Accurate and Robust Alignment of Variable-stained Histologic Images Using a General-purpose Greedy Diffeomorphic Registration Tool
Variously stained histology slices are routinely used by pathologists to assess extracted tissue samples from various anatomical sites and determine the presence or extent of a disease. Evaluation of sequential slides is expected to enable a better understanding of the spatial arrangement and growth patterns of cells and vessels. In this paper we present a practical two-step approach based on diffeomorphic registration to align digitized sequential histopathology stained slides to each other, starting with an initial affine step followed by the estimation of a detailed deformation field.
💡 Research Summary
This paper addresses the challenge of aligning sequential histology slides that have been stained with different dyes, a common scenario in digital pathology. The authors propose a practical two‑step registration pipeline that first performs a global affine alignment and then refines the result with a diffeomorphic deformation field. The method leverages the open‑source “Greedy” registration tool, originally designed for 3‑D medical imaging, and adapts it to 2‑D whole‑slide images with minimal parameter tuning.
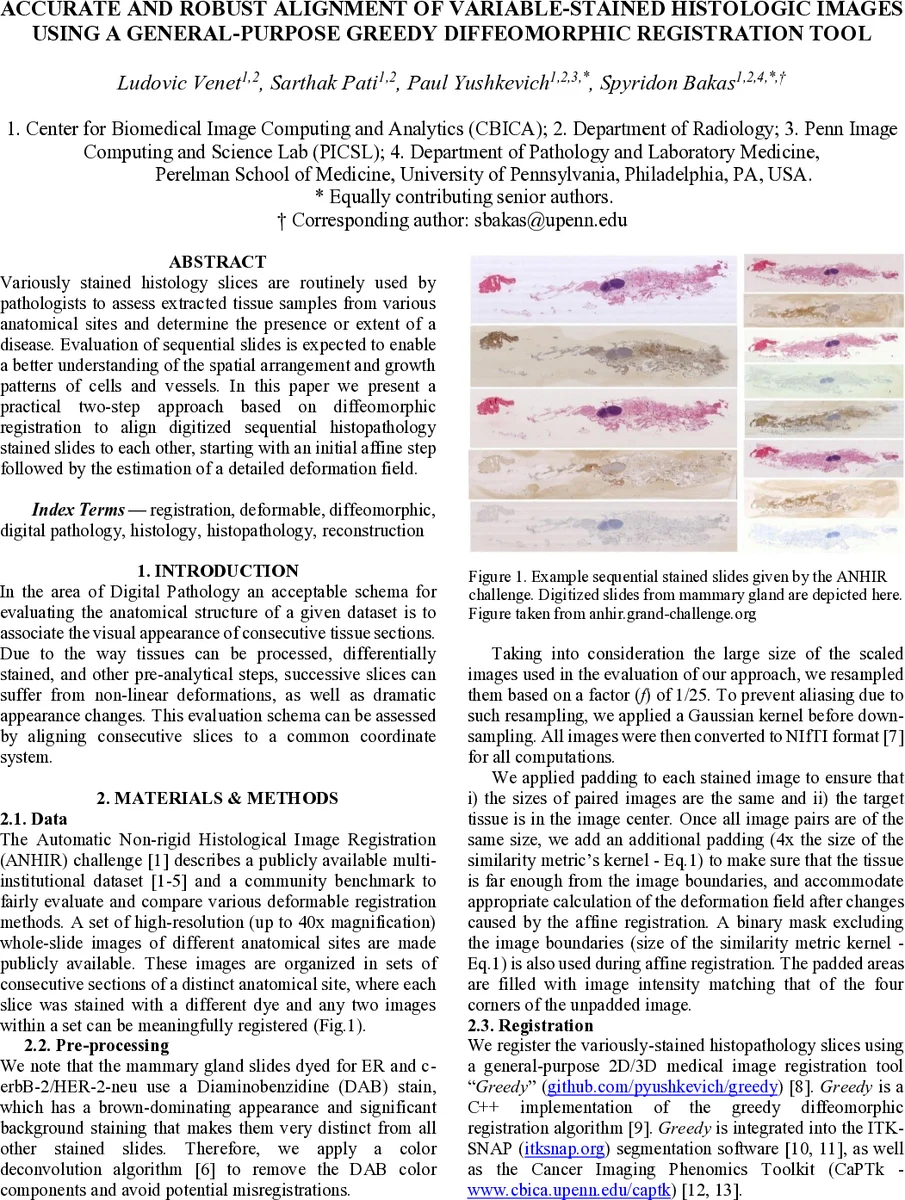
The dataset used for evaluation is the ANHIR (Automatic Non‑rigid Histological Image Registration) challenge collection, which provides high‑resolution whole‑slide images (up to 40× magnification) from multiple anatomical sites, each slice stained with a different dye (e.g., H‑E, DAB, PAS). Because DAB‑stained slides exhibit a strong brown background that can dominate similarity measures, the authors first apply the Ruifrok‑Johnston color deconvolution algorithm to remove the DAB component. All images are then down‑sampled by a factor of 1/25, preceded by Gaussian smoothing to avoid aliasing. Images are converted to NIfTI format, padded to equal dimensions, and an additional margin (four times the similarity‑metric kernel size) is added to ensure that tissue remains far from image borders during deformation computation.
Affine registration: An initial rigid transformation is obtained by a brute‑force search over 5,000 random rotations and translations, selecting the pose that maximizes Normalized Cross‑Correlation (NCC). This pose serves as the starting point for a Limited‑memory BFGS (LBFGS) optimization that refines the affine parameters. NCC is computed after scaling the fixed image so that the kernel captures sufficient structural information.
Diffeomorphic registration: The Greedy tool implements a “greedy diffeomorphic” algorithm that shares many concepts with the Symmetric Normalization (SyN) method but sacrifices symmetry for speed. NCC remains the similarity metric, and a multi‑resolution pyramid (4×, 2×, full resolution) is used with iteration counts of 100, 50, and 10 respectively. Regularization parameters (σ₁ = 6 px, σ₂ = 5 px) are selected via a grid search on the training set. Greedy applies recursive Gaussian smoothing to the velocity field at each iteration, ensuring that the Jacobian determinant stays positive, thus guaranteeing a diffeomorphic (topology‑preserving) transformation.
Evaluation: Performance is measured using the median relative Target Registration Error (rTRE), computed from manually annotated landmarks and normalized by the image diagonal. The affine‑only stage yields an average rTRE of 0.00473, while the full affine + diffeomorphic pipeline achieves 0.00279, the best score on the ANHIR leaderboard. Robustness, defined as the proportion of successfully registered pairs, is reported as 1.0.
Key contributions and insights:
- Demonstrates that a general‑purpose 3‑D registration engine can be repurposed for 2‑D histology with only minor adjustments.
- Shows that simple color deconvolution and careful padding effectively mitigate large appearance variations caused by different stains.
- Validates that NCC, combined with multi‑scale processing and efficient Gaussian smoothing, provides both speed and accuracy for very large whole‑slide images.
Limitations and future work: The current preprocessing targets only DAB removal; other complex staining combinations remain untested. The method’s performance on very fine structures (e.g., capillaries) is not separately quantified. Future directions include comparison with feature‑based (salient point) registration, integration into full 3‑D reconstruction pipelines, and development of automated parameter selection using learned multi‑scale features.
Overall, the study presents a robust, efficient solution for aligning variably stained histologic sections, facilitating downstream 3‑D tissue reconstruction and multimodal analyses that combine morphological and molecular information—an essential step toward more comprehensive digital pathology workflows, especially in oncology research.
Comments & Academic Discussion
Loading comments...
Leave a Comment