Cascaded V-Net using ROI masks for brain tumor segmentation
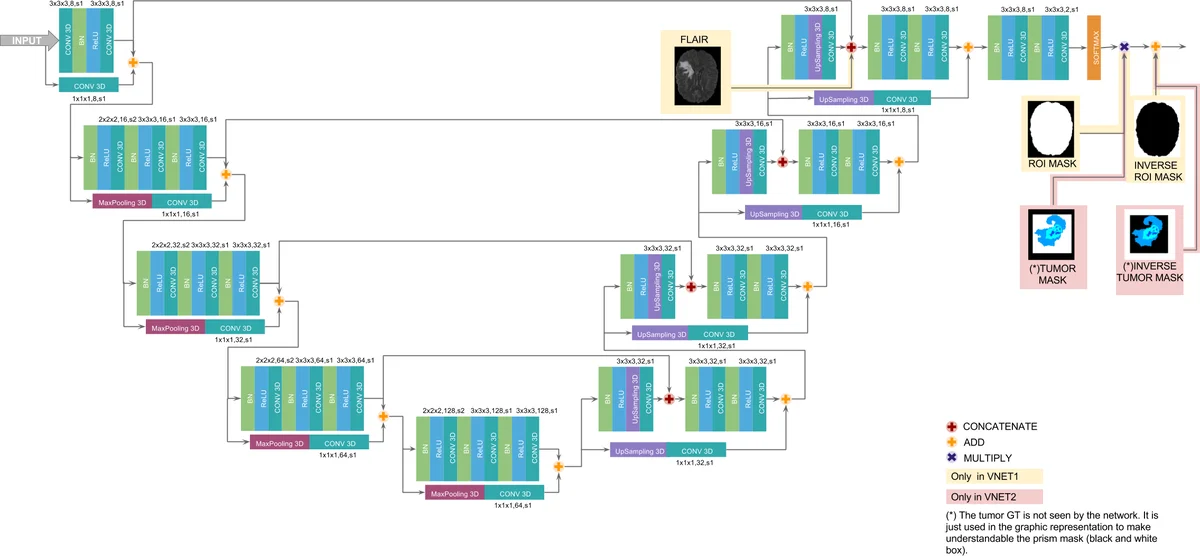
In this work we approach the brain tumor segmentation problem with a cascade of two CNNs inspired in the V-Net architecture \cite{VNet}, reformulating residual connections and making use of ROI masks to constrain the networks to train only on relevant voxels. This architecture allows dense training on problems with highly skewed class distributions, such as brain tumor segmentation, by focusing training only on the vecinity of the tumor area. We report results on BraTS2017 Training and Validation sets.
💡 Research Summary
This paper presents a two‑stage 3‑D convolutional neural network (CNN) architecture for automatic brain tumor segmentation that builds on the V‑Net design and incorporates region‑of‑interest (ROI) masking to address the severe class imbalance inherent in the task. The authors first modify the original V‑Net by reducing filter size from 5×5×5 to 3×3×3, replacing PReLU with ReLU, and inserting batch normalization before each non‑linearity. They also redesign the short residual connections: when spatial dimensions differ, they apply max‑pooling, repeat up‑sampling, and a 1×1×1 convolution to align the tensors, then add an identity‑mapping residual. These changes lower the total number of parameters, improve gradient flow, and make training more stable.
The overall pipeline consists of two cascaded networks. The first network performs a binary segmentation of the whole tumor (WT). It receives all four MR modalities (T1, T1‑c, T2, FLAIR) and is trained only on voxels inside a brain mask, thereby discarding background and skull. The loss is a Dice‑based coefficient tailored for highly imbalanced binary data. The output of this network is a raw WT mask.
The second network refines the WT mask into four classes: background, edema, enhancing tumor (ET), and non‑enhancing core (TC). It is trained only on voxels inside a rectangular ROI that tightly encloses the WT mask produced by the first stage. This ROI masking prevents the network from being overwhelmed by the vast majority of non‑tumor voxels and forces learning to focus on the tumor vicinity. The loss combines categorical cross‑entropy with Dice terms for each tumor sub‑region (WT, ET, TC) using empirically chosen weights (L₂ = XE + 0.5·(D_WT + D_ET + D_TC)).
Training is performed in a dense‑training regime: each mini‑batch contains a single whole‑patient volume rather than small patches. This approach leverages the full anatomical context while keeping memory requirements manageable. Data augmentation consists of left‑right (sagittal) reflections.
Experiments were conducted on the BraTS 2017 dataset (210 high‑grade, 75 low‑grade training scans; 46 validation scans). Evaluation metrics include Dice coefficient, Hausdorff distance, sensitivity, and specificity. For the whole‑tumor class, the method achieves Dice scores of 0.71–0.88 and Hausdorff distances of 5–9 mm, comparable to top‑performing entries in the BraTS challenge. However, performance drops markedly for the enhancing tumor and tumor core classes, with Dice scores around 0.63–0.64 and Hausdorff distances exceeding 11 mm. Sensitivity for ET and TC is also low (≈0.72), indicating many small tumor sub‑regions are missed. Training curves reveal that after roughly 18 epochs (≈3600 iterations) the Dice loss for the tumor core begins to overfit, while the cross‑entropy component continues to decrease, suggesting a tension between the two loss terms.
Qualitative visualizations confirm that the first stage reliably localizes the tumor mass, but the second stage often misclassifies sub‑regions, over‑segmenting edema at the expense of ET and TC. The authors attribute this to the persistent imbalance between large edema regions and the much smaller enhancing and necrotic cores, even after ROI masking.
In conclusion, the cascaded V‑Net with ROI masks is effective for coarse tumor localization but insufficient for precise delineation of small sub‑structures. The study highlights that dense‑training alone does not fully solve the class‑imbalance problem; additional strategies such as dynamic loss weighting, attention mechanisms, or more sophisticated multi‑scale sampling are required. Future work will explore these avenues to improve detection of the less common tumor components while preserving the efficiency gains of the proposed dense‑training framework.
Comments & Academic Discussion
Loading comments...
Leave a Comment