Robustness and Games Against Nature in Molecular Programming
Matter, especially DNA, is now programmed to carry out useful processes at the nanoscale. As these programs and processes become more complex and their envisioned safety-critical applications approach deployment, it is essential to develop methods for engineering trustworthiness into molecular programs. Some of this can be achieved by adapting existing software engineering methods, but molecular programming also presents new challenges that will require new methods. This paper presents a method for dealing with one such challenge, namely, the difficulty of ascertaining how robust a molecular program is to perturbations of the relative “clock speeds” of its various reactions. The method proposed here is game-theoretic. The robustness of a molecular program is quantified in terms of its ability to win (achieve its original objective) in games against other molecular programs that manipulate its relative clock speeds. This game-theoretic approach is general enough to quantify the security of a molecular program against malicious manipulations of its relative clock speeds. However, this preliminary report focuses on games against nature, games in which the molecular program’s opponent perturbs clock speeds randomly (indifferently) according to the probabilities inherent in chemical kinetics.
💡 Research Summary
The paper tackles a fundamental reliability issue in molecular programming: the variability of reaction rate constants, which can be thought of as “clock speeds” of individual chemical reactions. As DNA‑based molecular systems move toward safety‑critical applications such as biosensing, diagnostics, and targeted drug delivery, it becomes essential to quantify how robust a molecular program is when these clock speeds are perturbed. The authors propose a game‑theoretic framework that models a molecular program (represented as a chemical reaction network, CRN) as a player competing against other CRNs that can influence its reaction rates. A special “nature” player is introduced, which perturbs the rates randomly according to the stochastic mass‑action kinetics of chemistry, thereby providing a baseline for robustness against uncontrolled environmental fluctuations.
The technical foundation begins with a formal definition of CRNs: a finite set of species S and a finite set of reactions R, each reaction ρ = (r, p, k) consisting of reactant vector r, product vector p, and rate constant k. Under Gillespie’s stochastic mass‑action semantics, a reaction’s propensity in state x is k·V^{1‑arity(ρ)}·∏_{X∈S} x(X)^{r(ρ)(X)}. This yields a continuous‑time Markov chain whose trajectories (states together with sojourn times) constitute the observable behavior of the molecular program.
The game is defined for n players, each selecting a CRN from a strategy space N_i. The joint CRN N_σ is the union of all selected networks, and a trajectory τ of the combined system determines each player’s utility u_i(τ). The “nature” player has zero utility and does not care about the outcome; its only effect is to modify the rates of reactions belonging to other players. Expected utility U_i(σ, ξ) is computed by integrating u_i over the probability measure induced by the Markov chain, given an initial distribution ξ of species counts.
Robustness is quantified as α‑robustness: a CRN N₁ is α‑robust against opponents (N₂,…,N_n) if its expected utility when playing against them is at least an α‑fraction of its utility when the opponents are trivial (empty) CRNs. Formally, U₁((N₁,N₂,…,N_n), ξ) ≥ α·U₁((N₁,∅,…,∅), ξ′), where ξ′ corresponds to the same initial state for player 1 but empty initial states for the others. An α close to 1 indicates strong robustness.
Because many molecular systems interact only through catalysts, the authors restrict attention to “catalytic games.” In such games each player’s species set S_i splits into active species A_i (disjoint across players) and catalyst species C_i. Catalysts appear in reactions without net change (Δρ(C)=0), so they can only affect other players by altering reaction rates, not by consuming or producing active species. This restriction mirrors realistic DNA strand‑displacement implementations where auxiliary strands act as catalysts.
The framework is illustrated with the approximate majority algorithm, a classic CRN that decides which of two species X or Y is initially in the majority and drives the system to a state where the majority species occupies the entire population. The original CRN R consists of two reactions: 2X + Y → 3X (rate ∝ x(x‑1) y) X + 2Y → 3Y (rate ∝ x y(y‑1)). To embed this in a catalytic game, the authors introduce catalyst species A and B, yielding R′: 2X + Y + A → 3X + A X + 2Y + B → 3Y + B. If a and b (the counts of A and B) are equal and constant, R′ behaves exactly like R but uniformly faster. To model random perturbations, a “nature” CRN N_k is added: A → B (rate k) B → A (rate k), which swaps A and B at a high rate k (chosen as 10⁹). This causes a and b to fluctuate randomly, thereby randomly scaling the two competing reaction rates.
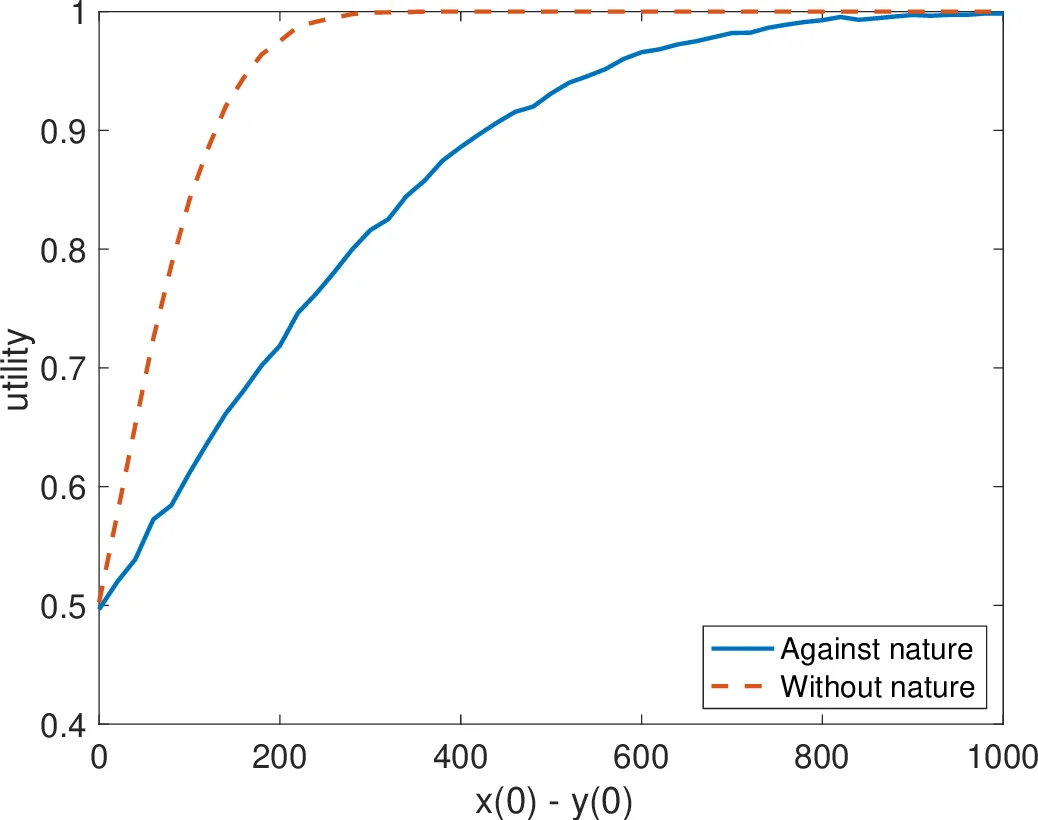
Simulation experiments were performed in MATLAB SimBiology. Initial conditions were set to a(0)=b(0)=100, total active population x + y = 10 000, and the initial imbalance Δ = x(0) − y(0) varied from 0 to 1 000 in steps of 10. For each Δ, one million stochastic runs were executed both with and without the nature CRN. Success was defined as the system reaching a state where either x or y becomes zero (the other species holds the entire population). Without nature, success rates were ≈99 % across all Δ. With nature (k = 10⁹), success rates dropped modestly but remained above 70 % for every Δ; for example, at Δ = 240 (x = 5 120, y = 4 880) success fell from 99 % to 76 %. Consequently, R′ is at least 0.7‑robust against N_{10⁹} under the considered initial distributions.
The authors argue that this game‑theoretic approach provides a systematic, quantitative measure of robustness that can be directly linked to design specifications. It also generalizes to adversarial settings where a malicious molecular program deliberately manipulates reaction rates, offering a pathway toward security analysis of molecular systems. Limitations include the binary utility model (success/failure) and the focus on a single opponent; extending the framework to multi‑objective utilities (e.g., time to convergence, energy consumption) and to equilibrium analysis with multiple strategic players is identified as future work. Experimental validation with actual DNA strand‑displacement implementations is also suggested.
In conclusion, the paper introduces a novel, mathematically grounded method for assessing the robustness of molecular programs against stochastic perturbations of reaction rates. By framing the problem as a game against nature, it bridges concepts from software engineering, stochastic chemical kinetics, and game theory, laying groundwork for more reliable and secure molecular computing systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment