Alignment of dynamic networks
Networks can model real-world systems in a variety of domains. Network alignment (NA) aims to find a node mapping that conserves similar regions between compared networks. NA is applicable to many fields, including computational biology, where NA can guide the transfer of biological knowledge from well- to poorly-studied species across aligned network regions. Existing NA methods can only align static networks. However, most complex real-world systems evolve over time and should thus be modeled as dynamic networks. We hypothesize that aligning dynamic network representations of evolving systems will produce superior alignments compared to aligning the systems’ static network representations, as is currently done. For this purpose, we introduce the first ever dynamic NA method, DynaMAGNA++. This proof-of-concept dynamic NA method is an extension of a state-of-the-art static NA method, MAGNA++. Even though both MAGNA++ and DynaMAGNA++ optimize edge as well as node conservation across the aligned networks, MAGNA++ conserves static edges and similarity between static node neighborhoods, while DynaMAGNA++ conserves dynamic edges (events) and similarity between evolving node neighborhoods. For this purpose, we introduce the first ever measure of dynamic edge conservation and rely on our recent measure of dynamic node conservation. Importantly, the two dynamic conservation measures can be optimized using any state-of-the-art NA method and not just MAGNA++. We confirm our hypothesis that dynamic NA is superior to static NA, under fair comparison conditions, on synthetic and real-world networks, in computational biology and social network domains. DynaMAGNA++ is parallelized and it includes a user-friendly graphical interface.
💡 Research Summary
The paper addresses a fundamental limitation of existing network alignment (NA) techniques, which have been confined to static graphs, by introducing the first method capable of aligning dynamic networks: DynaMAGNA++. Building on the state‑of‑the‑art static aligner MAGNA++, the authors extend the alignment objective to incorporate temporal information intrinsic to evolving systems.
Two novel conservation measures are defined. First, Conserved Event Time (CET) quantifies the total duration during which a pair of aligned edges is simultaneously active in both networks, while Non‑Conserved Event Time (NCET) measures the total duration when only one of the two edges is active. These metrics replace the static S3 edge‑conservation score and capture the temporal overlap of interactions rather than mere presence/absence. Second, a dynamic node‑conservation score is derived from a time‑aware graphlet degree vector (GDV). The authors aggregate graphlet counts over time, reduce dimensionality with principal component analysis, and compute cosine similarity between the reduced vectors of candidate node pairs.
The combined objective function is α·S_E + (1‑α)·S_N, where S_E is the CET/NCET‑based edge score and S_N is the dynamic GDV‑based node score. The weight α is set to 0.5, mirroring the balance used in MAGNA++. Optimization proceeds via a genetic‑algorithm framework identical to MAGNA++: an initial population of random alignments evolves over generations through crossover and mutation, directly maximizing the new objective.
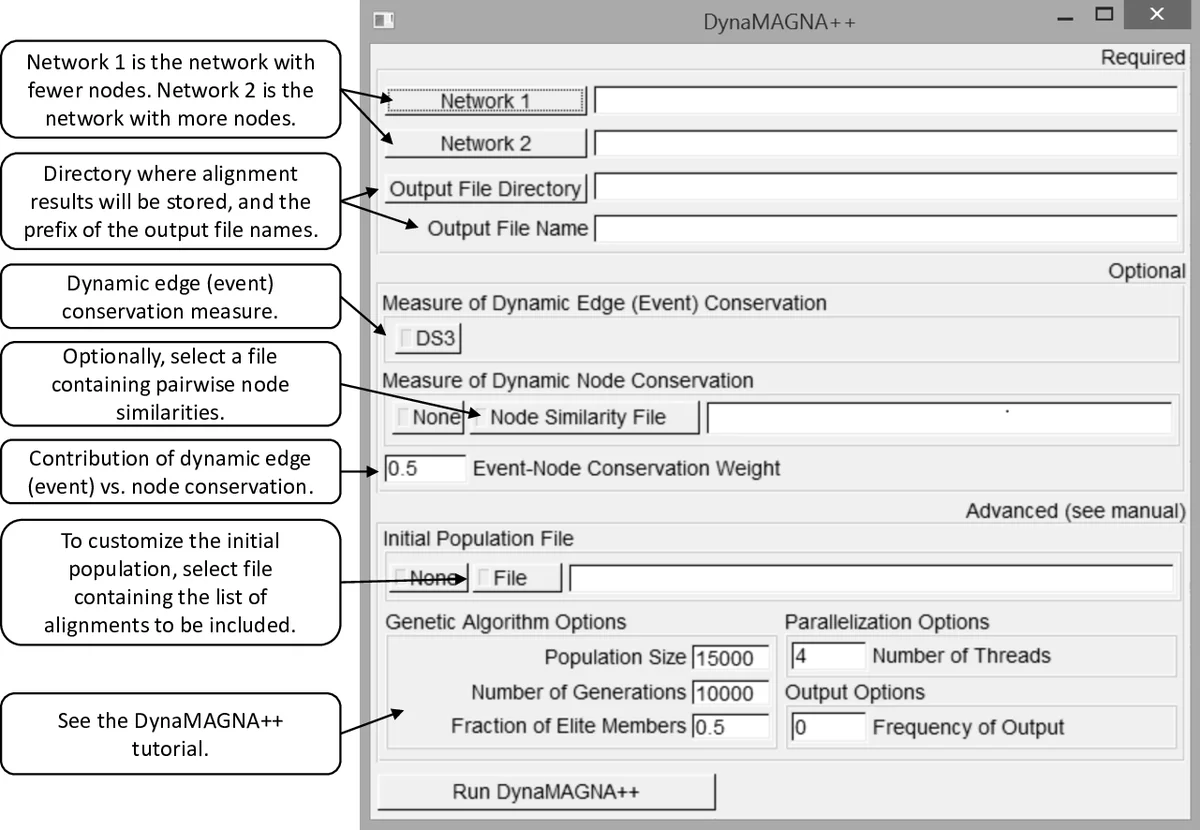
Implementation details include multi‑core parallelization for scalability and a user‑friendly graphical interface that allows non‑experts to load dynamic network files, adjust parameters, and visualize results.
Experimental evaluation covers three categories of data: (1) synthetic dynamic networks with controlled noise, (2) real protein‑protein interaction (PPI) networks from yeast and human that have been temporally resolved, and (3) social‑media networks (Twitter ↔ Facebook) where friendships appear and disappear over time. For each dataset, the authors compare DynaMAGNA++ against six established static aligners (MAGNA++, IsoRank, MI‑GRAAL, GHOST, NETAL, and WAVE) under identical initialization and parameter settings.
Results consistently show that DynaMAGNA++ achieves higher CET scores (average improvement ≈ 12 %) and lower NCET scores (≈ 9 % reduction), indicating superior temporal edge alignment. Functional validation on PPI data—using Gene Ontology term transfer—reveals an 8–11 % increase in prediction accuracy, while structural validation (shared clustering coefficient) improves by roughly 10 % across all domains. Runtime remains practical; thanks to parallelism, most experiments finish within one to two hours, comparable to static MAGNA++.
The authors discuss limitations: the current implementation handles only pairwise alignment, leaving multi‑network alignment as future work; the graphlet‑based dynamic node measure may be less effective for extremely rapid or noisy temporal data; and CET/NCET are sensitive to timestamp precision, requiring preprocessing for noisy datasets.
Future directions suggested include integrating dynamic graph neural networks to learn richer temporal similarity functions, extending the evolutionary search to multi‑objective frameworks for simultaneous alignment of many networks, and developing online algorithms for streaming‑data scenarios.
In summary, DynaMAGNA++ opens a new research avenue by demonstrating that explicitly modeling temporal dynamics yields measurable gains in alignment quality across biological and social domains, while providing a scalable and accessible tool for the community.
Comments & Academic Discussion
Loading comments...
Leave a Comment