Knowledge Transfer with Medical Language Embeddings
Identifying relationships between concepts is a key aspect of scientific knowledge synthesis. Finding these links often requires a researcher to laboriously search through scien- tific papers and databases, as the size of these resources grows ever l…
Authors: Stephanie L. Hyl, Theofanis Karaletsos, Gunnar R"atsch
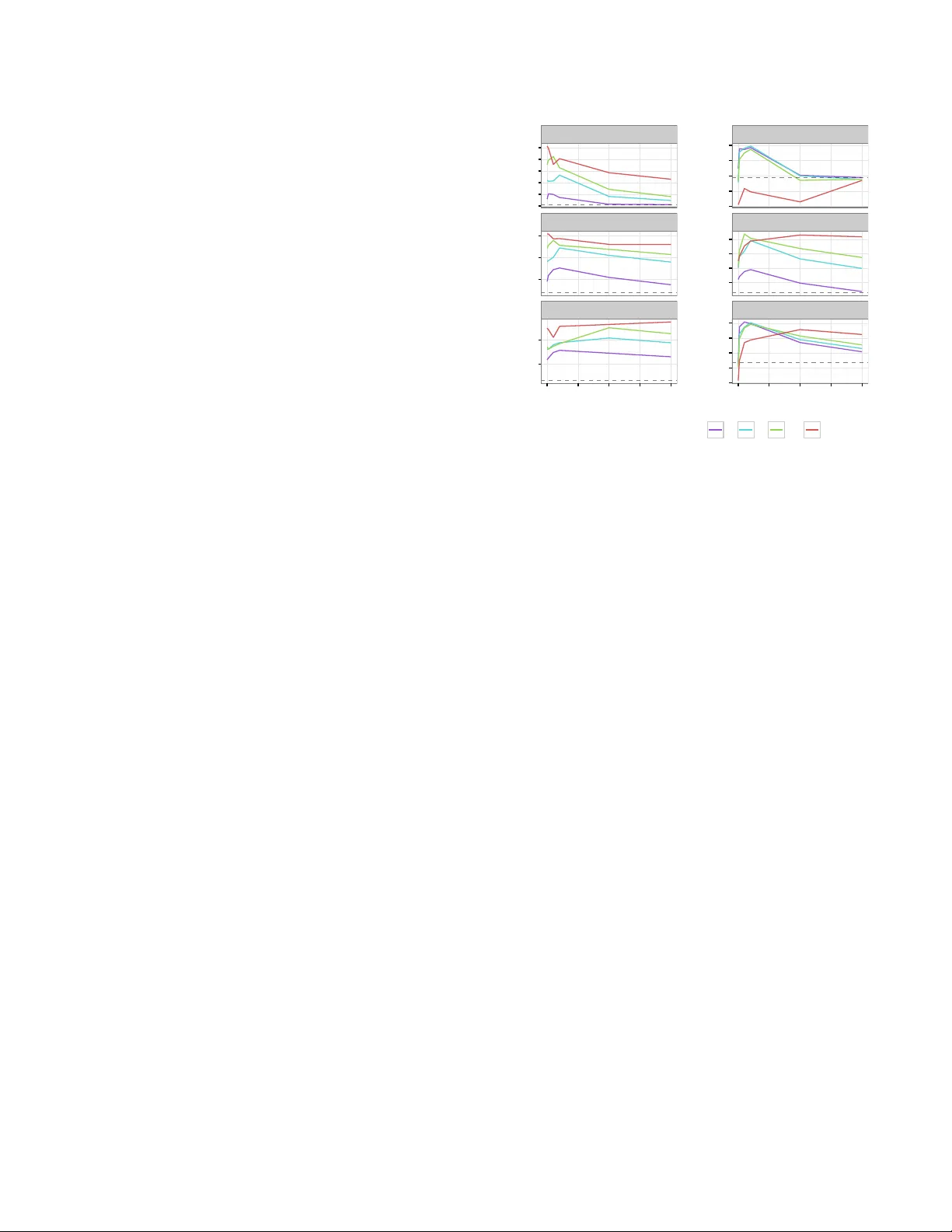
Kno wledge T ransfer with Medical Language Em b eddings ∗ Stephanie L. Hyland † ‡ Theofanis Karaletsos ‡ Gunnar R¨ atsc h ‡ Abstract Iden tifying relationships betw een concepts is a k ey aspect of scien tific knowledge syn thesis. Finding these links often requires a researcher to lab oriously searc h through scien- tific papers and databases, as the size of these resources gro ws ever larger. In this pap er we describe how distribu- tional seman tics can b e used to unify structured kno wledge graphs with unstructured text to predict new relationships b et w een medical concepts, using a probabilistic generative mo del. Our approac h is also designed to ameliorate data sparsit y and scarcit y issues in the medical domain, which mak e language mo delling more c hallenging. Sp ecifically , w e integrate the medical relational database (SemMedDB) with text from electronic health records (EHRs) to p erform kno wledge graph completion. W e further demonstrate the abilit y of our mo del to predict relationships b et ween tok ens not app earing in the relational database. 1 In tro duction The accelerating pace of scien tific progress presents both c hallenge and opportunity to researchers and health- care providers. Reading and comprehending the ever- gro wing b o dy of literature is a difficult but necessary part of kno wledge disco v ery and syn thesis. This is particularly imp ortan t for biomedical researc h, where therap eutic breakthroughs may rely on insights derived from disparate subfields. Curating literature at suc h breadth and scale is infeasible for individuals, neces- sitating the dev elopment of domain-sp ecific computa- tional approaches. W e present here a method using language emb e d- dings . Suc h an embedding is a representation of the tok ens of a language (such as words, or ob jects in a con trolled vocabulary) as elements of a vector space. Seman tic similarity is then captured b y v ector similar- it y , t ypically through Euclidean or cosine distance. The dimensionalit y of the space is typically m uch less than ∗ Supported by the Memorial Hospital and the Sloan Kettering Institute (MSKCC; to G.R.). Additional supp ort for S.L.H. was provided by the T ri-Institutional T raining Program in Computa- tional Biology and Medicine. † T ri-Institutional T raining Program in Computational Biology and Medicine, W eill Cornell Medical College ‡ Computational Biology Program, Memorial Sloan Kettering Cancer Center (MSKCC) the size of the vocabulary , so this pro cedure allo ws to- k ens to b e represen ted more compactly while also cap- turing semantics. Such represen tations can be used as features in downstream language-pro cessing tasks. In our case, we aim to exploit the embedding itself to dis- co ver new relationships b etw een tokens. This is p ossi- ble b ecause our embedding pro cedure defines a proba- bilit y distribution o v er tok en-relationship-tok en triples, allo wing for questions such as ‘is abdominal pain more lik ely to b e associated with acute appendicitis or pulmonary tuberculosis ?’, or ‘ho w is radium related to carcinoma ?’ 1 The tokens of in terest are chiefly Concept Unique Iden tifiers (CUIs) from the Unified Medical Lan- gauage System (UMLS) [3]. These represen t dis- crete me dic al c onc epts , which ma y require sev eral w ords to describ e, for example: C0023473: chronic myelogenous leukemia . W e consider it more mean- ingful and interesting to consider relationships b etw een CUIs rather than words themselves, when p ossible. W e exploit the exisence of SemMedDB [9], a database of se- man tic predications in the form of sub ject-relationship- ob ject triples, where the sub jects and ob jects are such CUIs. These w ere deriv ed from PubMed abstracts using the to ol SemRep [16]. W e combine this structured data with unstructured text consisting of clinical notes writ- ten b y physicians at Memorial Sloan Kettering Cancer Cen ter (MSK CC). 2 Related W ork Neural language mo dels [2] are an approach to learning em b eddings whic h use a w ord’s represen tation to pr e dict its surrounding context. This relies on the fact that w ords with similar meanings ha ve similar contexts (the distributional hypothesis of language [17]), which forces their represen tations to b e similar. Intriguingly , it was observ ed [13] [4] that the geometry of the resulting space preserv ed functional r elationships betw een terms. An example is a consistent offset v ector existing b et ween ‘Berlin’ and ‘German y’, and ‘Dublin’ and ‘Ireland’, seemingly represen ting the relationship capital city of country . This prop erty has b een exploited to p erform knowledge-base completion, for example [4] [18] 1 These are real examples from SemMedDB . [20], ho wev er these approac hes hav e restricted their atten tion to edge-disco very within a knowledge graph. T o extend suc h a graph we therefore developed a mo del [7] whic h can combine structured and unstructured data sources while explicitly mo delling the types of relationships present in the structured data. Despite the p opularity of language em b eddings in the broader natural language pro cessing (NLP) com- m unity , the biomedical domain has y et to fully exploit them. P edersen et al. [15] highlight the need to per- form domain-sp e cific NLP and discuss measures of se- man tic relatedness. Other recent applications include using representations of nominal elemen ts of the EHR to predict hospital readmission [11], identifying adverse drug reactions [6], and clinical concept extraction [8]. 3 Approac h 3.1 Mo del W e briefly describ e the bf mo del; see our earlier pap er [7] for more details. This is a probabilis- tic generative mo del ov er directed sub ject-relationship- ob ject triples ( S , R , O ). Sub ject and ob ject are b oth to- k ens from the v o cabulary (e.g., UMLS CUIs), although follo wing [12] and [5] we giv e them indep enden t repre- sen tations. This is formulated mathematically through an energy function, (3.1) E ( S, R, O | Θ) = − v O · G R c S k v O kk G R c S k En tities S and O are represented as v ectors, while each represen tation R corresp onds to an affine tr ansforma- tion on the vector space. Intuitiv ely , our energy func- tion is the cosine distance b et ween (the represen tation of ) O and S under the c ontext of R , where this context- sp ecific similarity is ac hieved by first transforming the represen tation of S by the affine transformation asso ci- ated to R . This energy function defines a Boltzmann probabil- it y distribution ov er ( S, R, O ) triples, (3.2) P ( S, R, O | Θ) = 1 Z (Θ) e −E ( S,R,O | Θ) where the denominator is the partition function, Z (Θ) = P s,r,o e −E ( s,r,o | Θ) . Equation 3.2 defines the probabilit y of observing a triple ( S, R , O ), given the em b edding Θ, whic h is the set of all vectors { c s , v o } s,o ∈ tokens and matrices { G r } r ∈ relationships . 3.2 T raining T o learn the em b edding (the parame- ters Θ consisting of all word v ectors c s , v o , and the relationship matrices G r ), we maximise the join t proba- bilit y of a set of true triples ( S, R, T ) under this mo del. Lik ely pairs hav e a high cosine similarity (low energy) in the context of their shared relationship, requiring simi- lar vector represen tations. W e employ stochastic max- im um lik eliho od for learning, approximating gradients of the partition function using p ersistent con trastive di- v ergence [19]. In all cases, w e p erform early stopping using a held- out v alidation set. The hyperparameters of the mo del are as follows: vector dimension is 100, batc h size is 100, w e use 3 rounds of Gibbs sampling to get mo del samples, of which w e maintain one p ersisten t Marko v chain. The learning rate is 0 . 001 and we use a l 2 regulariser with strength 0.01 on G r parameters. T o make learning more stable, we use Adam [10] with hyperparameters as suggested in the original paper. 3.3 Prediction Equation 3.2 defines a join t distri- bution ov er triples. Ho wev er, we are often interested in c onditional probabilities: given a pair of en tities S and O , whic h R most lik ely exists b etw een them (if any)? Suc h a distribution ov er R (or equiv alently S , O ) can easily be derived from the join t distribution, for exam- ple: (3.3) P ( R | S, O ; Θ) = e −E ( S,R,O | Θ) P r e −E ( S,r,O | Θ) The cost of calculating the conditional probability is at w orst linear in the size of the v o cabulary , as the (gen- erally intractable) partition function is not required. 4 Exp erimen ts 4.1 Data preparation W e train the model on tw o t yp es of data: unstructur e d ( EHR ) and structur e d ( SemMedDB ). The unstructured data is a corpus of de-iden tified clinical notes written by ph ysicians at MSK CC. W e pro cess raw text by replacing n umbers with generic tokens suc h as HEIGHT or YEAR , and re- mo ving most punctuation. In total, the corpus contains 99,334,543 sen tences, of which 46,242,167 are unique. This demonstrates the prev alence of terse language and sen tence fragmen ts in clinical text; for example the fragmen t no known drug allergies appears 192,334 times as a sen tence. W e identify CUIs in this text by greedily matching against strings asso ciated with CUIs (eac h CUI can hav e multiple such strings). This results in 45,402 unique CUIs, leaving 270,100 non-CUI w ord tok ens. W e note that the MetaMap [1] to ol is a more sophisticated approac h for this task, but found it to o inefficien t to use on a dataset of our size. T o generate ( S, R, O ) triples, we consider t wo words in a appears in a sentence with relationship if they are within a fiv e-word window of eac h other. The structured data ( SemMedDB ) consists of CUI - relationship - CUI statemen ts, for example C0027530(Neck) is LOCATION OF C0039979(Thoracic Duct) or C0013798(Electrocardiogram) DIAGNOSES C0026269(Mitral Valve Stenosis) . These w ere deriv ed from PubMed abstracts using SemRep [16]. SemMedDB contains 82,239,653 such statements, of whic h 16,305,000 are unique. This cov ers 237,269 unique CUIs. Since the distribution of CUI/token frequencies has a long tail in b oth data sources, we threshold tokens by their frequency . Firstly , tok ens (w ords of CUIs) must app ear at least 100 times in either dataset, and then at least 50 times in the pruned datasets. That is, in the first round we remov e sen tences (in EHR ) or statemen ts (in SemMedDB ) containing ‘rare’ tokens. In addition, the 58 relationships in SemMedDB also exhibit a long-tailed frequency distribution, so we retain only the top tw ent y . F rom this po ol of ( S, R, O ) triples (from EHR and SemMedDB ) we create fixed test sets (see next subsection) and smaller datasets with v arying relative abundances of eac h data type, using 0, 10, 50, 100, 500, and 1000 thousand training examples. The final list of tok ens has size W = 45 , 586, with 21 relationships: t wen ty from SemMedDB and an additional appears in sentence with from EHR . Of the W tok ens, 7,510 app ear in both data sources. These ov erlapping tok ens are critical to ensure embeddings derived from the kno wledge graph are consistent with those derived from the free text, allo wing information transfer. 4.2 Kno wledge-base completion Exp erimen tal design As the mo del defines con- ditional distributions for eac h element of a triple given the remaining tw o (Equation 3.3), we can test the abil- it y to predict new comp onen ts of a kno wledge graph. F or example, by selecting the b est R given S and O , w e predict the relationship (the type of edge) b et ween tok ens S and O . Without loss of generality , we describe the pro ce- dure for generating the test set for the R task. W e select a random set of S, O pairs app earing in the data. F or each pair, we record all entities r whic h app ear in a triple with them, removing these triples from the train- ing set. The S , O → { r i } i task is then recorded in the test set. Evidently , there may b e many correct com- pletions of a triple; in this case we exp ect the mo del to distribute probability mass across all answers. How b est to ev aluate this is task-dep endent; we consider both the r ank and the c ombine d pr ob ability mass in these exp er- imen ts. Results Figure 1 sho ws results for the task of predicting R given S and O . The mo del pro duces a ranking of all possible R s (high probability → low rank) and we rep ort the mean reciprocral rank of the ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● 0.60 0.65 0.70 0.75 0.80 10 100 # SemMedDB (structured) examples (thousands) mean reciprocal rank of correct relationship Model type ● ● ● ● ● ● ● ● 1ofN bf bf++ word2vec Figure 1: With more evidence from the kno wledge graph, the model better predicts the correct relationship for a given ( S , O ) pair. bf++ has an additional 100,000 triples from EHR : with little structured data, so m uch off- task information is harmful, but provides some b enefit when there is enough signal from the knowledge graph. Baselines are a random forest taking [ f ( S ) : f ( O )] as an input to predict the label R , where the feature represen tation f is either a 1-hot enco ding ( 1ofN ) or 200-dimensional word2vec v ectors trained on PubMed. 1ofN prov ed to o computationally exp ensive for large data. lowest-r anke d correct answ er ov er the test set. W e use this metric to ev aluate the utilit y of these predictions in prioritising hypotheses to test: w e would like any correct answ er to b e rank ed highly , and don’t apply a penalty for a failure to capture alternative answers. Results for our mo del are marked by bf 2 and bf++ . The latter model uses an additional 100,000 training examples from the EHR : these are ‘off-task’ information. As a baseline we consider a random forest trained to predict R given the concatenation [ f ( S ) : f ( O )], where the represen tation f is either: a) 1ofN : each tok en has a binary vector of length W ( W = 45,586), b) word2vec : eac h token has a 200-dimensional vector obtained by running word2vec [12] trained on PubMed [14]. W e note that the PubMed corpus con tains o ver 2 billions tok ens, far more data than was a v ailable to bf . W e additionally trained TransE [4] on this data, but it pro ved unsuited to the task (data not shown). As we can see, adding examples from SemMedDB impro ves performance for all mo del types, but bf seems to make b etter use of the additional data. In spite of 2 bf stands for ‘br ´ ı-focal’, which means word meaning in Irish. its very large input v ector size (2 W = 91172), 1ofN struggles, lik ely as it treats all tokens as independent en tities. W e note that for bf++ , performance is de gr ade d when the amoun t of structured data is low. This is consisten t with earlier observ ations on non-medical data [7], as the quantit y of ‘off-task’ information added is in this case comparable to that of ‘on-task’. Interestingly ho wev er, the mo del appears sligh tly b etter able to exploit more structured data when some ‘semantic bac kground’ is provided by EHR . 4.3 Information transfer Exp erimen tal design As mentioned, the mo del is capable of com bining structured and unstructured data. In [7] we observed that classification p erformance on a knowledge base could b e impro v ed by addition of unstructured data. How ever, the task in that case was quite ‘easy’; the model simply needed to differentiate b et ween true and false triples. Here w e consider the harder problem of correctly selecting which entit y w ould complete the triple. In addition to p ossibly improving performance, access to unstructured data pro vides the opp ortunit y to augment the knowlede base. That is, w e can predict relationships for tok ens not app e aring in SemMedDB . This uses the join t em b edding of all tokens in to one vector space, regardless of their data source. The geometric action of the relationships learned from SemMedDB can then b e applied to the representation of any token, suc h as those uniquely found in EHR . W e note that this pro cedure amounts to lab el tr ansfer from structured to unstructured examples, which can b e understo o d as a form of semi-supervised learning. T o generate ground truth for this task, we select some tok ens { T i } (these could appear as S or O en tities) found in b oth SemMedDB and EHR and remov e them from SemMedDB , recording them to use in the test set. Put another w ay , as in the previous setting, during the ‘random’ selection of S, O (still wlog) pairs, we mak e sure all of these recording them to use in the test set. Put another w ay , as in the previous setting, during the ‘random’ selection of S, O (still wlog) pairs, we mak e sure all T i in the deletion list are included, alongside an y other tok ens whic h appear in a SemMedDB -deriv ed relationship with them. The task is then to use purely semantic similarit y gleaned from EHR to place these tok ens in the embedding space such that the action of relationship op erators is still meaningful. Results Figure 2 shows results on all three tasks (predicting S , R , O giv en the remaining t wo), as a function of the typ e of test example . The righ t column of results is for test en tities in volving at least one element not app e aring in SemMedDB . As w e are no w interested ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● S, generic S, EHR−only R, generic R, EHR−only T , gener ic T , EHR−only 4 × 10 − 5 5 × 10 − 5 6 × 10 − 5 7 × 10 − 5 8 × 10 − 5 9 × 10 − 5 3.15 × 10 − 5 3.17 × 10 − 5 3.19 × 10 − 5 3.21 × 10 − 5 3.23 × 10 − 5 0.06 0.08 0.1 0.055 0.06 0.065 0.07 4 × 10 − 5 5 × 10 − 5 2.6 × 10 − 5 2.8 × 10 − 5 3.0 × 10 − 5 3.2 × 10 − 5 3.4 × 10 − 5 0 250 500 750 1000 0 250 500 750 1000 # EHR (unstructured) examples (thousands) probability mass of true entities # SemMedDB (thousands) ● ● ● ● 10 50 100 500 Figure 2: T otal probabilit y mass assigned to correct answ ers for all tasks. The right column sho ws results for test triples where at least one of S and O is found only in EHR , and therefore represen ts the know le dge tr ansfer setting. Information ab out relationships found in SemMedDB m ust b e transferred through the join t em b edding to enable these predictions. Grey dotted lines represent a random-guessing baseline. in the emb e ddings themselves w e rep ort the probabilit y mass of true entities, feeling this b etter captures the information contained in the embeddings. That is, it is no longer sufficien t for the mo del to correctly predict a single answer, we wan t it to assign appropriate probabilit y mass to al l correct answ ers. The dotted grey lines demonstrate the random baseline, where all tok ens are equally likely . The probability mass assigned by the baseline is therefore equal to k/W (or k/R ) where k is the a verage n umber of correct options in that task type. There are sev eral observ ations to be made here: • Most of the time, p erformance is best with a non- zero, but r elatively smal l amount of EHR data ( x - axis). This supports our observ ations that off-task information impro v es embeddings, but can ‘dro wn out’ signal if it dominates relative to the on-task examples. This can b e improv ed b y including a pre-factor on gradient contributions from the off- task data to adjust their con tribution relativ e to the structured examples, as demonstrated in our previous work [7]. • The EHR-only setting is muc h harder, as antici- pated. In the case of S and O it is comparable to the random baseline. F or R how ever, the mo del successfully assigns probabilit y mass when there is enough SemMedDB data a v ailable. • The S and O tasks are not symmetric. The S task features slightly more correct options on a verage than O (1.87 and 1.5 resp ectively , for the generic task), but this do es not accoun t for the difference in proportional performance relativ e baseline, espe- cially at low EHR abundance. A p ossible explana- tion is the energy function (Equation 3.1): it does not treat S -type and O -t yp e v ariables identically . Ho wev er, exp erimen ts using the F rob enius norm of G R in the denominator of E did not remov e asym- metry , so it is likely that the tasks are simply not equiv alent. This could arise due to bias in the di- rectionalit y of edges in the knowledge graph. W e conclude that it is possible to use the joint em- b edding pro cedure to predict R for pairs of S , O en tities ev en if they do not app ear in SemMedDB . F or the harder S and O tasks, the mo del generally succeeds in improv- ing visibly ov er baseline, but its assignmen ts are still quite ‘soft’. This ma y reflect premature stopping during training (most results rep orted w ere b efore 50 ep ochs had elapsed), an insufficien tly p o w erful model form ula- tion, or an excess of noise in the training data. Man y predicates in SemMedDB are v ague, and some relation- ships lend themselves to a one-to-many situation, for example part of , or location of . A core assumption in our mo del is that a token with fixed vector represen- tation can b e transformed by a single affine transforma- tion to be similar to its partner in a relationship. Many- to-one (or vice-versa) type relationships requires that m ultiple unique locations m ust be mapped to the same p oin t, which necessitates a rank-deficien t linear op er- ator or a more complex transformation function (one whic h is lo cally-sensitiv e, for example). F uture work in relational mo delling m ust carefully address the issue of man y-to-many and hierarc hical relationships. 5 Discussion Distributed language represen tations hav e seen limited application in healthcare to date, but present a p o- ten tially v ery p o w erful to ol for analysis and discov- ery . W e hav e demonstrated their use in knowledge syn thesis and text mining using a probabilistic gener- ativ e mo del which com bines structured and unstruc- tured data. These embeddings can further be used in do wnstream tasks, for example to reduce v ariation in language use b et ween doctors (by iden tifying and col- lapsing similar terms), for ‘fuzzy’ term-matc hing, or as inputs to c omp ositional approaches to represent larger structures such as sentences, documents, or even pa- tien ts. Expressive knowledge representations suc h as these will b e facilitate richer clinical data analysis in the future. References [1] Ar onson, A. R. Effective mapping of biomedical text to the umls metathesaurus: the metamap program. In Pr o c e e dings of the AMIA Symp osium (2001), American Medical Informatics Asso ciation, p. 17. [2] Bengio, Y., Ducharme, R., Vincent, P., and Janvin, C. A neural probabilistic language mo del. J. Mach. L earn. R es. 3 (Mar. 2003), 1137–1155. [3] Bodenreider, O. The unified medical language system (umls): integrating biomedical terminology . Nucleic A cids R ese ar ch 32 (2004), D267–D270. [4] Bordes, A., Usunier, N., Garcia-Duran, A., We- ston, J., and Y akhnenko, O. T ranslating embed- dings for modeling m ulti-relational data. In A dvanc es in Neur al Information Pr o c essing Systems (NIPS) (2013), pp. 2787–2795. [5] Goldber g, Y., and Levy, O. word2v ec explained: deriving mik olov et al.’s negative-sampling w ord- em b edding metho d. arXiv pr eprint (2014). [6] Henriksson, A. Represen ting clinical notes for ad- v erse drug even t detection. In Pr o c e e dings of the Sixth International Workshop on Health T ext Mining and In- formation Analysis (2015), Asso ciation for Computa- tional Linguistics, pp. 152–158. [7] Hyland, S. L., Karaletsos, T., and R ¨ atsch, G. A generative mo del of words and relationships from m ultiple sources. In Pr o c e e dings of the 30th AAAI Confer enc e on A rtificial Intel ligence (2016). [8] Jonnala gadda, S., Cohen, T., Wu, S., and Gon- zalez, G. Enhancing clinical concept extraction with distributional seman tics. Journal of biome dic al infor- matics 45 , 1 (2012), 129–140. [9] Kilicoglu, H., Shin, D., Fiszman, M., Rosembla t, G., and Rindflesch, T. C. Semmeddb: a pubmed- scale rep ository of biomedical semantic predications. Bioinformatics 28 , 23 (2012), 3158–3160. [10] Kingma, D., and Ba, J. Adam: A metho d for sto c hastic optimization. arXiv pr eprint arXiv:1412.6980 (2014). [11] Kr omp aß, D., Esteban, C., Tresp, V., Sedlma yr, M., and Ganslandt, T. Exploiting latent em b ed- dings of nominal clinical data for predicting hospital readmission. KI - K¨ unstliche Intel ligenz 29 , 2 (2014), 153–159. [12] Mik olov, T., Chen, K., Corrado, G., and Dean, J. Efficient estimation of word representations in v ector space. arXiv pr eprint arXiv:1301.3781 (2013). [13] Mik olov, T., Sutskever, I., Chen, K., Corrado, G. S., and Dean, J. Distributed representations of w ords and phrases and their comp ositionalit y . In A dvanc es in Neur al Information Pr o cessing Systems (NIPS) (2013), pp. 3111–3119. [14] Moen, S. P. F. G. H., and Ananiadou, T. S. S. Distributional semantics resources for biomedical text pro cessing. LBM (2013). [15] Pedersen, T., P akhomov, S. V., P a tw ardhan, S., and Chute, C. G. Measures of semantic similarity and relatedness in the biomedical domain. Journal of Biome dic al Informatics 40 , 3 (2007), 288 – 299. [16] Rindflesch, T. C., and Fiszman, M. The in terac- tion of domain knowledge and linguistic structure in natural language processing: interpreting h yp ern ymic prop ositions in biomedical text. Journal of Biome dic al Informatics 36 , 6 (2003), 462 – 477. Unified Medical Language System. [17] Sahlgren, M. The distributional hypothesis. Italian Journal of Linguistics 20 , 1 (2008), 33–53. [18] Socher, R., Chen, D., Manning, C. D., and Ng, A. Reasoning with neural tensor net works for knowledge base completion. In A dvances in Neur al Information Pr o c essing Systems (NIPS) (2013), pp. 926–934. [19] Tieleman, T. T raining restricted boltzmann ma- c hines using appro ximations to the lik eliho od gradi- en t. In International Conferenc e on Machine L e arning (ICML) (2008), pp. 1064–1071. [20] Weston, J., Bordes, A., Y akhnenko, O., and Usunier, N. Connecting language and knowledge bases with embedding mo dels for relation extraction. In Confer enc e on Empiric al Metho ds in Natur al L an- guage Pr o c essing (EMNLP) (2013), pp. 1366–1371.
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment