Association of genomic subtypes of lower-grade gliomas with shape features automatically extracted by a deep learning algorithm
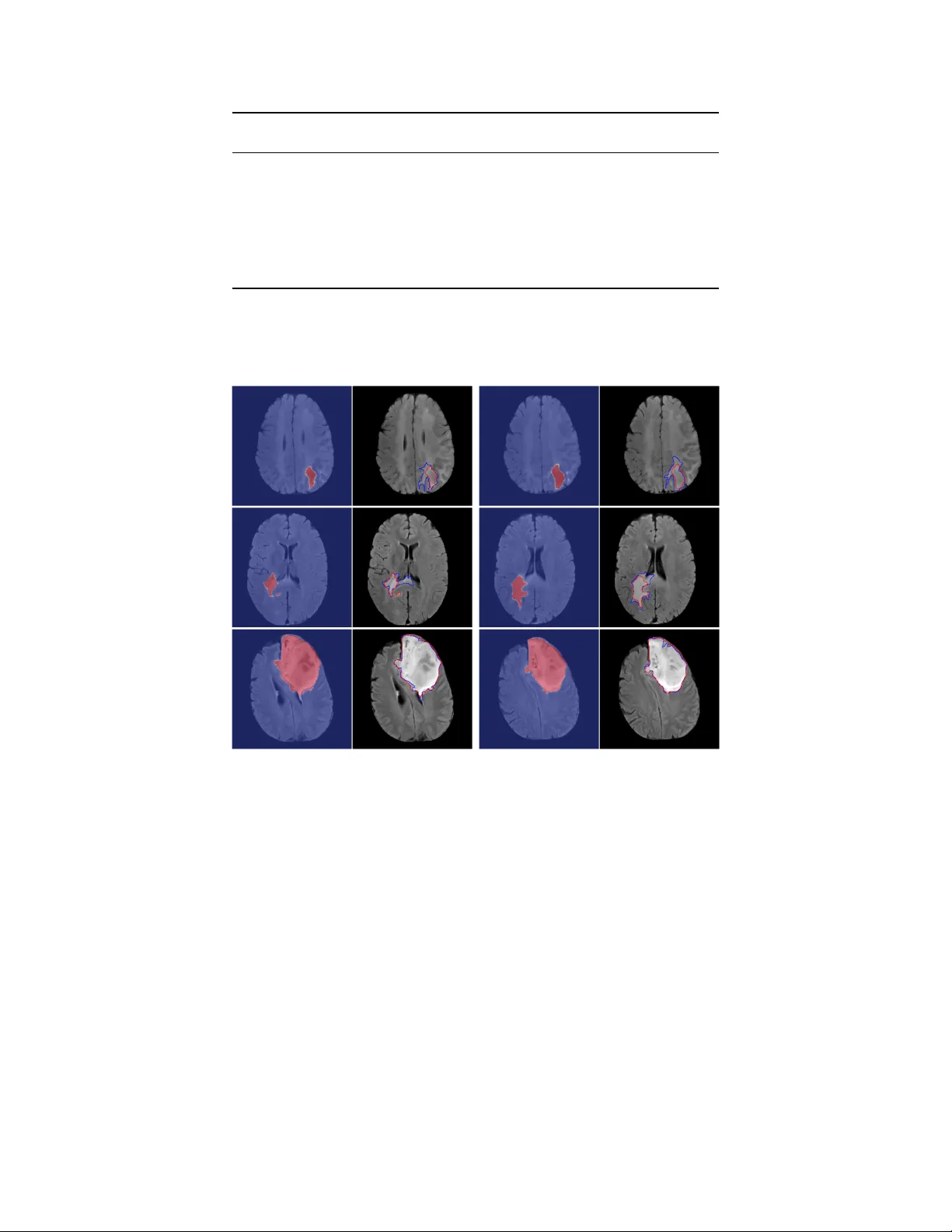
Recent analysis identified distinct genomic subtypes of lower-grade glioma tumors which are associated with shape features. In this study, we propose a fully automatic way to quantify tumor imaging characteristics using deep learning-based segmentati…
Authors: Mateusz Buda, Ashirbani Saha, Maciej A Mazurowski
Asso ciation of genomic subt yp es of lo w er-grade gliomas with shap e features automatically extracted b y a deep learning algorithm Mateusz Buda 1 , Ashirbani Saha 1 , Maciej A. Mazuro wski 1, 2 1 Departmen t of Radiology , Duke Universit y , Durham, NC 2 Departmen t of Electrical and Computer Engineering, Duk e Univ ersity , Durham, NC Abstract Recen t analysis identified distinct genomic subt yp es of low er-grade glioma tumors which are asso ciated with shap e features. In this study , w e propose a fully automatic w ay to quantify tumor imaging c haracteristics using deep learning-based segmentation and test whether these c haracteristics are predictiv e of tumor genomic subt yp es. W e used preop erative imaging and genomic data of 110 patien ts from 5 institutions with lo wer-grade gliomas from The Cancer Genome Atlas. Based on automatic deep learning seg- men tations, we extracted three features which quan tify tw o-dimensional and three-dimensional c haracteristics of the tumors. Genomic data for the analyzed cohort of patien ts consisted of previously iden tified genomic clusters based on IDH mutation and 1p/19q co-deletion, DNA meth ylation, gene expression, DNA copy num b er, and microRNA expression. T o analyze the relationship betw een the imaging features and genomic clusters, we conducted the Fisher exact test for 10 h ypotheses for each pair of imaging feature and genomic subt ype. T o account for m ultiple h yp othesis testing, we applied a Bonferroni correction. P-v alues lo wer than 0.005 were considered statistically significant. W e found the strongest association b etw een RNASeq clusters and the b ounding ellipsoid v olume ratio ( p < 0 . 0002) and b et ween RNASeq clusters and margin fluctuation ( p < 0 . 005). In addition, we identified asso ciations b et ween b ounding ellipsoid v olume ratio and all tested molecular subtypes ( p < 0 . 02) as well as betw een angular standard deviation and RNASeq cluster ( p < 0 . 02). In terms of automatic tumor segmen tation that was used to ge nerate the quan titative image characteristics, our deep learning algorithm achiev ed a mean Dice co efficien t of 82% which is comparable to human performance. Keyw ords: deep learning; brain segmen tation; radiogenomics; MRI; LGG 1 1 In tro duction Lo wer-grade gliomas (LGG) are a group of WHO grade I I and grade II I brain tumors. As opp osed to grade I which are often curable by surgical resection, grade I I and I I I are infiltrativ e and tend to recur and evolv e to higher-grade lesion. Predicting patient outcomes based on histopathologi- cal data for these tumors is inaccurate and suffers from in ter-observ er v ariability [ 1 ]. One of the promising metho ds that might address this issue is defining subtypes of LGG through clustering of patien ts based on DNA methylation, gene expression, DNA copy n umber, and microRNA expres- sion [ 1 ]. It was shown that the clusters iden tified in suc h wa y are to a large extent in agreemen t with another basic molecular subtype based on IDH (IDH1 and IDH2) m utation and 1p/19q co- deletion [ 1 , 2 ]. P atients with tumors from different molecular groups substan tially differ in terms of t ypical course of the disease and ov erall surviv al [ 3 ]. A new research direction in cancer, called radiogenomics, aims at inv estigating the relationship b et w een tumor genomic c haracteristics and medical imaging [ 4 ]. Imaging can pro vide important information before surgery or in cases when resection is not p ossible. V ery recen t studies in this area hav e disco vered an asso ciation of tumor shap e features extracted from MRI with its genomic subt yp es [ 5 , 6 ]. How ev er, the first step when extracting tumor features w as the manual segmen tation of MRI. Such annotation is costly , time consuming and results in annotations with high inter-rater v ariance [ 7 ]. Deep learning is a new field of machine learning that is recen tly rev olutionizing the automated analysis of images [ 8 , 9 ]. There are man y examples of successful applications of deep learning in medical imaging [ 10 , 11 , 12 , 13 , 14 ] and more sp ecifically in brain MRI segmentation [ 15 ]. In recent y ears, progress in deep learning for automatic brain segmentation matured to a level that ac hiev es p erformance of a skilled radiologist [ 16 ]. Most of these efforts are focused on glioblastoma rather than LGG. Dev elopment of mo dels that yield high qualit y segmentation of LGG in brain MRI w ould p otentially allow for automatization of the pro cess of tumor genomic subtype identification through imaging that is fast, inexp ensiv e, and free of in ter-reader v ariabilit y . In this study , w e combine the field of deep learning and radiogenomic and prop ose a fully automatic algorithm for quan tification of tumor shap e and test whether the assessed shape features are prognostic of tumor molecular subt yp es. Developing imaging-biomark ers that could inform of tumor genomics w ould pro vide the information to clinicians sooner in a non-inv asiv e w ay and in some cases could allo w for b etter stratification of tumors where resection is not p erformed. In this 2 study , w e show promise for ev entually developing such imaging-based biomark ers. The reminder of this pap er is organized as follows. Section 2 describ es data used in our study whereas section 3 describes segmen tation mo del, features used for tumor quan tification, and sta- tistical metho ds. Then, in section 4 we show results for the segmentation algorithm and prediction of genomic subt yp es. In section 5 w e discuss our findings. Finally , sections 6 and 7 are dev oted to limitations and conclusions of the study , resp ectiv ely . 2 Dataset 2.1 P atien t p opulation The data used in this study w as obtained from The Cancer Genome A tlas (TCGA) and The Cancer Imaging Archiv e (TCIA). W e identified 120 patien ts from TCGA lo wer-grade glioma collection 1 who had preop erativ e imaging data av ailable, con taining at least a fluid-atten uated inv ersion recov ery (FLAIR) sequence. T en patients had to b e excluded since they did not ha ve genomic cluster information av ailable. The final group of 110 patients w as from the follo wing 5 institutions: Thomas Jefferson Univ ersity (TCGA-CS, 16 patients), Henry F ord Hospital (TCGA-DU, 45 patien ts), UNC (TCGA-EZ, 1 patient), Case W estern (TCGA-FG, 14 patients), Case W estern – St. Joseph’s (TCGA-HT, 34 patien ts) from TCGA LGG collection. The complete list of patients used in this study is included in Online Resource 1. The en tire set of 110 patients was split into 22 non- o verlapping subsets of 5 patients eac h. This was done for ev aluation with cross-v alidation. 2.2 Imaging data Imaging data was obtained from The Cancer Imaging Arc hive 2 whic h con tains the images corre- sp onding to the TCGA patien ts and is sp onsored by the National Cancer Institute. W e used all mo dalities when a v ailable and only FLAIR in case any other modality w as missing. There were 101 patien ts with all sequences av ailable, 9 patients with missing p ost-con trast sequence, and 6 with missing pre-con trast sequence. The complete list of av ailable sequences for each patient is included in Online Resource 1. The n umber of slices v aried among patien ts from 20 to 88. In order to capture the original pattern of tumor growth, we only analyzed preop erativ e data. The assessment of tumor shap e was based on FLAIR abnormalit y since enhancing tumor in LGG is rare. 1 https://cancergenome.nih.gov/cancersselected/lowergradeglioma 2 https://wiki.cancerimagingarchive.net/display/Public/TCGA- LGG 3 A researc her in our lab oratory , who w as a medical school graduate with experience in neuroradi- ology imaging, manually annotated FLAIR images by dra wing an outline of the FLAIR abnormalit y on eac h slice to form training data for the automatic segmen tation algorithm. W e used soft ware dev elop ed in our lab oratory for this purp ose. A board eligible radiologist v erified all annotations and mo dified those that w ere identified as incorrect. Dataset of registered images together with man ual segmentation masks for each case used in our study is released and made publicly av ailable at the follo wing link: https://kaggle.com/mateuszbuda/lgg- mri- segmentation . 2.3 Genomic data Genomic data used in this study consisted of DNA meth ylation, gene expression, DNA copy n umber, and microRNA expression, as w ell as IDH m utation 1p/19q co-deletion measurement. Sp ecifically , in our analysis we consider six previously identified molecular classifications of LGG that are kno wn to b e correlated with some tumor shap e features [ 6 ]: 1. Molecular subtype based on IDH mutation and 1p/19q co-deletion (three subtypes: IDH m utation-1p/19q co-deletion, IDH mutation-no 1p/19q co-deletion, IDH wild type) 2. RNASeq clusters (4 clusters: R1-R4) 3. DNA meth ylation clusters (5 clusters: M1-M5) 4. DNA cop y num b er clusters (3 clusters: C1-C3) 5. microRNA expression clusters (4 clusters: mi1-mi4) 6. Cluster of clusters (3 clusters: co c1-coc3) 3 Metho ds 3.1 Automatic segmen tation Figure 1 shows the o verview of the segmen tation algorithm. The follo wing phases comprise the fully automatic algorithm for obtaining the segmen tation mask: image preprocessing, segmentation, and p ost-processing. Then, once the segmen tation masks are generated, w e extracted shap e features that were iden tified as predictive of molecular subt yp es. The following sections provide details on eac h of the steps. Source co de of the algorithm described in this section is also av ailable at the follo wing link: https://github.com/mateuszbuda/brain- segmentation . 4 Figure 1: A sc hema sho wing data processing steps of our system for molecular subtype inference from a sequence of brain MRI 3.1.1 Prepro cessing Images v aried significan tly b et ween patien ts in terms of size. The prepro cessing of the image sequences consisted of the follo wing steps: • Scaling of the images to the common frame of reference. • Remo v al of the skull to fo cus the analysis on the brain region (a.k.a., skull stripping). • Adaptiv e window and lev el adjustment based on the image histogram to normalize intensities of tissues b et w een cases. • Z-score normalization of the en tire data set. The details of all the pre-pro cessing steps are included in the Online Resource 2. 3.1.2 Segmen tation The main segmentation step was performed using a fully conv olutional neural net work with the U-Net architecture [ 10 ] sho wn in Figure 2. It comprises four lev els of blo c ks con taining tw o con- v olutional la yers with ReLU activ ation function and one max p ooling lay er in the encoding part and up-conv olutional la yers instead in the deco ding part [ 17 , 18 , 19 , 20 , 21 ]. Consistent with the U-Net arc hitecture, from the enco ding lay ers we use skip connections to the corresp onding lay ers in the deco ding part. They pro vide a shortcut for gradient flo w in shallo w lay ers during the training phase [ 22 ]. Man ual segmen tation serv ed as a grou nd truth for training a mo del for automatic segmentation. W e trained tw o netw orks, one for cases with three sequences av ailable (pre-con trast, FLAIR, and p ost-con trast) and the other that used only FLAIR. F or the second net w ork, instead of missing sequences w e used neigh b oring FLAIR slices from b oth sides of a slice of in terest as additional c hannels. Since in this scenario the tw o sequences, whic h o ccupied channel 1 and channel 3 of the input are not av ailable, we filled these channels with neighboring tumor slices to pro vide additional information to the net work. 5 Input 256x256x3 Conv 3x3, ReLU 256x256x32 Max pooling 2x2 128x128x32 Concatenation Conv 3x3, ReLU 128x128x64 Max pooling 2x2 64x64x64 Conv 3x3, ReLU 64x64x128 Max pooling 2x2 32x32x128 Conv 3x3, ReLU 32x32x256 Max pooling 2x2 16x16x256 Conv 3x3, ReLU 16x16x512 Conv 3x3, ReLU 16x16x512 Up-conv 2x2 32x32x256 Conv 3x3, ReLU 32x32x256 Up-conv 2x2 64x64x128 Conv 3x3, ReLU 64x64x128 Up-conv 2x2 128x128x64 Conv 3x3, ReLU 128x128x64 Up-conv 2x2 256x 256x32 Conv 3x3, ReLU 256x256x32 Conv 1x1, Sigmoid 256x256x1 Conv 3x3, ReLU 256x256x32 Conv 3x3, ReLU 128x128x64 Conv 3x3, ReLU 64x64x128 Conv 3x3, ReLU 32x32x256 Conv 3x3, ReLU 32x32x256 Conv 3x3, ReLU 64x64x128 Conv 3x3, ReLU 128x128x64 Conv 3x3, ReLU 256x256x32 Figure 2: U-Net architecture used for skull stripping and segmentation. Below eac h la yer sp ecification w e pro vide dimensionality of a single example that this la yer outputs The n umber of slices containing tumor w as considerably low er than those with only background class present. Therefore, to accoun t for this fact, we applied ov ersampling with data augmen tation that was pro ved to help in training conv olutional neural netw orks [ 23 ]. W e did it by ha ving three instances of eac h tumor slice in our training set. F or one o versampled slice w e applied random rotation b y 5 to 15 degrees and for the other slice w e applied random scale b y 4% to 8%. T o further reduce the imbalance b etw een tumor and non-tumor pixels, w e discarded empty slices that did not con tain an y brain or other tissue after applying skull stripping. This step has b een undertak en since training a fully conv olutional neural netw ork with image s that do not contain an y p ositiv e v oxels can be highly detrimental. Please note that a significant ma jority of vo xels in the abnormal slices are still normal and therefore sufficien t negative data is a v ailable for training. 6 3.1.3 P ost-pro cessing T o further impro v e the accuracy , w e implemented a post-pro cessing algorithm that remov es false p ositiv es. Specifically , w e extracted all tumor volumes using connected comp onen ts algorithm on a three-dimensional segmentation mask for each patien t. W e did it using 6-connected pixels in three dimensions, i.e. neigh b oring pixels are defined as b eing connected along primary axes. Ev entually , w e included in the final segmentation mask only the pixels comprising the largest connected tu- mor volume. This p ost-pro cessing strategy b enefits extraction of shap e features (describ ed in the follo wing section) since they are sensitive to isolated false p ositive pixel segmentations. 3.1.4 Extraction of shap e features W e consider three shape features of a segmen ted tumor that w ere identified as imp ortan t in the con text of low er grade glioma radiogenomic [ 6 ]: A ngular standar d deviation (ASD) is the av erage of the radial distance standard deviations from the centroid of the mass across ten equiangular bins in one slice, as describ ed in [ 24 ]. Before calculating the v alue of this feature, w e normalize radial distances to ha ve mean equal one. Angular standard deviation of a tumor shap e is a quan titative measure of v ariation in the tumor margin within relatively small parts of the tumor. It also captures non-circularit y of the tumor, i.e. lo w v alue indicates circle like shap e. Bounding el lipsoid volume r atio (BEVR) is the ratio b et ween the v olume of segmented FLAIR abnormalit y and its minim um b ounding ellipsoid. This feature captures the irregularit y of the tumor in three dimensions. If the tumor fits well into its b ounding ellipsoid (high v alue of BEVR), it is considered more regular while if more space in the b ounding ellipsoid is unfilled, the shap e is considered irregular. Mar gin fluctuation (MF) is computed as follows. First, we find the centroid of the tumor and distances from it to all pixels on the tumor boundary in one slice. Then, we apply a veraging filter of length equal to 10% of the tumor p erimeter measured in the num b er of pixels. Margin fluctuation is the standard deviation of the difference b et ween v alues b efore and after smo othing, i.e. applying a veraging filter. Similarly as in ASD, radial distances are normalized to hav e a mean of one. This is done in order to remov e the impact of tumor size on the v alue of this feature. Margin fluctuation is a tw o-dimensional feature that quan tifies the amoun t of high frequency c hanges, i.e. smoothness of the tumor b oundary and w as previously used for analysis of spiculation in breast tumors [ 25 , 26 ]. 7 3.2 Statistical analysis Our hypothesis was that fully automatically-assessed shap e features are predictive of tumor molec- ular subt yp es. Since we considered 6 definitions of molecular subt yp es based on genomic assa ys and multiple imaging features, we fo cused our analysis on the relationships betw een imaging and genomics that w ere found significan t (with man ual tumor segmen tation) in a previous study [ 6 ]. Sp ecifically , those were the follo wing relationships: b ounding ellipsoid v olume ratio with RNASeq, miRNA, CNC, and COC, the relationship of Margin fluctuation with RNASeq, and the relationship of angular standard deviation with IDH/1p19q, RNASeq, Methylation, CNC, and COC resulting in 10 sp ecific hypotheses. T o assess statistical significance of these asso ciations, we conducted the Fisher exact test (fisher.test function in R) for each of 10 com binations of imaging and genomics. F or the purp ose of this test, we turned eac h contin uous imaging v ariable v alue into a num b er from 1 to 4 based on which quartile of the feature v alue it fell into. F or each of the imaging and genomic feature com binations, we used only the cases that had b oth: imaging and genomic subtype data a v ailable. W e conducted a total of 10 statistical tests for eac h pair of imaging feature and genomic subt yp e for our primary hypothesis. T o accoun t for m ultiple h yp othesis testing, w e applied a Bonferroni correction. P-v alues lo w er than 0.005 (0.05/10) w ere considered statistically significant for our primary radiogenomics h yp otheses. Additionally , w e ev aluated performance of the deep learning-based segmentation itself. W e used Dice similarit y coefficient [ 27 ] as the ev aluation metric which measures the ov erlap b et ween the segmen tation provided by the algorithm and the man ually-annotated gold standard. In the ev aluation pro cess, we used cross-v alidation. Sp ecifically , we divided our en tire dataset in to 22 subsets, each con taining exactly 5 cases. The mo del training was conducted on the training subsets and then the model w as applied to the test cases. This was rep eated 22 times un til eac h subset s erv ed once as the test set. The cases then w ere p ooled for the analysis as describ ed ab o ve. The n umber of cases included in the training and test sets (whic h determines the num b er of folds) is a trade-off b et ween computational cost of training m ultiple mo dels and having more data to train eac h of them. The t wo extremes of this approach are lea ve-one-out strategy which results in one-case folds and the other is 50% split whic h giv es 2 folds. W e found folds of 5 patien ts to b e a go od balance b et ween a training set size and computational cost. 8 4 Results The patients’ c haracteristics are sho wn in T able 1. The a verage patien t age w as 47. Fift y six of the patients were w omen, fifty three w ere men, and the gender of one was unknown. The tumors’ c haracteristics are provided in T able 2. There w ere three histological types of tumors: oligo dendroglioma (47), astro cytoma (33), oligoastro cytoma (29), and one unknown. Grade of the tumors in our data included 51 cases of grade I I, 58 of grade I I I, and grade of one tumor was unkno wn. Characteristic P atients (N=110) Age (y ears) Median 47 Range 20-75 Gender F emale 56 Male 32 Not a v ailable 1 T able 1: Patien t characteristic. Age for one patient w as missing and was ignored in the calculation. The results of the radiogenomic analysis are shown in Figure 3. W e confirmed our primary h y- p othesis for t wo pairs of imaging features and genomic subt yp e. W e found the strongest asso ciation b et w een RNASeq cluster and the b ounding ellipsoid volume ratio ( p < 0 . 0002) along with margin fluctuation ( p < 0 . 005). In addition, we iden tified considerable correlations for the b ounding el- lipsoid volume ratio and all tested molecular subtypes ( p < 0 . 02) as well as for angular standard deviation and RNASeq cluster ( p < 0 . 02). In T able 3 w e provide R OC AUC scores for the task of discriminating eac h RNASeq cluster from all other subtypes based on shap e features extracted from segmentation masks obtained with the deep learning algorithm and compare them to man ual segmentations. W e selected RNASeq cluster since it sho wed the strongest association with shap e features. In addition, w e included ROC A UC based on tw o demographic v ariables, i.e. age and gender. The results sho w that deep learning w as able to pro vide tumor segmen tations of a quality that allow ed for extraction of shap e features that match man ual segmen tations. Sp ecifically , cluster R2 was distinguished from all other clusters 9 based on in versed bounding ellipsoid v olume ratio with AUC of 0.80 and 0.78 for deep learning- based and manual segmentations, respectively . In terms of angular standard deviation, A UC for deep learning w as 0.73 and for man ual segmen tations w as 0.72. The predictive v alue of demographic v ariables for R2 cluster was notably lo wer with AUC=0.66 for age and A UC=0.50 for gender. A detailed comparison of man ual and automatic segmentation for the task of discriminating cluster R2 from all other clusters with resp ect to sensitivity , sp ecificit y , p ositiv e predictiv e v alue, and negativ e predictive v alue is given in Online Resource 1. In terms of tumor segmen tation, our deep learning algorithm ac hiev ed mean Dice co efficient of 82% and median Dice co efficien t of 85%. F or the 101 cases with all sequences av ailable, mean and median Dice co efficien t was the same as for all 110 cases. F or the remaining 9 cases segmen ted based on FLAIR sequence only , mean and median Dice co efficien t was 82% and 88%, resp ectively . Characteristic Cases (N=110) Histologic t yp e and grade Astro cytoma Grade I I 8 Grade I II 25 Oligoastro cytoma Grade I I 14 Grade I II 21 Oligo dendroglioma Grade I I 29 Grade I II 18 Not a v ailable 1 IDH-1p/19q subt yp e IDH m utation, 1p/19q co-deletion 26 IDH m utation, no 1p/19q co-deletion 56 IDH wild t yp e 25 Not a v ailable 3 T able 2: T umor characteristic. 10 R1 R2 R3 R4 RNASeq cluster 0.2 0.25 0.3 0.35 0.4 0.45 0.5 0.55 Bounding ellipsoid volume ratio mi1 mi2 mi3 mi4 miRNA cluster 0.2 0.25 0.3 0.35 0.4 0.45 0.5 0.55 Bounding ellipsoid volume ratio C1 C2 C3 CNC cluster 0.2 0.25 0.3 0.35 0.4 0.45 0.5 0.55 Bounding ellipsoid volume ratio coc1 coc2 coc3 COC cluster 0.2 0.25 0.3 0.35 0.4 0.45 0.5 0.55 Bounding ellipsoid volume ratio R1 R2 R3 R4 RNASeq cluster 0.02 0.04 0.06 0.08 0.1 Margin fluctuation IDHmut-codel IDHmut-nocodel IDHwt Molecular subtype (IDH, 1p/19q) 0.05 0.1 0.15 0.2 0.25 Angular standard deviation R1 R2 R3 R4 RNASeq cluster 0.05 0.1 0.15 0.2 0.25 Angular standard deviation M1 M2 M3 M4 M5 Methylation cluster 0.05 0.1 0.15 0.2 0.25 Angular standard deviation C1 C2 C3 CNC cluster 0.05 0.1 0.15 0.2 0.25 Angular standard deviation coc1 coc2 coc3 COC cluster 0.05 0.1 0.15 0.2 0.25 Angular standard deviation Figure 3: Box and whisker plots demonstrating the relationship betw een tested genomic clusters and imaging features that quantify tumor shape Examples of segmen tations that we obtained alongside with ground truth masks for cases of v ary- ing performance of our segmen tation algorithm are presented in Figure 4. Due to max po oling la yers included in the U-Net arc hitecture which allo w for pro cessing of large volumes on currently a v ailable computers, the automated segmentation is less sensitive to high curv ature and sulcation and therefore, the pro duced masks tend to b e more smo oth comparing to manual segmentations. 11 V ariable R1 R2 R3 R4 Deep learning ASD 0.26 0.73 0.53 0.47 Deep learning 1/BEVR 0.23 0.80 0.36 0.53 Age 0.29 0.66 0.72 0.41 Gender 0.44 0.50 0.58 0.52 T able 3: R OC A UCs of shap e features and demographic v ariables for the task of discriminating one RNASeq cluster from all others. Figure 4: Examples of automatic segmentation for low (top), mo derate (middle), and high (b ottom) agree- men t with ground truth. Their Dice coefficient is 50%, 82% and 95% resp ectiv ely . In each pair, the first image sho ws a heatmap of ra w mo del output and in the second image blue outline corresp onds to ground truth and red to postpro cessed automatic segmentation output. Images show FLAIR mo dalit y after prepro cessing and skull stripping. Giv en small sample size, we additionally assessed the stabilit y and p erformance of our segmen- tation model using predictions generated for noisy input. F or eac h input image, w e applied additiv e Gaussian noise with zero mean and standard deviation equal to 10% and 20% of standard devia- tion computed on the training data. In the first case with 10% noise level, mean Dice co efficien t decreased to 81%. After increasing noise level to 20%, mean Dice co efficien t further decreased to 79%. In b oth cases, median Dice co efficien t was 85%. 12 5 Discussion In this study , w e were able to demonstrate that fully automatically-assessed imaging features of lo w grade gliomas are asso ciated with tumor molecular subtypes established using genomic analysis. The strength of these asso ciations w as sho wn to b e mo derate. Deep learning algorithms were used to segmen t the tumors. Using imaging to predict tumor genomics is of very high imp ortance and if accurate mo dels are developed, it could b e incorporated in the curren t treatment paradigm in a v ariet y of w ays. In the simplest scenario, if the mo del is highly accurate, it could simply replace the genomic analysis altogether. Curren t state of the art radiogenomic models, suc h as the one shown in this pap er whic h represen ts mo derate predictive p erformance, do not yet justify such substitution. Ho wev er, there are other w ays in whic h ev en mo dels with mo derate p erformance could contribute v aluable information. The imaging data is av ailable early in the pro cess and therefore approximate assessmen t of tumor biology b efore the surgery could still b e of help in guiding the next steps. The appro ximate imaging surrogate of molecular subt yp e w ould also be of particular use for patien ts that do not immediately undergo surgical excision of the tumor. In such case, in the absence of tissue analysis, the appro ximate classification by imaging could b e of very high v alue since genomic subt yp es are highly correlated with patient outcomes. If biopsy results are av ailable for a tumor that has not b een fully resected, the imaging surrogate of subtype can still b e of use given a p otential high intra-tumor heterogeneit y of the lesions and therefore a p ossibility that a local biopsy do es not accurately reflect the ov erall genomics of a tumor. Imaging offers a complete view of a tumor. Finally , even if the ov erall accuracy is not p erfect, it migh t b e p ossible to op erate at a high p ositiv e predictiv e v alue or high predictiv e v alue and the surrogate imaging-based mo dels of genomic subtypes could b e still used for triaging patients for genomic tests, ev en if it is a small minorit y of the patients. F or example, if based on imaging there is a high confidence that a tumor is of a particular aggressiv e subt yp e, the patien t could b e treated accordingly without additional exp ensiv e and in v asive genomic testing. If on the other hand the imaging-based mark er has lo w confidence, then additional genomic test could b e ordered. An imp ortan t step tow ard accurate and repro ducible assessment of imaging features of low er- grade gliomas is accurate segmen tation of the tumors. While the annotation by radiologists is considered a gold standard, a considerable inter observer v ariability has b een documented for this task. F or the whole tumor segmentation of LGG on brain MRI, Dice co efficien t b et ween tw o 13 exp ert raters is 84% with standard deviation of 2% [ 7 ]. This demonstrates that our algorithm falls within acceptable lev el precision. At the same time, our algorithm provides a fully reproducible and consisten t wa y of tumor quan tification for future cases. Automatic segmentation of tumors such as the one show ed in this study has multiple adv antages. First, it addresses the inter-observ er v ariability described ab o ve. Since there is only one reader (the computer algorithm), the inter-observ er v ariability is non-existen t. F urthermore, it addresses the problem of intra-observ er v ariability . The algorithm is deterministic which means that giv en the same image, the algorithm will alwa ys perform an iden tical assessmen t. Finally , application of a computer algorithm is inexpensive and fast. The performance of our segmentation algorithm in terms of the mean Dice co efficien t w as 82% whic h puts it on a par with exp ert human readers. This w as ac hieved with the help of deep learning which has demonstrated similar phenomenal p erformance in other applications. Our results sho w that RNASeq R2 cluster, compared to other clusters, is associated with the tumors of notably higher irregularit y of shap e as quan tified by bounding ellipsoid volume ratio, an- gular standard deviation and margin fluctuation. R2 cluster is in turn link ed to considerably p o orer o verall surviv al as compared with R1, R3, and R4 [ 1 ]. The same conclusion can be dra wn for molec- ular subtype of IDH wild type which indicates less fa vorable prognosis that is close to glioblastoma prognosis [ 1 ]. It is asso ciated with relativ ely high angular standard deviation. This is consistent with findings obtained in the previous study for shap e features extracted from manually segmen ted tumors [ 6 ]. This p oints out to a conclusion th at angular standard deviation, margin fluctuation, b ounding ellipsoid volume ratio, and p oten tially other features that measure the irregularit y of tumor shap e may be prognostic of patien t’s outcome. 6 Limitations This study had limitations. It constitutes only a first step to ward imaging-based surrogates of genomic subtypes. Sp ecifically , only three imaging features w ere considered. While these features w ere selected based on prior evidence of their effectiveness and therefore are of high importance, they constitute a small sample of different features that could b e calculated including texture and enhancemen t of the tumor and its surroundings. F urthermore, a fairly limited sample size was used in the study (110 patien ts) since data that con tains comprehensiv e genome-wide assays alongside with imaging is still rare. While no separate v alidation set w as a v ailable in this study , w e utilized 14 a commonly used cross-v alidation technique, which splits the data in to training and test sets to a void a p ositive ev aluation bias. Regarding segmentation algorithms, there are many methods for performing automatic seg- men tation of brain tumors that could be considered for comparison and to further improv e our results [ 28 , 29 ]. First, regarding general approac hes to segmen tation, in [ 16 ] deep learning with sliding-windo w approach w as used. An impro vemen t that significantly reduces computational com- plexit y is a fully conv olutional neural netw ork which allows for pro cessing en tire image in one for- w ard pass [ 30 , 31 ]. In U-Net architecture, used in our study , additional skip connections b et w een enco der and decoder parts of the netw ork are used [ 10 ]. Second, regarding net work arc hitecture, differen t types hav e b een prop osed, e.g. ResNet [ 32 ], Inception [ 33 ], and DenseNet [ 34 ], whic h w ere incorp orated in segmentation mo dels. Finally , v arious optimization functions for training deep learning segmentation models w ere prop osed. The most commonly applied is cross-en trop y loss, used also in classification mo dels. How ever, for highly imbalanced segmen tation tasks, loss functions based on Dice similarit y co efficien t outperformed other loss functions in man y applica- tions [ 22 , 35 , 36 , 37 ]. 7 Conclusions In conclusion, w e demonstrated that features of MRI, extracted in a fully automatic manner using deep learning algorithms, w ere associated with tumor molecular subt yp es of lo w er-grade gliomas determined using genomic assays. This shows promise for repro ducible non-in v as iv e imaging-based surrogates of tumor genomics in brain cancer. References [1] Cancer Genome Atlas Research Net work. Comprehensive, in tegrative genomic analysis of diffuse lo wer-grade gliomas. New England Journal of Me dicine , 372(26):2481–2498, 2015. [2] Chang-Ming Zhang and Daniel J Brat. Genomic profiling of lo wer-grade gliomas uncov ers cohesiv e disease groups: implications for diagnosis and treatmen t. Chinese journal of c anc er , 35(1):12, 2016. [3] Jeanette E Ec k el-Passo w, Daniel H Lachance, Annette M Molinaro, Kyle M W alsh, Paul A 15 Dec ker, Hugues Sicotte, Melike P ekmezci, T erri Rice, Matt L Kosel, and Iv an V Smirnov. Glioma groups based on 1p/19q, idh, and tert promoter mutations in tumors. New England Journal of Me dicine , 372(26):2499–2508, 2015. [4] Maciej A Mazurowski. Radiogenomics: what it is and why it is imp ortant. Journal of the A meric an Col le ge of R adiolo gy , 12(8):862–866, 2015. [5] Maciej A Mazurowski, Kal Clark, Nicholas M Czarnek, P arisa Shamsesfandabadi, Kather- ine B P eters, and Ashirbani Saha. Radiogenomic analysis of low er grade glioma: a pilot m ulti-institutional study shows an asso ciation b et ween quantitativ e image features and tumor genomics. In Me dic al Imaging 2017: Computer-Aide d Diagnosis , volume 10134, page 101341T. In ternational So ciet y for Optics and Photonics, 2017. [6] Maciej A Mazuro wski, Kal Clark, Nic holas M Czarnek, P arisa Shamsesfandabadi, Katherine B P eters, and Ashirbani Saha. Radiogenomics of lo w er-grade glioma: algorithmically-assessed tumor shap e is asso ciated with tumor genomic subt yp es and patient outcomes in a m ulti- institutional study with the cancer genome atlas data. Journal of neur o-onc olo gy , 133(1): 27–35, 2017. [7] Bjo ern H Menze, Andras Jak ab, Stefan Bauer, Jay ashree Kalpathy-Cramer, Keyv an F arahani, Justin Kirby , Y uliya Burren, Nicole P orz, Johannes Slotb o om, and Roland Wiest. The mul- timo dal brain tumor image segmentation b enc hmark (brats). IEEE tr ansactions on me dic al imaging , 34(10):1993–2024, 2014. [8] Ian Go odfellow. Deep learning of represen tations and its application to computer vision. 2015. [9] Anastasia Ioannidou, Elisa vet Chatzilari, Spiros Nik olop oulos, and Ioannis Kompatsiaris. Deep learning adv ances in computer vision with 3d data: A survey . A CM Computing Surveys (CSUR) , 50(2):20, 2017. [10] Olaf Ronneb erger, Philipp Fischer, and Thomas Brox. U-net: Conv olutional net works for biomedical image segmentation. In International Confer enc e on Me dic al image c omputing and c omputer-assiste d intervention , pages 234–241. Springer, 2015. [11] ¨ Ozg ¨ un C ¸ i¸ cek, Ahmed Ab dulk adir, So eren S Lienk amp, Thomas Brox, and Olaf Ronneb erger. 3d u-net: learning dense v olumetric segmen tation from sparse annotation. In International 16 c onfer enc e on me dic al image c omputing and c omputer-assiste d intervention , pages 424–432. Springer, 2016. [12] F austo Milletari, Nassir Na v ab, and Seyed-Ahmad Ahmadi. V-net: F ully conv olutional neural net works for volumetric medical image segmen tation. In 2016 F ourth International Confer enc e on 3D Vision (3D V) , pages 565–571. IEEE, 2016. [13] Hao Chen, Qi Dou, Lequan Y u, Jing Qin, and Pheng-Ann Heng. V o xresnet: Deep vo xelwise residual net works for brain segmentation from 3d mr images. Neur oImage , 170:446–455, 2018. [14] Maciej A Mazuro wski, Mateusz Buda, Ashirbani Saha, and Mustafa R Bashir. Deep learning in radiology: An o verview of the concepts and a surv ey of the state of the art with fo cus on mri. Journal of Magnetic R esonanc e Imaging , 49(4):939–954, 2019. [15] Mohammad Hav aei, Nicolas Guizard, Hugo Laro c helle, and Pierre-Marc Jo doin. Deep learn- ing trends for fo cal brain pathology segmen tation in mri. In Machine L e arning for He alth Informatics , pages 125–148. Springer, 2016. [16] Mohammad Hav aei, Axel Davy , David W arde-F arley , Antoine Biard, Aaron Courville, Y osh ua Bengio, Chris Pal, Pierre-Marc Jo doin, and Hugo Laro chelle. Brain tumor segmentation with deep neural net works. Me dic al image analysis , 35:18–31, 2017. [17] Xa vier Glorot, An toine Bordes, and Y oshua Bengio. Deep sparse rectifier neural net works. In Pr o c e e dings of the fourte enth international c onfer enc e on artificial intel ligenc e and statistics , pages 315–323, 2011. [18] Y ann LeCun and Y oshua Bengio. Con volutional netw orks for images, sp eech, and time series. The handb o ok of br ain the ory and neur al networks , 3361(10):1995, 1995. [19] Y ann LeCun, L´ eon Bottou, Y oshua Bengio, and Patric k Haffner. Gradient-based learning applied to do cumen t recognition. Pr o c e e dings of the IEEE , 86(11):2278–2324, 1998. [20] Alex Krizhevsky , Ilya Sutskev er, and Geoffrey E Hin ton. Imagenet classification with deep con volutional neural net works. In A dvanc es in neur al information pr o c essing systems , pages 1097–1105, 2012. [21] Hy eonw o o Noh, Seungho on Hong, and Boh yung Han. Learning deconv olution netw ork for 17 seman tic segmentation. In Pr o c e e dings of the IEEE international c onfer enc e on c omputer vision , pages 1520–1528, 2015. [22] Mic hal Drozdzal, Eugene V orontso v, Gabriel Chartrand, Samuel Kadoury , and Chris Pal. The imp ortance of skip connections in biomedical image segmentation. In De ep L e arning and Data L ab eling for Me dic al Applic ations , pages 179–187. Springer, 2016. [23] Mateusz Buda, Atsut o Maki, and Maciej A Mazurowski. A systematic study of the class im balance problem in conv olutional neural net works. Neur al Networks , 106:249–259, 2018. [24] Harris Georgiou, Michael Ma vroforakis, Nik os Dimitrop oulos, Dionisis Cav ouras, and Sergios Theo doridis. Multi-scaled morphological features for the characterization of mammographic masses using statistical classification schemes. A rtificial Intel ligenc e in Me dicine , 41(1):39–55, 2007. [25] Mary ellen L Giger, Carl J Vyb orn y , and Rob ert A Schmidt. Computerized characterization of mammographic masses: analysis of spiculation. Canc er L etters , 77(2-3):201–211, 1994. [26] Scott Pohlman, Kimerly A P ow ell, Nancy A Obuc howski, William A Chilcote, and Sharon Grundfest-Broniato wski. Quantitativ e classification of breast tumors in digitized mammo- grams. Me dic al Physics , 23(8):1337–1345, 1996. [27] Lee R Dice. Measures of the amoun t of ecologic association b et ween sp ecies. Ec olo gy , 26(3): 297–302, 1945. [28] Geert Litjens, Thijs Ko oi, Babak Eh teshami Bejnordi, Arnaud Arindra Adiy oso Setio, F rancesco Ciompi, Mohsen Ghafo orian, Jero en Awm V an Der Laak, Bram V an Ginneken, and Clara I S´ anc hez. A surv ey on deep learning in medical image analysis. Me dic al image analysis , 42:60–88, 2017. [29] Zeynettin Akkus, Alfiia Galimzianov a, Assaf Ho ogi, Daniel L Rubin, and Bradley J Erickson. Deep learning for brain mri segmen tation: state of the art and future directions. Journal of digital imaging , 30(4):449–459, 2017. [30] Vija y Badrinara yanan, Alex Kendall, and Rob erto Cip olla. Segnet: A deep conv olutional enco der-decoder architecture for image segmen tation. IEEE tr ansactions on p attern analysis and machine intel ligenc e , 39(12):2481–2495, 2017. 18 [31] Jonathan Long, Ev an Shelhamer, and T revor Darrell. F ully con volutional net works for se- man tic segmentation. In Pr o c e e dings of the IEEE c onfer enc e on c omputer vision and p attern r e c o gnition , pages 3431–3440, 2015. [32] Qiao Zhang, Zhip eng Cui, Xiaoguang Niu, Shijie Geng, and Y u Qiao. Image segmen tation with pyramid dilated conv olution based on resnet and u-net. In International Confer enc e on Neur al Information Pr o c essing , pages 364–372. Springer, 2017. [33] Christian Szegedy , Vincen t V anhouc ke, Sergey Ioffe, Jon Shlens, and Zbigniew W o jna. Re- thinking the inception arc hitecture for computer vision. In Pr o c e e dings of the IEEE c onfer enc e on c omputer vision and p attern r e c o gnition , pages 2818–2826, 2016. [34] Simon J ´ egou, Michal Drozdzal, David V azquez, Adriana Romero, and Y oshua Bengio. The one hundred lay ers tiramisu: F ully conv olutional densenets for seman tic segmen tation. In Pr o c e e dings of the IEEE Confer enc e on Computer Vision and Pattern R e c o gnition Workshops , pages 11–19, 2017. [35] Carole H Sudre, W enqi Li, T om V ercauteren, Sebastien Ourselin, and M Jorge Cardoso. Gen- eralised dice ov erlap as a deep learning loss function for highly un balanced segmentations. In De ep le arning in me dic al image analysis and multimo dal le arning for clinic al de cision supp ort , pages 240–248. Springer, 2017. [36] Lucas Fidon, W enqi Li, Luis C Garcia-P eraza-Herrera, Jinendra Ek ana yak e, Neil Kitchen, S ´ ebastien Ourselin, and T om V ercauteren. Generalised wasserstein dice score for im balanced m ulti-class segmen tation using holistic conv olutional netw orks. In International MICCAI Br ainlesion Workshop , pages 64–76. Springer, 2017. [37] Jiac hi Zhang, Xiaolei Shen, Tianqi Zhuo, and Hong Zhou. Brain tumor segmen tation based on refined fully conv olutional neural net works with a hierarc hical dice loss. arXiv pr eprint arXiv:1712.09093 , 2017. 19
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment