Batch Incremental Shared Nearest Neighbor Density Based Clustering Algorithm for Dynamic Datasets
Incremental data mining algorithms process frequent updates to dynamic datasets efficiently by avoiding redundant computation. Existing incremental extension to shared nearest neighbor density based clustering (SNND) algorithm cannot handle deletions to dataset and handles insertions only one point at a time. We present an incremental algorithm to overcome both these bottlenecks by efficiently identifying affected parts of clusters while processing updates to dataset in batch mode. We show effectiveness of our algorithm by performing experiments on large synthetic as well as real world datasets. Our algorithm is up to four orders of magnitude faster than SNND and requires up to 60% extra memory than SNND while providing output identical to SNND.
💡 Research Summary
The paper addresses the problem of clustering dynamic datasets that undergo frequent insertions and deletions. While the static Shared Nearest Neighbor Density‑based clustering algorithm (SNND) yields high‑quality clusters, re‑executing it after every change is computationally prohibitive. An earlier incremental extension, InSD, mitigates some redundancy but suffers from two critical limitations: it cannot process deletions and it handles insertions only one point at a time, leading to repeated full‑graph updates and reclustering.
The authors propose BISD (Batch Incremental Shared Nearest Neighbor Density‑based clustering), a batch‑oriented incremental algorithm that overcomes both shortcomings. BISD first partitions a set of updates into an insertion batch (n_add) and a deletion batch (n_del). For each data point it maintains two neighbor lists: the standard k‑nearest‑neighbors (KNN) used for clustering, and an extended w‑nearest‑neighbors list (w ≥ k) that serves as a buffer for incremental updates.
During the insertion phase, BISD computes the w‑list for every new point by measuring distances to all existing points and to other new points. Existing points are classified into three categories: T1_add (points whose KNN list actually changes because a new point becomes one of their k‑nearest neighbors), T2_add (points whose KNN list remains unchanged but at least one of their current KNN neighbors belongs to T1_add), and U_add (unaffected points). The deletion phase mirrors this process: after removing points in n_del, points whose KNN list shrinks are placed in T1_del, points whose KNN list stays the same but have a neighbor in T1_del go to T2_del, and the rest to U_del.
BISD then merges the two phases by forming T1 = (T1_add ∪ T1_del) − n_del and T2 = ((T2_add ∪ T2_del) − T1) − n_del. The SNN graph, stored as adjacency lists, is updated selectively: all vertices in T1 have their adjacency lists fully rebuilt, while vertices in T2 are updated only for edges that involve a neighbor from T1. This selective updating drastically reduces the amount of work compared with rebuilding the entire graph from scratch.
After the graph has been refreshed, BISD runs the original SNND clustering routine: vertices are labeled as core (degree ≥ δ_core), non‑core, or outlier; core vertices form initial clusters; clusters are merged when core vertices are linked; and non‑core vertices are assigned to the cluster with the strongest link. Because the final SNN graph is identical to the one SNND would produce on the fully recomputed dataset, the clustering output of BISD (C_BISD) is guaranteed to be identical to SNND’s output (C_SNND).
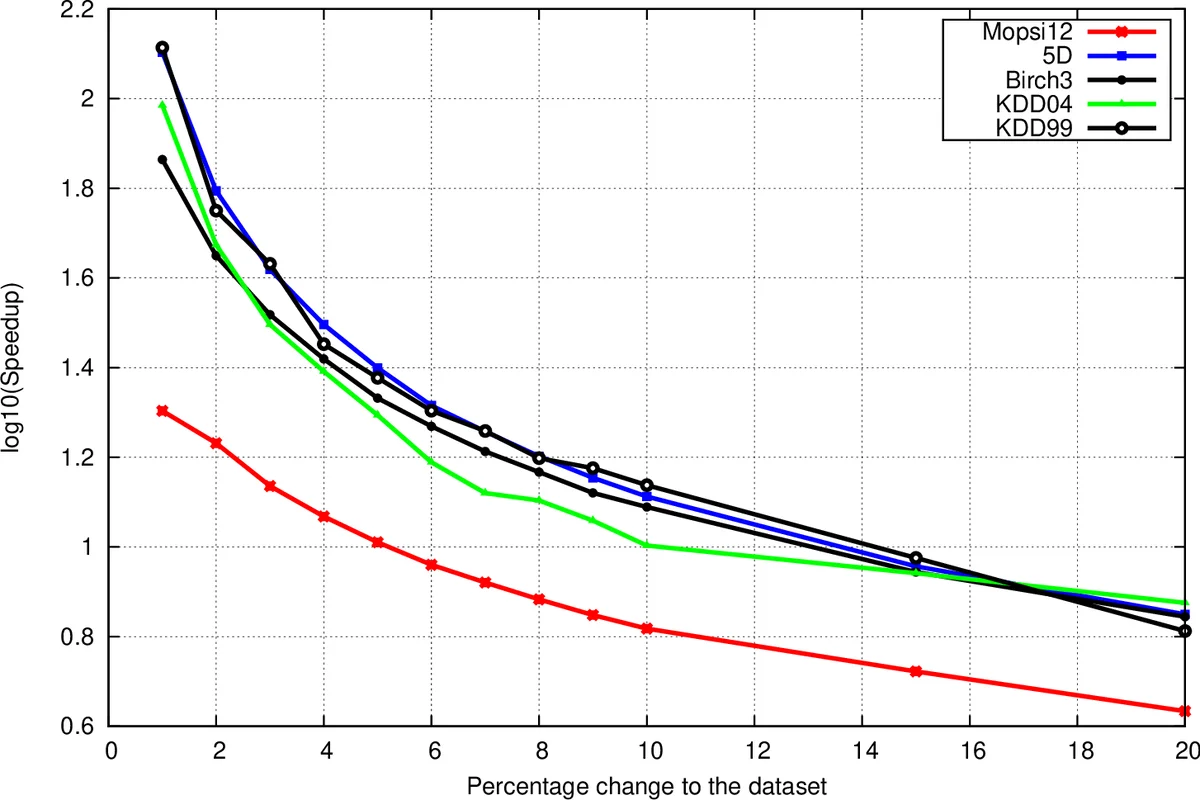
The authors evaluate BISD on five datasets (three real‑world: Mopsi12, KDDCup’99, KDDCup’04; two synthetic: Birch3, 5D) ranging from 13 k to 100 k points and 2 to 70 dimensions. Implemented in C++ and run on a 32 GB RAM, 1.56 GHz machine, BISD achieves speed‑ups of up to four orders of magnitude over SNND and up to four times faster than InSD, while incurring at most a 60 % memory overhead due to storing the extended neighbor lists. The speed‑up diminishes as the proportion of changed points grows because more vertices fall into T1 or T2, but BISD consistently outperforms both baselines.
In conclusion, BISD provides a practical solution for incremental clustering of dynamic data: it supports both insertions and deletions, aggregates batch updates to avoid redundant work, and delivers exactly the same clustering result as the original static algorithm with bounded additional memory. The paper suggests extending this batch‑incremental framework to other density‑based clustering methods as future work.
Comments & Academic Discussion
Loading comments...
Leave a Comment