GANPOP: Generative Adversarial Network Prediction of Optical Properties from Single Snapshot Wide-field Images
We present a deep learning framework for wide-field, content-aware estimation of absorption and scattering coefficients of tissues, called Generative Adversarial Network Prediction of Optical Properties (GANPOP). Spatial frequency domain imaging is u…
Authors: Mason T. Chen, Faisal Mahmood, Jordan A. Sweer
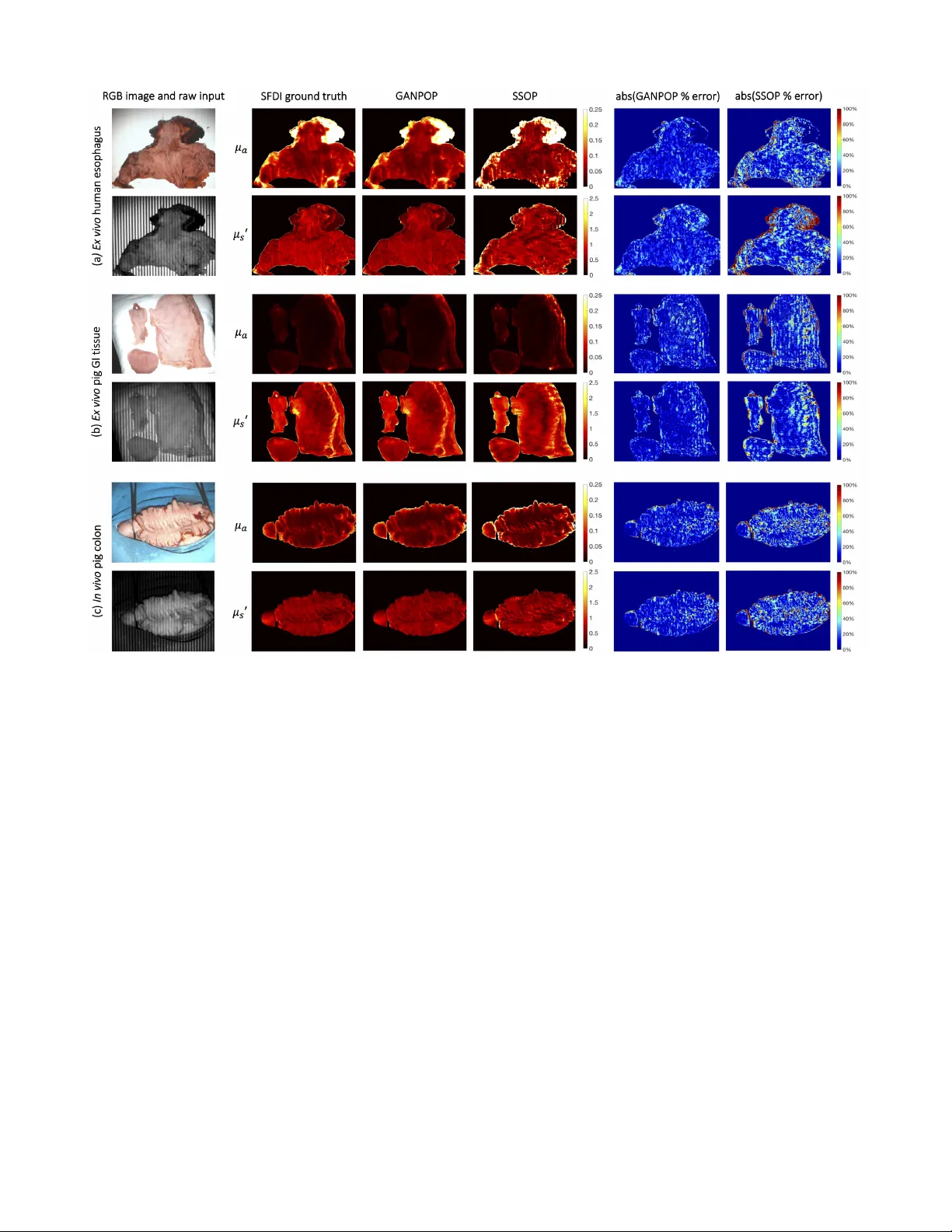
1 GANPOP: Generati v e Adversarial Netw ork Prediction of Optical Properties from Single Snapshot W ide-field Images Mason T . Chen, Faisal Mahmood, Jordan A. Sweer , and Nicholas J. Durr Abstract —W e present a deep learning framework for wide- field, content-aware estimation of absorption and scattering coef- ficients of tissues, called Generative Adversarial Network Predic- tion of Optical Properties (GANPOP). Spatial frequency domain imaging is used to obtain gr ound-truth optical properties from in vivo human hands, freshly resected human esophagectomy samples and homogeneous tissue phantoms. Images of objects with either flat-field or structur ed illumination are paired with register ed optical pr operty maps and ar e used to train conditional generative adversarial networks that estimate optical properties from a single input image. W e benchmark this approach by com- paring GANPOP to a single-snapshot optical property (SSOP) technique, using a normalized mean absolute error (NMAE) metric. In human gastrointestinal specimens, GANPOP estimates both reduced scattering and absorption coefficients at 660 nm from a single 0.2 mm -1 spatial fr equency illumination image with 58% higher accuracy than SSOP . When applied to both in vivo and ex vivo swine tissues, a GANPOP model trained solely on human specimens and phantoms estimates optical properties with approximately 43% improvement over SSOP , indicating adaptability to sample variety . Moreo ver , we demonstrate that GANPOP estimates optical properties fr om flat-field illumination images with similar error to SSOP , which requires structured- illumination. Given a training set that appr opriately spans the target domain, GANPOP has the potential to enable rapid and accurate wide-field measur ements of optical pr operties, ev en from con ventional imaging systems with flat-field illumination. Index T erms —optical imaging, tissue optical properties, neural networks, machine learning, spatial frequency domain imaging I . I N T R O D U C T I O N T HE optical properties of tissues, including the absorption ( µ a ) and reduced scattering ( µ 0 s ) coefficients, can be useful clinical biomarkers for measuring trends and detecting abnormalities in tissue metabolism, tissue oxygenation, and cellular proliferation [1]–[5]. Optical properties can also be used for contrast in functional or structural imaging [6], [7]. Thus, quantitativ e imaging of tissue optical properties can facilitate more objective, precise, and optimized management of patients. T o measure optical properties, it is generally necessary to decouple the effects of scattering and absorption, which both influence the measured intensity of remitted light. Separation of these parameters can be achie ved with temporally or spatially resolved techniques, which can each be performed with measurements in the real or frequency domains. Spatial Frequency Domain Imaging (SFDI) decouples absorption from All authors are with the Department of Biomedical Engineering, Johns Hopkins University , Baltimore, MD, 21218. Contact e-mail: ndurr@jhu.edu. Fig. 1. Proposed conditional Generative Adversarial Network (cGAN) ar- chitecture. The generator is a combination of ResNet and U-Net and is trained to produce optical property maps that closely resemble SFDI output. The discriminator is a three-layer classifier that operates on a patch level and is trained to classify the output of the generator as ground truth (real) or generated (fake). The discriminator is updated using a history of 64 previously-generated image pairs. scattering by characterizing the tissue modulation transfer function to spatially modulated light [8], [9]. This approach has significant advantages in that it can easily be implemented with a consumer grade camera and projector , and achieve rapid, non-contact mapping of optical properties. These ad- vantages make SFDI well-suited for applications that benefit from wide-field characterization of tissues, such as image- guided surgery [10], [11] and wound characterization [12]– [14]. Additionally , recent work has explored the use of SFDI for improving endoscopic procedures [15], [16]. Although SFDI is finding a growing number of clinical applications, there are remaining technical challenges that limit its adoption. First, SFDI requires structured light projection with carefully-controlled working distance and calibration, which is especially challenging in an endoscopic setting. Second, it is dif ficult to achie ve real-time measurements. Con ventional SFDI requires a minimum of six images per wa velength (three distinct spatial phases at two spatial frequen- cies) to generate a single optical property map. A lookup table (LUT) search is then performed for optical property fitting. The recent development of real-time single snapshot imaging of optical properties (SSOP) has reduced the number of images required per wa velength from 6 to 1, considerably shorten- 2 ing acquisition time [17]. Howe ver , SSOP introduces image artifacts arising from single-phase projection and frequency filtering, which corrupt the optical property estimations. T o reduce barriers to clinical translation, there is a need for optical property mapping approaches that are simultaneously fast and accurate while requiring minimal modifications to con ventional camera systems. Here, we introduce a deep learning framew ork to predict optical properties directly from single images. Deep networks, especially con volutional neural networks (CNNs), are growing in popularity for medical imaging tasks, including computer- aided detection, segmentation, and image analysis [18]–[20]. W e pose the optical property estimation challenge as an image- to-image translation task and employ generativ e adversarial networks (GANs) to efficiently learn a transformation that is robust to input v ariety . First proposed in [21], GANs ha ve improv ed upon the performance of CNNs in image generation by including both a generator and a discriminator . The former is trained to produce realistic output, while the latter is tasked to classify generator output as real or fake. The two components are trained simultaneously to outperform each other , and the discriminator is discarded once the generator has been trained. When both components observe the same type of data, such as text labels or input images, the GAN model becomes conditional. Conditional GANs (cGANs) are capable of making structured predictions by incorporating non-local, high-lev el information. Moreover , because they can automatically learn a loss function instead of using a hand- crafted one, cGANs have the potential to be an effecti ve and generalizable solution to various image-to-image translation tasks [22], [23]. In medical imaging, cGANs have been proven successful in many applications, such as image synthesis [24], noise reduction [25], and sparse reconstruction [26]. In this study , we train cGAN networks on a series of structured or flat- field illumination images paired with corresponding optical property maps (Fig. 1). W e demonstrate that the GANPOP approach produces rapid and accurate estimation from input images from a wide variety of tissues using a relatively small set of training data. I I . R E L AT E D W O R K A. Diffuse reflectance imaging Optical absorption and reduced scattering coef ficients can be measured using temporally or spatially resolved diffuse re- flectance imaging. Approaches that rely on point illumination inherently have a limited field of view [27], [28]. Non-contact, hyperspectral imaging techniques measure the attenuation of light at different wav elengths, from which the concentrations of tissue chromophores, such as oxy- and deoxy-hemoglobin, water , and lipids, can be quantified [29]. A recent study has also proposed using a Bayesian framew ork to infer tissue oxygen concentration by recov ering intrinsic multispectral measurements from RGB images [30]. Howe ver , these meth- ods fail to unambiguously separate absorption and scattering coefficients, which poses a challenge for precise chromophore measurements. Moreov er , accurate determination of both pa- rameters is critical for the detection and diagnosis of diseases [1], [5]. B. Single snapshot imaging of optical pr operties SSOP achiev es optical property mapping from a single structured light image. Using Fourier domain filtering, this method separates DC (planar) and AC (spatially modulated) components from a single-phase structured illumination image [17]. A grid pattern can also be applied to simultaneously ex- tract optical properties and three-dimensional profile measure- ments [31]. When tested on homogeneous tissue-mimicking phantoms, this method is able to recov er optical properties within 12% for absorption and 6% for reduced scattering using con ventional profilometry-corrected SFDI as ground truth. C. Machine learning in optical pr operty estimation Despite its prev alence and increasing importance in the field of medical imaging, machine learning has only recently been explored for optical property mapping. This includes a random forest re gressor to replace the nonlinear model in version [32], and using deep neural networks to reconstruct optical properties from multifrequency measurements [33]. Both of these approaches aim to bypass the time-consuming LUT step in SFDI. Howe ver , they require dif fuse reflectance measurements from multiple images to achie ve accurate results and consider each pixel independently . I I I . C O N T R I B U T I O N S W e propose an adv ersarial framew ork for learning a content- aware transformation from single illumination images to opti- cal property maps. In this work, we: 1) dev elop a data-driv en model to estimate optical properties directly from input reflectance images; 2) demonstrate adv antages of structured versus flat-field light as an input to determine optical properties; 3) perform cross-validated experiments, comparing our tech- nique with model-based SSOP and other deep learning- based methods; and 4) acquire and make publicly-av ailable a dataset of registered flat-field-illumination images, structured- illumination images, and ground-truth optical properties of a variety of ex vivo and in vivo tissues. I V . M E T H O D S For training and testing of the GANPOP model, single structured or flat-field illumination images were used, paired with re gistered optical property maps. T o generate ground truth optical properties, con ventional six-image SFDI was imple- mented. GANPOP performance was analyzed and compared to other techniques both in unseen tissues of the same type as the training data (new e x vivo esophagus) and in different tissue types ( in vivo and ex vivo swine gastrointestinal tissues). A. Har dwar e In this study , all images were captured using a commercially av ailable SFDI system (Reflect RS TM , Modulated Imaging Inc.). A schematic of the system is sho wn in Fig. 2. Cross polarizers were utilized to reduce the effect of specular re- flections, and images were acquired in a custom-built light 3 enclosure to minimize ambient light. Ra w images, after 2x2 pixel hardware binning, were 520 × 696 pixels, with a pixels size of 0.278 mm in the object space. Fig. 2. Overvie w of conv entional SFDI illumination patterns, hardware, and processing flow . SFDI captures six frames (three phase offsets at two different spatial frequencies) to generate an absorption and reduced scattering map. DC stands for planar illumination images and AC stands for spatially modulated images. T o calculate optical properties, acquired data is demodulated, cali- brated against a reference phantom, and in verted using a lookup table. B. SFDI gr ound truth optical pr operties Ground truth optical property maps were generated using con ventional SFDI with 660 nm light following the method from Cuccia et al. [9]. First, images of a calibration phan- tom with homogeneous optical properties and the tissue of interest are captured under spatially modulated light. W e used a flat polydimethylsiloxane-titanium dioxide (PDMS-T iO 2 ) phantom with reduced scattering coef ficient of 0.957 mm -1 and absorption coef ficient of 0.0239 mm -1 at 660 nm. W e project spatial frequencies of 0 mm -1 (DC) and 0.2 mm -1 (A C), each at three different phase offsets ( 0 , 2 3 π , and 4 3 π ) for this study . A C images are demodulated at each pixel x using: M AC ( x ) = √ 2 3 · s ( I 1 ( x ) − I 2 ( x )) 2 + ( I 2 ( x ) − I 3 ( x )) 2 + ( I 3 ( x ) − I 1 ( x )) 2 , (1) where I 1 , I 2 , and I 3 represent images at the three phase offsets. The spatially varying DC amplitude is calculated as the av erage of the three DC images. Diffuse reflectance at each pixel x is then computed as: R d ( x ) = M AC ( x ) M AC,r ef ( x ) · R d,predicted . (2) Here, M AC,r ef denotes the demodulated A C amplitude of the reference phantom, and R d,predicted is the dif fuse re- flectance predicted by Monte Carlo models. Where indicated, we corrected for height and surface angle variation of each pixel from depth maps measured via profilometry . Profilometry measurements were obtained by projecting a spatial frequency of 0.15 mm -1 and calculating depth at each pixel [34]. Finally , µ a and µ 0 s are estimated by fitting R d, 0 mm − 1 and R d, 0 . 2 mm − 1 into an LUT previously created using Monte Carlo simulations [35]. C. Single Snapshot Optical Properties (SSOP) SSOP was implemented as the model-based alternativ e of GANPOP . This method separates DC and AC components from a single-phase structured-light image by frequency fil- tering with a 2D band-stop filter and a high-pass filter [17]. Both filters are rectangular windows that isolate the frequency range of interest while preserving high-frequency content of the image. In this study , cutoff frequencies f DC = [0.16 mm -1 , 0.24 mm -1 ] and f AC = [0, 0.16 mm -1 ] were selected [31]. M DC can subsequently be recovered through a 2D in verse Fourier transform, and the A C component is obtained using an additional Hilbert transform. Fig. 3. Detailed architecture of the proposed generator . W e use a fusion network that combines properties from ResNet and U-Net, including both short and long skip connections in the form of feature addition. Each residual block contains five con volution layers, with short skips between the first and the fifth layer . D. GANPOP Ar chitectur e The GANPOP architecture is based on an adversarial training framework. When used in a conditional GAN-based image-to-image translation setup, this framework has the abil- ity to learn a loss function while a voiding the uncertainty inherent in using hand-crafted loss functions [23], [36]. The generator is tasked with predicting pixel-wise optical proper- ties from SFDI images while the discriminator classifies pairs of SFDI images and optical property maps as being real or fake (Fig. 1). The discriminator additionally gi ves feedback to the generator over the course of training. The generator employs a modified U-Net consisting of an encoder and a decoder with skip connections [37]. Howe ver , unlike the original U-Net, the GANPOP network includes properties of a ResNet, including short skip connections within each lev el [38] (Fig. 3). Each residual block is a 3-layer b uilding block with an additional con volutional layer on both sides. This ensures that the number 4 of input features matches that of the residual block and that the network is symmetric [39]. Moreover , GANPOP generator replaces the U-Net concatenation step with feature addition, making it a fully residual network. Using n as the total number of layers in the encoder-decoder network and i as the current layer , long skip connections are added between the i th and the ( n − i ) th layer in order to sum features from the two lev els. After the last layer in the decoder, a final con volution is applied to shrink the number of output channels and is followed by a T anh function. Regular ReLUs are used for the decoder and leaky ReLUs (slope = 0.2) for the encoder . W e chose a receptiv e field of 70 × 70 pixels for our discriminator because this window captures two periods of AC illumination in each direction. This discriminator is a three-layer classifier with leaky ReLUs (slope = 0.2), as discussed in [23]. The dis- criminator makes classification decisions based on the current batch as well as a batch randomly sampled from 64 previously generated image pairs. Both networks are trained iteratively and the training process is stabilized by incorporating spectral normalization in both the generator and the discriminator [40]. The conditional GAN objective for generating optical property maps from input images ( G : X → Y ) is: L GAN ( G, D ) = E x,y ∼ p data ( x,y ) [( D ( x, y ) − 1) 2 ] + E x ∼ p data ( x ) [ D ( x, G ( x )) 2 ] , (3) where G is the generator , D the discriminator , and p data is the optimal distribution of the data. W e empirically found that a least squares GAN (LSGAN) objectiv e [41] produced slightly better performance in predicting optical properties than a traditional GAN objectiv e [21], and so we utilize LSGAN in the networks presented here. An additional L 1 loss term was added to the GAN loss to further minimize the distance from the ground truth distribution and stabilize adversarial training: L 1 ( S ) = E x,y ∼ p data ( x, y )[ || y − G ( x ) || 1 ] . (4) The full objective can be expressed as: L ( G, D ) = L GAN ( G, D ) + λ L 1 ( G ) , (5) where λ is the regularization parameter of the L 1 loss term. This optimization problem was solved using an Adam solver with a batch size of 1 [42]. The training code w as implemented using Pytorch 1.0 on Ubuntu 16.04 with Google Cloud. For all experiments, λ was set to 60. A total of 200 epochs was used with a learning rate of 0.0002 for half of the epochs and the learning rate was linearly decayed for the remaining half. Both networks were initialized from a Gaussian distribution with a mean and standard deviation of 0 and 0.02, respectiv ely . Con ventional neural networks typically operate on three- channel (or RGB) images as input and output, with each channel representing red (R), green (G), or blue (B). In this study , four separate networks (N 1 to N 4 ) were trained for image-to-image translation with a v ariety of input and output parameters, summarized in T able I. For input, I AC and I DC represent single-phase raw images at 0.2 mm -1 and 0 spatial frequency , respectiv ely . M DC ,ref and M AC,r ef are the demodulated DC and AC amplitude of T ABLE I S U MM A RY O F NE T W O RK S TR A I N ED I N T H I S S T UDY . Input Output N i R channel G channel R channel G channel N 1 I AC M DC,r ef I AC M AC,ref µ a µ 0 s N 2 I AC M DC,r ef I AC M AC,ref µ a,prof µ 0 s,prof N 3 I DC M DC,r ef I DC M AC,ref µ a µ 0 s N 4 I DC M DC,r ef I DC M AC,ref µ a,prof µ 0 s,prof Fig. 4. Example input-output pair used in N 1 showing each individual channel as well as the combined RGB images. R and G channel of the output contain µ a and µ 0 s , respectively . Thus, a high absorption appears red while a high scattering appears green. the calibration phantom. Blue channels are left as zeros in all cases. It is important to note that M AC,r ef and M DC ,ref are measured only once during calibration before the imaging experiment and thus do not add to the total acquisition time. The purpose of these two terms is to account for drift of the system over time and correct for non-uniform illumination, making the patch used in the network origin-independent. These two calibration images are also required by traditional SFDI and the SSOP approaches. A network without calibration was empirically trained, and it produced 230% and 58% larger error than with calibration in absorption and scattering coefficients, respectively . A single output image contains both µ a and µ 0 s in different channels. T wo dedicated networks were empirically trained for estimating µ a and µ 0 s independently , but no accuracy benefits were observed. Optical property maps calculated by non-profile-corrected SFDI were used as ground truth for N 1 and N 3 . W e also assessed the ability of GANPOP to learn both optical property estimation and sample height and surface normal correction by training and testing with profilometry-corrected data ( N 2 and N 4 ). All optical property maps for training and testing were nor- malized to have a consistent representation in the 8-bit images commonly used in CNNs [43]. W e defined the maximum value of 255 to be 0.25 mm -1 for µ a and 2.5 mm -1 for µ 0 s . 5 Additionally , each image of size 520 × 696 was segmented at a random stride size into multiple patches of 256 × 256 pixels and paired with a registered optical property patch for training, as shown in Fig. 4. E. T issue Samples 1) Ex vivo human esophagus: Eight ex vivo human esophagectomy samples were imaged at Johns Hopkins Hos- pital for training and testing of our networks. All patients were diagnosed with esophageal adenocarcinoma and were scheduled for an esophagectomy . The research protocol was approv ed by the Johns Hopkins Institutional Re vie w Board and consents were acquired from all patients prior to each study . All samples were handled by a trained pathologist and imaged within one hour after resection [44]. Example raw images of a specimen captured by the SFDI system are sho wn in Fig. 2, 4, and 11(a). All samples consisted of the distal esophagus, the gastroesophageal junction, and the proximal stomach. The samples contain complex topography and a relativ ely wide range of optical properties (0.02-0.15 mm -1 for µ a and 0.1-1.5 mm -1 for µ 0 s at λ = 660 nm), making it suitable for training a generalizable model that can be applied to other tissues with non-uniform surface profiles. An illumination wav elength of 660 nm was chosen because it is close to the optimal wav elength for accurate tissue oxygenation measurements [45]. In this study , six ex vivo human esophagus samples were used for training of the GANPOP model and two used for testing. A leav e-two-out cross v alidation method was imple- mented, resulting in four iterations of training for each net- work. Performance results reported here are from an average of these four iterations. 2) Homogeneous phantoms: The four GANPOP networks were also trained on a set of tissue-mimicking silicone phan- toms made from PDMS-TiO 2 (P4, Eager Plastics Inc.) mixed with India ink as absorbing agent [46]. T o ensure homoge- neous optical properties, the mixture was thoroughly combined and poured into a flat mold for curing. In total, 18 phantoms with unique combinations of µ a and µ 0 s were fabricated, and their optical properties are summarized in Fig. 5. In this study , six tissue-mimicking phantoms were used for training and twelv e for testing. W e intentionally selected phantoms for training that had optical properties not spanned by the esophagus training samples (highlighted by green ellipses in Fig. 6), in order to develop GANPOP networks capable of estimating the widest range of optical properties. 3) In vivo samples: T o provide the network with in vivo samples that were perfused and oxygenated, se ven human hands with different le vels of pigmentation (Fitzpatrick skin types 1-6) were imaged with SFDI. T wo were used for training and fi ve for testing. 4) Swine tissue: Four specimens of upper gastrointestinal tracts that included stomach and esophagus were harvested from four different pigs for ex vivo imaging with SFDI. Optical properties of these samples are summarized in Fig. 7. Additionally , we imaged a pig colon in vivo during a surgery . The li ve study was performed with approv al from Fig. 5. V alidation of SSOP model-based prediction of optical properties with 18 homogeneous tissue phantoms. SSOP prediction from a single input image demonstrates close agreement with SFDI prediction from six input images. Johns Hopkins Univ ersity Animal Care and Use Committee (A CUC). All swine tissue images were used exclusiv ely for testing optical property prediction. F . P erformance Metric Normalized Mean Absolute Error (NMAE) was used to ev aluate the performance of different methods, which was calculated using: N M AE = P T i =1 | p i − p i,ref | P T i =1 p i,ref . (6) p i and p i,ref are pixel values of predicted and ground- truth data, and T is the total number of pixels. The metric was calculated using SFDI output as ground truth. A smaller NMAE v alue indicates better performance. V . R E S U L T S A. SSOP validation For benchmarking, SSOP was implemented as a model- based counterpart of GANPOP . For independent validation, we applied SSOP to 18 homogeneous tissue phantoms (Fig. 5). Each v alue was calculated as the mean of a 100 × 100-pixel region of interest (R OI) from the center of the phantom, with error bars sho wing standard deviations. SSOP demonstrates high accuracy in predicting optical properties of the phantoms, with an average percentage error of 2.35% for absorption and 2.69% for reduced scattering. 6 Fig. 6. Scatter plot showing optical property pairs estimated by GANPOP compared to ground truth from conv entional SFDI on 12 tissue phantoms. The 2D histogram in the background illustrates the distribution of training pixels among all optical property pairs, determined by SFDI. Green ellipses indicate dense pixel counts due to homogeneous phantoms used in training. T esting samples in the red box fell outside of the training range but were accurately predicted by GANPOP . B. GANPOP test in homogeneous phantoms Phantom optical properties predicted by N 1 are plotted with ground truth in Fig. 6. Each optical property reported is the average v alue of a 100 × 100 R OI of a homogeneous phantom, with error bars sho wing standard deviations. On av erage, GANPOP produced 3.06% error for absorption and 1.26% for scattering. The scatter plot in Fig. 6 is overlaid on a 2D histogram of pix el counts for each ( µ a , µ 0 s ) pair used in an example training iteration. Green ellipses indicate training samples from homogeneous phantoms. The three testing results enclosed by red boxes ha ve optical properties outside of the range spanned by the training data but were still reasonably estimated by the GANPOP network. C. GANPOP test on ex vivo human esophagus GANPOP and SSOP were tested on the ex vivo human esophagus samples. NMAE scores were calculated for the two testing samples from each of four-fold cross validation iterations, and the av erage values from the four networks tested on a total of eight samples are reported in Fig. 8. Results from N 2 , N 4 , and SSOP are also compared to profilometry- corrected ground truth and shown in the same bar chart. On av erage, GANPOP produced approximately 58% higher accuracy with A C input than SSOP . Example optical property maps of a testing sample generated by N 1 are shown in Fig. 11(a). D. GANPOP test on ex vivo pig samples Each of the four GANPOP networks were tested on ex vivo esophagus and stomach samples from four pigs. A verage NMAE scores for GANPOP and SSOP method were calcu- lated for all eight pig tissue specimens (four esophagi and four Fig. 7. Histogram of optical property distribution of testing pixels from pig samples. Compared to training samples, pig tissues tested in this study had, on average, lower absorption coefficients and higher scattering coefficients. Fig. 8. Accuracy of GANPOP in predicting optical properties of ex vivo hu- man esophagus samples. A verage NMAE for absorption (green) and scattering coefficients (blue) are reported for the SSOP technique as well as GANPOP networks. SFDI was used for ground truth. OP and OP’ stand for profile- corrected and uncorrected optical properties, respectively . stomachs) and are summarized in Fig. 9. Background regions, which were absorbing paper , were manually masked in the calculation, and the reported scores are the av erage v alues of 779,101 tissue pixels. The optical properties of the pig samples are also shown in a 2D histogram in Fig. 7. Despite the fact that some testing samples had optical properties not co vered by the training set, GANPOP outperforms SSOP in terms of av erage accuracy and qualitative image quality (Fig. 11). E. GANPOP test on in vivo pig colon The networks were additionally tested on an in vivo pig colon. A verage NMAE scores for GANPOP and SSOP are reported in Fig. 10 as av erage v alues of 118,594 pixels. The generated maps are shown in Fig. 11(c). The proposed technique produces more accurate results than SSOP when compared to both uncorrected and profile-corrected ground truth data. 7 T ABLE II P E RF O R M AN C E C O M P A R IS O N O F T HE P RO PO S E D F R A ME W O R K AG A I NS T M OD E L - B A SE D S SO P AN D OT H ER D EE P LE A R N IN G A RC H I TE C T U RE S WH E N T E ST E D O N PR O FIL E - U NC O R RE C T E D DAT A ( N 1 ) . P E R FO R M A NC E I S M E AS U R E D I N T E R MS O F N O R MA L I Z ED M EA N AB S O L UT E E RR OR ( NM A E ) . Data type ResNet UNet ResNet-UNet ResNet GAN UNet GAN SSOP Proposed µ a µ 0 s µ a µ 0 s µ a µ 0 s µ a µ 0 s µ a µ 0 s µ a µ 0 s µ a µ 0 s Human esophagus 0.227 0.143 0.161 0.153 0.203 0.140 0.232 0.156 0.176 0.165 0.301 0.290 0.124 0.121 In vivo pig colon 0.614 0.729 0.320 0.486 0.609 0.583 0.795 0.769 0.335 0.377 0.246 0.235 0.139 0.131 Ex vivo pig GI tissue 2.954 0.175 0.344 0.378 2.842 0.177 3.138 0.175 0.574 0.410 0.152 0.106 0.080 0.068 In vivo human hands 0.373 0.100 0.123 0.109 0.249 0.081 0.353 0.106 0.162 0.099 0.092 0.058 0.075 0.055 Overall 1.042 0.287 0.237 0.281 0.976 0.245 1.129 0.301 0.312 0.263 0.198 0.172 0.104 0.094 Fig. 9. Plots of av erage NMAE for absorption (green) and scattering coefficients (blue) of ex vivo pig stomach and esophagus samples. Metrics are calculated against SFDI ground truth. OP and OP’ stand for profile-corrected and uncorrected optical properties, respectiv ely . Fig. 10. Plots of av erage NMAE for absorption (green) and scattering coefficients (blue) of the in vivo pig colon. Metrics are calculated against SFDI ground truth. OP and OP’ stand for profile-corrected and uncorrected optical properties, respectiv ely . F . Comparative analysis of existing deep networks Sev eral deep learning architectures were explored for the purpose of optical property mapping, including con ventional U-Net [37] and ResNet [38], both stand-alone and integrated in a cGAN framework [23], [39]. The NMAE performance of each architecture was compared to GANPOP . All the networks were four-fold cross v alidated, and the testing dataset included eight e x vivo human esophagi, four e x vivo pig GI samples, one in vivo pig colon, and five in vivo hands (T able II). V I . D I S C U S S I O N In this study , we have described a GAN-based technique for end-to-end optical property mapping from single structured and flat-field illumination images. Compared to the original pix2pix paradigm [23], the generator of our model adopted a fusion of U-Net and ResNet architectures for several reasons. First, a fully residual network effecti vely resolved the issue of vanishing gradients, allowing us to stably train a relativ ely deep neural network [39]. Second, the use of both long and short skip connections enables the network to learn from the structure of the images while preserving both low and high frequency details. The information flow both within and between levels is important for the prediction of optical properties, as demonstrated by the improved performance over a U-Net or ResNet approach. Moreover , as shown in T able II, the inclusion of a discriminator significantly improved the performance of the fusion generator . This was especially apparent in the case for pig data, likely due to this testing tissue differing considerably from the training samples. W e hypothesize that the cGAN architecture enforced the similarity between generated images and ground truth while preventing the generator from depending too much on the context of the image. Overall, the GANPOP method outperformed the other deep networks by a significant margin on all data types (T able II). Additionally , we empirically found that a least squares GAN outperformed a conv entional GAN when trained for 200 epochs. Howe ver , as discussed in [47], this improvement could potentially be matched by a con ventional GAN with more training. Compared to phantom ground truth in Fig. 6, GANPOP esti- mated optical properties with standard deviations on the same order of magnitude as con ventional SFDI. Additionally , the GANPOP networks exhibited potential to extrapolate phantom optical properties that were not present in the training samples (highlighted by the red boxes in Fig. 6). This provides e vidence that these networks have successfully learned the relationship between diffuse reflectance and optical properties, and are able to infer beyond the range of training data. Fig. 8, 9, and 10 sho w that GANPOP with AC input consistently outperformed SSOP when tested on these types of data. From Fig. 7, it is evident that optical properties of the pig samples differed considerably from those of human esophagi used for training. Nev ertheless, GANPOP exhibited more accurate estimation than the model-based SSOP bench- mark. Moreover , a single network was trained for estimating 8 Fig. 11. Example results for AC input to non-profilometry-corrected optical properties ( N 1 ). From left to right: RGB image and raw structured illumination image, SFDI ground truth, GANPOP output, SSOP output, percent difference between GANPOP and ground truth, and percent difference between SSOP and ground truth; From top to bottom: (a) ex vivo human esophagus, (b) ex vivo pig stomach and esophagus, and (c) in vivo pig colon. both µ a and µ 0 s due to its lo wer computational cost and potential benefits in learning the relationships between the two parameters in tissues. Compared to SSOP , GANPOP optical property maps con- tain fe wer artifacts caused by frequency filtering (Fig. 11). For both GANPOP and SSOP optical property estimation, a relativ ely large error is present on the edge of the sample. This is caused by the transition between tissue and the background, which poses problems for SFDI ground truth, and would be less significant for in vivo imaging. Artifacts caused by patched input are visible in GANPOP images, which can be reduced by using a larger patch size. Ho wev er, this was not implemented in our study due to the size and the number of the specimens av ailable for training. In our benchmarking with SSOP , we implemented the first version of the technique, which does not correct for sample height and surface angle variations. Recent dev elopments have enabled these corrections by utilizing a more complex illumination pattern and additional processing steps [31]. W e implemented the original version of SSOP because it allowed comparing identical input images for both SSOP and GANPOP . In addition to training GANPOP models to estimate optical properties from objects assumed to be flat ( N 1 and N 3 ), we also trained networks that directly estimate profilometry- corrected optical properties ( N 2 and N 4 ). For the same A C input, these models generated improved results over SSOP when tested on human and pig data. Moreov er, when compared against profile-corrected ground truth, they produced 35.7% less error for µ a and 44.7% for µ 0 s than did uncorrected GANPOP results from N 1 and N 3 . This means that GANPOP is capable of inferring surface profile from a single fringe image and adjusting measured diffuse reflectance accordingly . In experiment N 3 and N 4 , when trained on DC illumination images, the GANPOP model became less accurate. Nev er- theless, these netw orks conv erged during training, and albeit less accurate, the ex vivo human results still produced a lower NMAE than SSOP . Hence, gi ven a sufficiently large training dataset, GANPOP has the potential to enable rapid and accurate wide-field measurements of optical properties from con ventional camera systems. This could be useful for applications such as endoscopic imaging of the GI tract, where the range of tissue optical properties is limited and modification of the hardware system is challenging. In terms of speed, GANPOP requires capturing one sam- 9 ple image instead of six, thus significantly shortening data acquisition time. For optical property extraction, the model dev eloped here without optimization takes approximately 0.04 s to process a 256 × 256 image on an NVIDIA T esla P100 GPU. Therefore, this technique has the potential to be applied in real time for fast and accurate optical property mapping. In terms of adaptability , random cropping ensures that our trained models work on an y 256 × 256 patches within the field of vie w . Additionally , while the models were trained on the same calibration phantom at 660 nm, they could theoretically be applied to other references or w av elengths by scaling the av erage M DC ,ref and M AC,r ef . For future work, a more generalizable model could be trained on a wider range of optical properties and imaging geometries, though this would inevitably incur a higher com- putational cost and necessitate a much larger dataset for train- ing. For e xample, all input images used here were acquired at an approximately-constant working distance. Incorporating monocular depth estimates into the prediction may enable GANPOP to account for large differences in working distance [48], [49]. This could be particularly useful for endoscopic screening where constant imaging geometries are dif ficult to achie ve. Ha ving a model trained on images at multiple wa velengths, this technique can be modified to provide critical information in real time, such as tissue oxygenation and metabolism biomarkers. Accuracy in this application may also benefit from training adversarial networks to directly estimate these biomarkers rather than using optical properties as intermediate representations. By similar extension, future research may dev elop networks to directly estimate disease diagnosis and localization from structured light images. V I I . C O N C L U S I O N W e have proposed a deep learning-based approach to optical property mapping (GANPOP) from single snapshot wide-field images. This model utilizes a conditional Generativ e Adv er- sarial Network consisting of a generator and a discriminator that are iteratively trained in concert with one another . Using SFDI-determined optical properties as ground truth, GANPOP produces significantly more accurate optical property maps than a model-based SSOP benchmark. Importantly , we ha ve demonstrated that GANPOP can estimate optical properties with con ventional flat-field illumination, potentially enabling optical property mapping in endoscopy without modifications for structured illumination. This method lays the foundation for future work in incorporating real-time, high-fidelity optical property mapping and quantitativ e biomarker imaging into endoscopy and image-guided surgery applications. A C K N O W L E D G M E N T This work was supported in part with funding from the NIH T railblazer A ward (R21 EB024700). W e would lik e to thank Dr . Darren Roblyers group at Boston Univ ersity for sharing SFDI software. R E F E R E N C E S [1] R. Richards-Kortum and E. Sevick-Muraca, “Quantitative optical spec- troscopy for tissue diagnosis, ” Annual Review of Physical Chemistry , vol. 47, no. 1, pp. 555–606, 1996. [2] R. A. Drezek, M. Guillaud, T . G. Collier, I. Boiko, A. Malpica, C. E. MacAulay , M. Follen, and R. R. Richards-Kortum, “Light scattering from cervical cells throughout neoplastic progression: influence of nuclear morphology , dna content, and chromatin texture, ” Journal of biomedical optics , vol. 8, no. 1, pp. 7–17, 2003. [3] B. W . Maloney , D. M. McClatchy , B. W . Pogue, K. D. Paulsen, W . A. W ells, and R. J. Barth, “Revie w of methods for intraoperative margin detection for breast conserving surgery , ” Journal of biomedical optics , vol. 23, no. 10, p. 100901, 2018. [4] J. R. Mourant, M. Canpolat, C. Brocker , O. Esponda-Ramos, T . M. Johnson, A. Matanock, K. Stetter, and J. P . Freyer , “Light scattering from cells: the contribution of the nucleus and the effects of proliferative status, ” Journal of biomedical optics , vol. 5, no. 2, pp. 131–138, 2000. [5] Z. A. Steelman, D. S. Ho, K. K. Chu, and A. W ax, “Light-scattering methods for tissue diagnosis, ” Optica , vol. 6, no. 4, pp. 479–489, Apr 2019. [6] A. J. Lin, M. A. Koike, K. N. Green, J. G. Kim, A. Mazhar , T . B. Rice, F . M. LaFerla, and B. J. T romberg, “Spatial frequenc y domain imaging of intrinsic optical property contrast in a mouse model of alzheimer’ s disease, ” Annals of Biomedical Engineering , vol. 39, no. 4, pp. 1349– 1357, Apr 2011. [7] N. Shah, A. Cerussi, C. Eker, J. Espinoza, J. Butler , J. Fishkin, R. Hor- nung, and B. T romberg, “Noninv asive functional optical spectroscopy of human breast tissue, ” Pr oceedings of the National Academy of Sciences , vol. 98, no. 8, pp. 4420–4425, 2001. [8] N. D ¨ ognitz and G. W agni ` eres, “Determination of tissue optical proper- ties by steady-state spatial frequency-domain reflectometry , ” Lasers in medical science , vol. 13, no. 1, pp. 55–65, 1998. [9] D. J. Cuccia, F . Bevilacqua, A. J. Durkin, F . R. A yers, and B. J. T romberg, “Quantitation and mapping of tissue optical properties using modulated imaging, ” Journal of Biomedical Optics , vol. 14, no. 2, p. 024012, Apr . 2009. [10] M. R. Pharaon, T . Scholz, S. Bogdanoff, D. Cuccia, A. J. Durkin, D. B. Hoyt, and G. R. D. Evans, “Early detection of complete vascular occlusion in a pedicle flap model using quantitative [corrected] spectral imaging, ” Plastic and Reconstructive Sur gery , vol. 126, no. 6, pp. 1924– 1935, Dec. 2010. [11] S. Gioux, A. Mazhar , B. T . Lee, S. J. Lin, A. M. T obias, D. J. Cuccia, A. Stockdale, R. Oketokoun, Y . Ashitate, E. K elly , M. W einmann, N. J. Durr , L. A. Moffitt, A. J. Durkin, B. J. T romberg, and J. V . Frangioni, “First-in-human pilot study of a spatial frequency domain oxygenation imaging system, ” Journal of Biomedical Optics , vol. 16, no. 8, 2011. [12] M. Kaiser , A. Y afi, M. Cinat, B. Choi, and A. J. Durkin, “Noninv asive assessment of burn wound sev erity using optical technology: a review of current and future modalities, ” Burns: Journal of the International Society for Burn Injuries , vol. 37, no. 3, pp. 377–386, May 2011. [13] C. W einkauf, A. Mazhar, K. V aishnav , A. A. Hamadani, D. J. Cuccia, and D. G. Armstrong, “Near-instant nonin vasiv e optical imaging of tissue perfusion for vascular assessment, ” Journal of V ascular Surg ery , vol. 69, no. 2, pp. 555 – 562, 2019. [14] A. Y afi, F . K. Muakkassa, T . Pasupneti, J. Fulton, D. J. Cuccia, A. Mazhar , K. N. Blasiole, and E. N. Mostow , “Quantitativ e skin assessment using spatial frequency domain imaging (sfdi) in patients with or at high risk for pressure ulcers, ” Lasers in Surgery and Medicine , vol. 49, no. 9, pp. 827–834, 2017. [15] J. P . Angelo, M. van de Giessen, and S. Gioux, “Real-time endoscopic optical properties imaging, ” Biomedical Optics Express , vol. 8, no. 11, pp. 5113–5126, Oct. 2017. [16] S. Nandy , W . Chapman, R. Rais, I. Gonzlez, D. Chatterjee, M. Mutch, and Q. Zhu, “Label-free quantitative optical assessment of human colon tissue using spatial frequency domain imaging, ” T echniques in Colopr octology , vol. 22, no. 8, pp. 617–621, Aug. 2018. [17] J. V ervandier and S. Gioux, “Single snapshot imaging of optical prop- erties, ” Biomedical Optics Expr ess , vol. 4, no. 12, pp. 2938–2944, Nov . 2013. [18] K. Suzuki, “Overview of deep learning in medical imaging, ” Radiolog- ical Physics and T echnology , vol. 10, no. 3, pp. 257–273, Sep. 2017. [19] H. Shin, H. R. Roth, M. Gao, L. Lu, Z. Xu, I. Nogues, J. Y ao, D. Mollura, and R. M. Summers, “Deep conv olutional neural networks for computer-aided detection: Cnn architectures, dataset characteristics and transfer learning, ” IEEE T ransactions on Medical Imaging , vol. 35, no. 5, pp. 1285–1298, May 2016. 10 [20] N. T ajbakhsh, J. Y . Shin, S. R. Gurudu, R. T . Hurst, C. B. Kendall, M. B. Gotway, and J. Liang, “Con volutional neural netw orks for medical image analysis: Full training or fine tuning?” IEEE T ransactions on Medical Imaging , vol. 35, no. 5, pp. 1299–1312, May 2016. [21] I. Goodfellow , J. Pouget-Abadie, M. Mirza, B. Xu, D. W arde-Farley , S. Ozair, A. Courville, and Y . Bengio, “Generative Adversarial Nets, ” in Advances in Neural Information Processing Systems 27 , Z. Ghahramani, M. W elling, C. Cortes, N. D. Lawrence, and K. Q. W einberger , Eds. Curran Associates, Inc., 2014, pp. 2672–2680. [22] M. Mirza and S. Osindero, “Conditional generative adversarial nets, ” 2014. [23] P . Isola, J.-Y . Zhu, T . Zhou, and A. A. Efros, “Image-to-image transla- tion with conditional adversarial networks, ” 2017 IEEE Conference on Computer V ision and P attern Recognition (CVPR) , Jul 2017. [24] A. Diaz-Pinto, A. Colomer, V . Naranjo, S. Morales, Y . Xu, and A. F . Frangi, “Retinal image synthesis and semi-supervised learning for glau- coma assessment, ” IEEE T ransactions on Medical Imaging , pp. 1–1, 2019. [25] Q. Y ang, P . Y an, Y . Zhang, H. Y u, Y . Shi, X. Mou, M. K. Kalra, Y . Zhang, L. Sun, and G. W ang, “Low-dose ct image denoising using a generativ e adv ersarial network with wasserstein distance and perceptual loss, ” IEEE T ransactions on Medical Imaging , vol. 37, no. 6, pp. 1348– 1357, June 2018. [26] G. Y ang, S. Y u, H. Dong, G. Slabaugh, P . L. Dragotti, X. Y e, F . Liu, S. Arridge, J. Keegan, Y . Guo, and D. Firmin, “Dagan: Deep de- aliasing generative adversarial networks for fast compressed sensing mri reconstruction, ” IEEE T ransactions on Medical Imaging , v ol. 37, no. 6, pp. 1310–1321, June 2018. [27] J. Swartling, J. S. Dam, and S. Andersson-Engels, “Comparison of spa- tially and temporally resolved diffuse-reflectance measurement systems for determination of biomedical optical properties, ” Appl. Opt. , vol. 42, no. 22, pp. 4612–4620, Aug 2003. [28] J. R. W eber, D. J. Cuccia, A. J. Durkin, and B. J. T romberg, “Noncontact imaging of absorption and scattering in layered tissue using spatially modulated structured light, ” Journal of Applied Physics , v ol. 105, no. 10, p. 102028, 2009. [29] G. M. Palmer, R. J. V iola, T . Schroeder , P . S. Y armolenko, M. W . Dewhirst, and N. Ramanujam, “Quantitative Diffuse Reflectance and Fluorescence Spectroscopy: A T ool to Monitor Tumor Physiology In V ivo, ” Journal of biomedical optics , vol. 14, no. 2, p. 024010, 2009. [30] G. Jones, N. T . Clancy, Y . Helo, S. Arridge, D. S. Elson, and D. Stoy- anov , “Bayesian estimation of intrinsic tissue oxygenation and perfusion from r gb images, ” IEEE T ransactions on Medical Imaging , vol. 36, no. 7, pp. 1491–1501, July 2017. [31] M. van de Giessen, J. P . Angelo, and S. Gioux, “Real-time, profile- corrected single snapshot imaging of optical properties, ” Biomedical Optics Express , vol. 6, no. 10, pp. 4051–4062, Sep. 2015. [32] S. G. Swapnesh Panigrahi, “Machine learning approach for rapid and accurate estimation of optical properties using spatial frequency domain imaging, ” Journal of Biomedical Optics , vol. 24, no. 7, pp. 1 – 6 – 6, 2018. [33] Y . Zhao, Y . Deng, F . Bao, H. Peterson, R. Istfan, and D. Roblyer , “Deep learning model for ultrafast multifrequency optical property extractions for spatial frequency domain imaging, ” Optics letters , vol. 43, no. 22, pp. 5669–5672, 2018. [34] S. Gioux, A. Mazhar , D. J. Cuccia, A. J. Durkin, B. J. T romberg, and J. V . Frangioni, “Three-Dimensional Surface Profile Intensity Correction for Spatially-Modulated Imaging, ” Journal of biomedical optics , vol. 14, no. 3, p. 034045, 2009. [35] M. Martinelli, A. Gardner , D. Cuccia, C. Hayakawa, J. Spanier, and V . V enugopalan, “Analysis of single Monte Carlo methods for prediction of reflectance from turbid media, ” Optics Express , vol. 19, no. 20, pp. 19 627–19 642, Sep. 2011. [36] R. Chen, F . Mahmood, A. Y uille, and N. J. Durr, “Rethinking monocular depth estimation with adversarial training, ” 2018. [37] O. Ronneberger , P . Fischer , and T . Brox, “U-net: Conv olutional networks for biomedical image segmentation, ” Medical Imag e Computing and Computer-Assisted Intervention MICCAI 2015 , p. 234241, 2015. [38] K. He, X. Zhang, S. Ren, and J. Sun, “Deep residual learning for image recognition, ” 2016 IEEE Conference on Computer V ision and P attern Recognition (CVPR) , Jun 2016. [39] T . M. Quan, D. G. C. Hildebrand, and W .-K. Jeong, “Fusionnet: A deep fully residual con volutional neural network for image segmentation in connectomics, ” CoRR , vol. abs/1612.05360, 2016. [40] T . Miyato and M. Koyama, “cgans with projection discriminator , ” 2018. [41] X. Mao, Q. Li, H. Xie, R. Y . Lau, Z. W ang, and S. P . Smolley , “Least squares generative adversarial networks, ” 2017 IEEE International Con- fer ence on Computer V ision (ICCV) , Oct 2017. [42] D. P . Kingma and J. Ba, “ Adam: A method for stochastic optimization, ” 2014. [43] V . V anhoucke, A. Senior, and M. Z. Mao, “Improving the speed of neural networks on CPUs, ” in Deep Learning and Unsupervised F eatur e Learning W orkshop, NIPS 2011 , 2011. [44] J. A. Sweer, K. Salimian, T . Chen, R. J. Battafarano, and N. J. Durr , “W ide-field optical property mapping and structured light imaging of the esophagus with spatial frequency domain imaging, ” Journal of biophotonics , p. e201900005, 2019. [45] A. Mazhar , S. Dell, D. J. Cuccia, S. Gioux, A. J. Durkin, J. V . Frangioni, and B. J. T romberg, “W av elength optimization for rapid chromophore mapping using spatial frequenc y domain imaging, ” J ournal of Biomedical Optics , vol. 15, no. 6, p. 061716, Dec. 2010. [46] F . A yers, A. Grant, D. Kuo, D. J. Cuccia, and A. J. Durkin, “Fabrication and characterization of silicone-based tissue phantoms with tunable optical properties in the visible and near infrared domain, ” in Design and P erformance V alidation of Phantoms Used in Conjunction with Optical Measur ements of T issue , vol. 6870. International Society for Optics and Photonics, Feb . 2008, p. 687007. [47] M. Lucic, K. Kurach, M. Michalski, S. Gelly , and O. Bousquet, “ Are GANs Created Equal? A Large-Scale Study, ” arXiv:1711.10337 [cs, stat] , Nov . 2017, arXiv: 1711.10337. [48] F . Mahmood, R. Chen, and N. J. Durr, “Unsupervised reverse domain adaptation for synthetic medical images via adversarial training, ” IEEE transactions on medical imaging , vol. 37, no. 12, pp. 2572–2581, 2018. [49] F . Mahmood, R. Chen, S. Sudarsk y , D. Y u, and N. J. Durr, “Deep learning with cinematic rendering: fine-tuning deep neural networks using photorealistic medical images, ” Physics in Medicine & Biology , vol. 63, no. 18, p. 185012, sep 2018.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment