Accurate Congenital Heart Disease Model Generation for 3D Printing
3D printing has been widely adopted for clinical decision making and interventional planning of Congenital heart disease (CHD), while whole heart and great vessel segmentation is the most significant but time-consuming step in the model generation for 3D printing. While various automatic whole heart and great vessel segmentation frameworks have been developed in the literature, they are ineffective when applied to medical images in CHD, which have significant variations in heart structure and great vessel connections. To address the challenge, we leverage the power of deep learning in processing regular structures and that of graph algorithms in dealing with large variations and propose a framework that combines both for whole heart and great vessel segmentation in CHD. Particularly, we first use deep learning to segment the four chambers and myocardium followed by the blood pool, where variations are usually small. We then extract the connection information and apply graph matching to determine the categories of all the vessels. Experimental results using 683D CT images covering 14 types of CHD show that our method can increase Dice score by 11.9% on average compared with the state-of-the-art whole heart and great vessel segmentation method in normal anatomy. The segmentation results are also printed out using 3D printers for validation.
💡 Research Summary
The paper addresses a critical bottleneck in the workflow for 3D‑printed models of congenital heart disease (CHD): the labor‑intensive segmentation of the whole heart and great vessels from CT scans. Existing automatic segmentation methods, which have been tuned on normal cardiac anatomy, perform poorly on CHD cases because the disease often produces dramatic variations in chamber size, vessel topology, and the connections between the pulmonary artery (PA) and aorta (Ao). To overcome this, the authors propose a hybrid framework that combines deep learning for locally regular structures with graph‑matching algorithms for globally variable vascular topology.
The pipeline consists of three main stages. First, a 3‑D U‑Net (input 64³ voxels) segments the four cardiac chambers (LV, RV, LA, RA) and the myocardium. These structures are relatively invariant across CHD types, allowing the network to achieve high accuracy. Second, a 2‑D U‑Net processes each axial slice (512 × 512) to segment the blood pool and, importantly, a dedicated “blood‑pool boundary” class. This extra class facilitates the extraction of clear vessel edges, which are later used to build a connectivity graph. Third, the authors construct a graph where nodes correspond to vessel segments and branch points, and edges encode adjacency and geometric attributes (length, orientation). By matching this graph against a template graph representing normal cardiac anatomy, the algorithm assigns anatomical labels (e.g., aorta, pulmonary artery) to each vessel, even when the underlying topology deviates from the norm.
The authors assembled a novel CHD dataset comprising 68 contrast‑enhanced CT volumes (512 × 512 × 130‑340 voxels, voxel size ≈ 0.25 × 0.25 × 0.5 mm³). The cohort covers 14 CHD sub‑types, including common defects such as atrial septal defect (ASD) and ventricular septal defect (VSD), as well as rarer, highly anomalous forms like pulmonary atresia (PuA) and common arterial trunk (CAT). Ground‑truth annotations for seven structures (LV, RV, LA, RA, myocardium, Ao, PA) were created by experienced radiologists, each taking 1–1.5 h per case.
Training employed a combination of Dice loss and cross‑entropy loss, with data augmentation and learning‑rate scheduling identical to the baseline method Seg‑CNN (the current state‑of‑the‑art for normal anatomy). Four‑fold cross‑validation was performed, ensuring that each test fold contained a representative mix of CHD morphologies.
Quantitative results show a substantial improvement over Seg‑CNN. Across all 14 CHD types, the proposed method achieved mean Dice scores ranging from 78.5 % to 83.1 %, compared with 66.4 %–71.4 % for Seg‑CNN—a relative gain of 5.5 % to 16.1 % (average 11.9 %). The largest gain was observed for transposition of the great arteries (TGA), where both Ao and PA retain near‑normal geometry, while the smallest gain occurred for PuA, reflecting the extreme topological disruption in that condition. Standard deviations were comparable between the two methods, indicating that the improvement is consistent rather than driven by outliers.
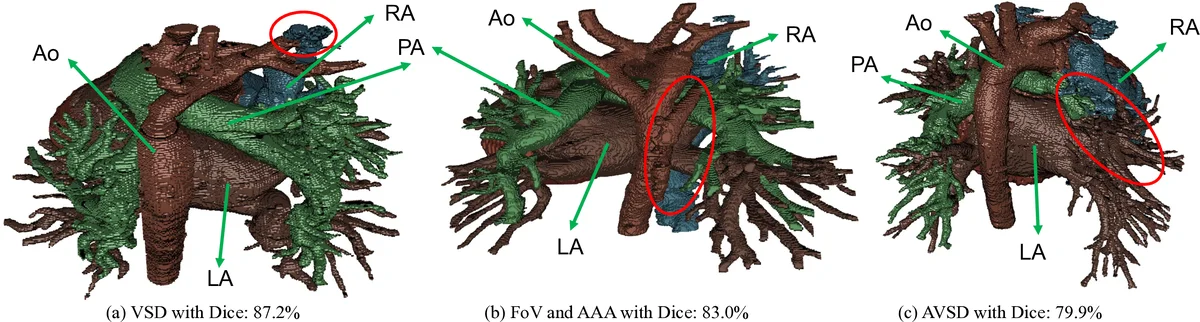
Qualitative visualizations (Fig. 4) illustrate that Seg‑CNN frequently mislabels the aorta as pulmonary artery when the PA is absent or severely altered, whereas the graph‑matching step correctly respects the connectivity constraints. The authors also printed selected segmentations on a commercial Sailner J501Pro printer. The printed models faithfully reproduced chamber volumes and vessel trajectories; only minor manual adjustments (e.g., adding thin coronary branches) were needed. Radiologists confirmed that the printed models were suitable for pre‑operative planning and educational purposes.
The study contributes three key advances: (1) a two‑stage deep‑learning‑plus‑graph‑matching pipeline tailored to the high variability of CHD anatomy; (2) a publicly released CHD CT dataset with comprehensive annotations covering a wide spectrum of defects; (3) empirical evidence that the approach not only outperforms the best existing method on Dice metrics but also yields clinically usable 3D‑printed models.
Limitations include reliance on accurate blood‑pool boundary detection; errors in this step can propagate to the graph construction and cause mislabeling. The current graph‑matching uses a fixed normal‑anatomy template, which may be insufficient for extremely rare or mixed anomalies. Moreover, the 2‑D blood‑pool segmentation does not exploit full 3‑D context, potentially limiting boundary precision.
Future work could explore more flexible graph representations (e.g., probabilistic or learned templates), integrate vessel‑specific shape priors, and replace the 2‑D U‑Net with a 3‑D network that jointly predicts blood‑pool and its boundaries. Extending the framework to incorporate hemodynamic simulation data would enable functional as well as geometric modeling, further enhancing its value for surgical planning and outcome prediction.
In summary, by intelligently combining deep learning for locally consistent structures with graph‑theoretic reasoning for globally variable vasculature, the authors deliver a robust, automated solution for whole‑heart and great‑vessel segmentation in CHD, paving the way for faster, more reliable 3D‑printed cardiac models in clinical practice.
Comments & Academic Discussion
Loading comments...
Leave a Comment