Tamper-Evident Complex Genomic Networks
Networks are important storage data structures now used to store personal information of individuals around the globe. With the advent of personal genome sequencing, networks are going to be used to store personal genomic sequencing of people. In contrast to social media networks, the importance of relationships in this genomic network is extremely significant. Losing connections between individuals thus implies losing relationship information (E.g. father or son etc.). There currently exists a considerably serious problem in the current approach to storing network data. Simply stated, network data is not tamper-evident. In other words, if some links or nodes were changed/removed/added by a malicious attacker, it would be impossible for the administrator to detect such changes. While, in the current age of social media networks, change in node characteristics and links can be bad in terms of relationships, in the case of networks for storing personal genomes, the results could be truly devastating. Here we present a scheme for building tamper-evident networks using a combination of Cryptographic and Ego-based Network analytic methods. Using actual published data-sets, we also demonstrate the utility and validity of the scheme besides demonstrating its working in various possible scenarios of usage. Results from the extensive experiments demonstrate the validity of the proposed approach.
💡 Research Summary
This paper addresses a critical security vulnerability in modern network data storage systems: the lack of tamper-evidence. While networks are increasingly used to store highly sensitive information, such as personal genomic sequencing data which encodes vital familial relationships, current architectures offer no inherent mechanism for administrators to detect if nodes or links have been maliciously altered, added, or deleted. The consequences of undetected tampering in genomic networks could be devastating, leading to corrupted family trees or erroneous medical insights.
To solve this problem, the authors propose a novel “cognitive digital footprinting” framework that creates a tamper-evident seal for any network by fusing concepts from network science and cryptography. The core idea leverages the sensitivity of network centrality measures—mathematical indices that quantify the importance or position of a node within a graph’s topology. The method specifically utilizes five key centrality metrics: Degree, Betweenness, Closeness, Eccentricity, and Eigenvector centrality.
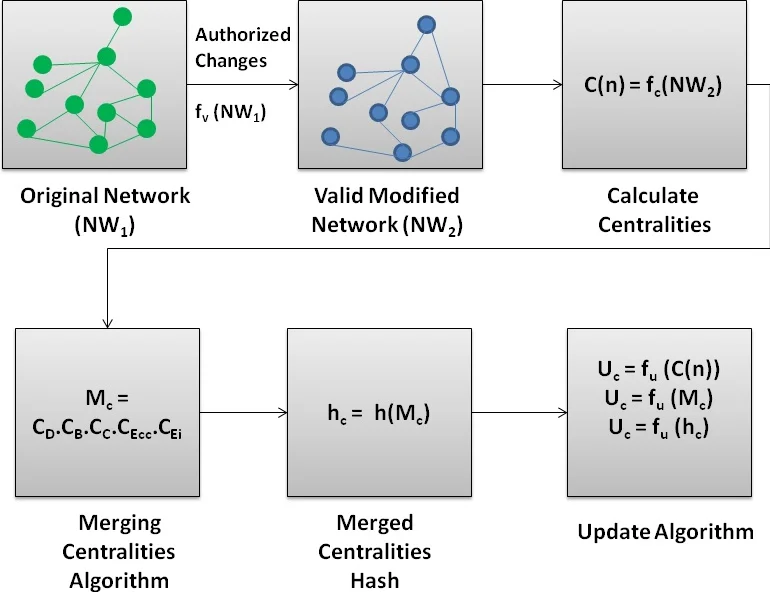
The proposed algorithm follows a systematic pipeline. First, the centrality values for every node in the network are calculated. Second, these multi-dimensional centrality profiles for all nodes are merged into a single, comprehensive metadata representation (M_c) that encapsulates the complete structural fingerprint of the network. Third, this metadata string is processed through a cryptographic hash function (e.g., SHA-256) to generate a fixed-length, unique digital fingerprint (h_c) of the network’s state. This hash value serves as the baseline for integrity.
In operational use, an administrator securely stores the hash of the original, trusted network state. Periodically, or after any suspected incident, the same process (centrality calculation -> merging -> hashing) is run on the current network. The newly generated hash is compared against the stored baseline. An exact match confirms the network’s integrity, while a mismatch provides undeniable proof that the network’s structure has been tampered with, as even a minuscule change in connections will alter centrality values and produce a completely different hash.
The validity of the approach is demonstrated through a proof-of-concept using the well-known Zachary Karate Club social network dataset. The experiments simulate three scenarios: (I) the original network, establishing the baseline hash; (II) a validly modified network (e.g., with a legitimate new link), which yields a different, but explainable hash; and (III) a maliciously tampered network. In the third scenario, the comparison of the current hash with the baseline would immediately reveal the discrepancy, enabling detection. This experiment concretely shows how structural changes are propagated through centrality metrics and magnified by the hash function.
The study’s significant contribution is its pragmatic and efficient hybrid approach. By focusing on the network’s relational topology rather than the potentially enormous content data (like genome sequences), it reduces computational overhead. Simultaneously, by incorporating cryptographic hashing, it provides a robust, non-repudiable integrity check. This method extends beyond genomic networks, offering a general-purpose solution for securing any complex network where the integrity of relationships—such as in financial transaction networks, supply chains, or critical infrastructure maps—is paramount.
Comments & Academic Discussion
Loading comments...
Leave a Comment