchison RM, Womelsdorf T, Allen EA, Bandettini PA, Calhoun V D, et al (2013) Dy namic functional connectivity : pr om ise, issues, and interpretations. N euroimage 80, 360 – 378. https://doi.org/10.101 6/j.neuroimag e.2013.05.079. Calhoun V. D, Miller R, Pearlson G , and Ada li T, (2014) T he chronnectome: time-v arying connectivity networks as the next fr ontier i n fMRI data discov ery .Neuron 84, 262 – 274 . https://doi.org /10.1016/j.neuron.2014.10.0 1. Mardia K. V, El -A toum, S (1976) Bayes ian inference for t he von Mi ses-Fisher distribution. B iometrika , 63, 203 – 206. http s://doi.org/10.230 7/2335106. Banerjee A, I nderj it S, Dhi llon, Ghosh, Sra S, R idgeway G (2005) Clustering on the uni t hypersphere using von Mises-Fisher distributions. J ournal of Machine Learning Research. 1345-1382. https://doi.org /10.1.1.76.908. Bronstein M. M, Bruna J , LeCun Y, Szlam A, Vandergheynst P (2017) Geometric deep l earning: Going beyond Euclidean data. IEEE Signal Process. Mag., vol. 34, no. 4, pp. 8 -42, Jul. https://doi.org /10.1109/MSP.2017.2693 418. Mirakhorli J, Amindav ar H (2017) Sem i -Supervised Hierarchical Semantic Object Parsing. IEEE Conference on S ignal Processing and Intelligent Sy stems. https://doi.org / 10.1109/I CSPIS.2017.8311588. Xu D, Zhu Y, Choy C. B, Fei-Fei L (2017) Scene Graph Generation by Iterative Message Passing. 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR).https://do i.org/10.1109/C VPR.2017.330. Simonov sky M, Komodak is N (2018) GraphVAE: Towards g eneration of sm all graphs using v ariational autoencoders. I nternational Conferenc e on Artifici al Neural Network s ICANN 2018. ht tp s: // do i .or g/ 1 0. 10 07 /9 78 -3 - 030 - 01418 -6 _4 1. Pan S, Hu R, Long G, Jiang J. F, Yao L, Zhang C (2018) adversarially regulariz ed graph autoencod er for graph embedding. Proceedings of the Twenty- Seventh I nternational Joint Conference on Artificial Intelligence (I J CAI-18). pp. 2609 – 261 5. https://doi.o rg/10.24963/i jcai.2018/362. Makhzani A, Shlens J , Jaitly N, Goodfellow I, Fr ey B (2016) Adversarial autoencoders. Internationa l Conference on Le arning Representat ions. abs/1511.05644. 8 Dai Q, Li Q, Tang J , and et.al (2018) Adversarial network embedding. In Proc. of 2018 AAAI Conf. on Artificial I ntelligence (AA AI’18), New Orl eans . arXiv: 1711.07838, (201 7 ). D Johnson D (2017) Learning g raphical sta te transitions. 5th I nternational C onference on Learning Representations, I CLR 2017 . Defferrard M, Bresso n X, Vanderg heynst P (2016) C onvolutional neu ral network s on graphs with fast localized spectr al filtering . I n NIPS, (2016). Levie R, Monti F , Bresson X, Brons tein M (2019) Ca yleynets: Graph conv olutional neur al network s with complex rational spectral fi lter s. IEEE Transactions on Signal Proces sing, vol. 67, no. 1, pp. 97 – 109 . https://doi.org /10.1109/TSP.2018.2879 624. Monti F, Boscaini D, Masci J , Rodola E, Svoboda J, Bronstein M (2017) Geometric deep learning on graphs and m anifolds using m ixture m odel cnns. I n Proceedin gs of the IEE E Conferenc e on Com puter Vision and Pattern Recognit ion, pages 5115 – 5124 .https://doi.org/ 10.1109/CVPR.2017.576. Verma N, Boyer E, Verbeek J (2018) FeaStNet: Feature- Steered Gr aph C onvolutions for 3D Shape Analysis. In CVPR. ar Xiv:1706.0520 6v2. Davidson T, Falorsi L, De Cao N, Kipf T, Tom czak J (2018) Hy perspherical Variational Au to-Encoders . 34th Conference on Unc ertainty in Artificial I ntelligence (UAI-18). Kingma D. P, Wel ling M (2014) Au to-Encoding Vari ational Bayes. I n ICLR. arXiv:1312.6114v 10. Kipf T. N, Well ing M (2016) Variational g raph auto- encoders. NIPS. Niepert M, Ahmed M, K ut zkov K (2016) Learning Convolutional Neural Networks for Graphs. I n I CML. 2014 – 2023. arXiv :1605.05273v4. Gao H, Wang Z, Ji S (201 8) Larg e-scale learnabl e graph convolutional n etworks. in Proceedings of the ACM S I GKDD International Conference on Know ledge Discovery & D ata Mining. ACM, pp. 1416 – 1424. https://doi.org /10.1145/3219819.3219 947 Henaff M, Bruna J and LeCun Y ( 2015) Deep convolutional networks on graph-structur ed data. https://arxiv.org /abs/1506.05163. Bruna J, Zaremba W, Szla m A, LeCun, Y (2014) Spectral networks and locally connecte d networks on graphs. In I CLR. arXiv:1312.6203v 3. Kipf N, Welling M (2017 ) Semi- supervised classification with graph convolutional networks . ICLR. arXiv:1609.02907v 4. Boscaini D, Masci J, Rodol` E, Bronstein M (2016) Learning shape correspondence with anisotropic convolutional neural ne tworks. In Advances in N eural Information Processing Sy stems, pages 3189 – 3197. Klokov R, Lem pi tsky V (20 17) Escape from cells: de ep K d-N etworks for the recognition of 3D po int cloud models. I n IEEE Int. Conf. on C omputer Vision (ICCV ). Sinha A, Bai J, Ramani K ( 2016) Deep l earning 3D shape surfaces using geometry images. In: European Conference on Com puter Vision. pp. 223 – 24 0. https://doi.o rg/10.1007/978- 3-319- 46466-4_14. 9 Such F.P, Sah S, Dominguez M, Pillai S, et al (2017) Robust spatial filtering with graph convolutiona l neural network s. IEEE Journal of Selected Topics in Signal Processing. https://doi.org /10.1109/JSTSP.2017.2726 981. Xu J, Durrett G (2018) Sph erical Latent Spac es for Stable Variational Autoencode rs. arxiv:1808.10805. Fisher R.S (1953) Dispersio n on a sphere. Proc. R. Soc. Lond. Ser. A Math. Phy s. Sci. 1953, 217, 295 – 305. Mardia K .V , El -Atoum S.A.M (1976) Bayesian inference for t he von m ises- fisher distribution . Biometrika, 63(1):203 – 206. https ://doi.org /10.2307/2335106. Larsen A.B.L, Sønderby S.K, Larochel le H, W inther O (2015) Autoencoding beyond pixels u sing a learned similarity m etric. ArXiv: 1512.09300. Wu J , Zhang C, X ue T , Freeman B, Tenenbaum J (2016) L earning a pr obabili stic latent space of object shapes via 3D generati ve- adversarial modeling. In: Advances in Neural Information Processing Systems. pp. 82 – 90, NI PS 2016. Le -Huu D, Paragios N (2017) Alternating direction g raph matching. IEEE CVPR, pp. 6253-6261, arXiv:1611.07583v 4. Yan J , Li C, Li Y, Cao G (2018) Adapt ive discrete hyperg raph m at ching. IEEE Trans. Cybern., vol. 48, no. 2, pp. 765 – 779, Feb. ht tps://doi.org /10.1109/TCYB.20 17.2655538. Yu J.G, X ia G.S, Samal A, Tian, J (2016) G lobally consistent correspondence of m ultiple feature se ts using proximal gausseid el relaxation. Pattern Recognition 51, 255 – 267. https://doi.org /10.1016/j.patcog.2015.09. 029. Wang F, Xia G, Xue N, Zhang Y, Pelillo M (2019) A Funct ional Represent ation for Grap h matching. arXiv:1901.05179. Wang F., Xue N., Zhang Y ., Bai X., X ia GS. (2018) A daptively Transforming Graph Match ing. Computer Vision – ECCV. ar Xiv:1807.10160v 1. Diederik P. King ma, Jimmy Ba (2017) Ad am: A method fo r stochastic optim i zation. arXiv:1412.6980v 9. Lan S, Holbrook A, Elias G.A, Fortin N, Ombao H, Shahbaba B (2019) Flexible Bayesian Dynamic Modeling of Covar iance and Correla tion Matrices. http s://arxiv.org/abs/1 711.02869v 5. Seiler C, Holm es S (2017) Multivariate Heterosceda sticity Models for Functional Brain Connectivity. Frontiers in Neu roscience, https://doi.o rg/10.3389/fnins.2017.00 696.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment