A convex model for non-negative matrix factorization and dimensionality reduction on physical space
A collaborative convex framework for factoring a data matrix $X$ into a non-negative product $AS$, with a sparse coefficient matrix $S$, is proposed. We restrict the columns of the dictionary matrix $A$ to coincide with certain columns of the data matrix $X$, thereby guaranteeing a physically meaningful dictionary and dimensionality reduction. We use $l_{1,\infty}$ regularization to select the dictionary from the data and show this leads to an exact convex relaxation of $l_0$ in the case of distinct noise free data. We also show how to relax the restriction-to-$X$ constraint by initializing an alternating minimization approach with the solution of the convex model, obtaining a dictionary close to but not necessarily in $X$. We focus on applications of the proposed framework to hyperspectral endmember and abundances identification and also show an application to blind source separation of NMR data.
💡 Research Summary
**
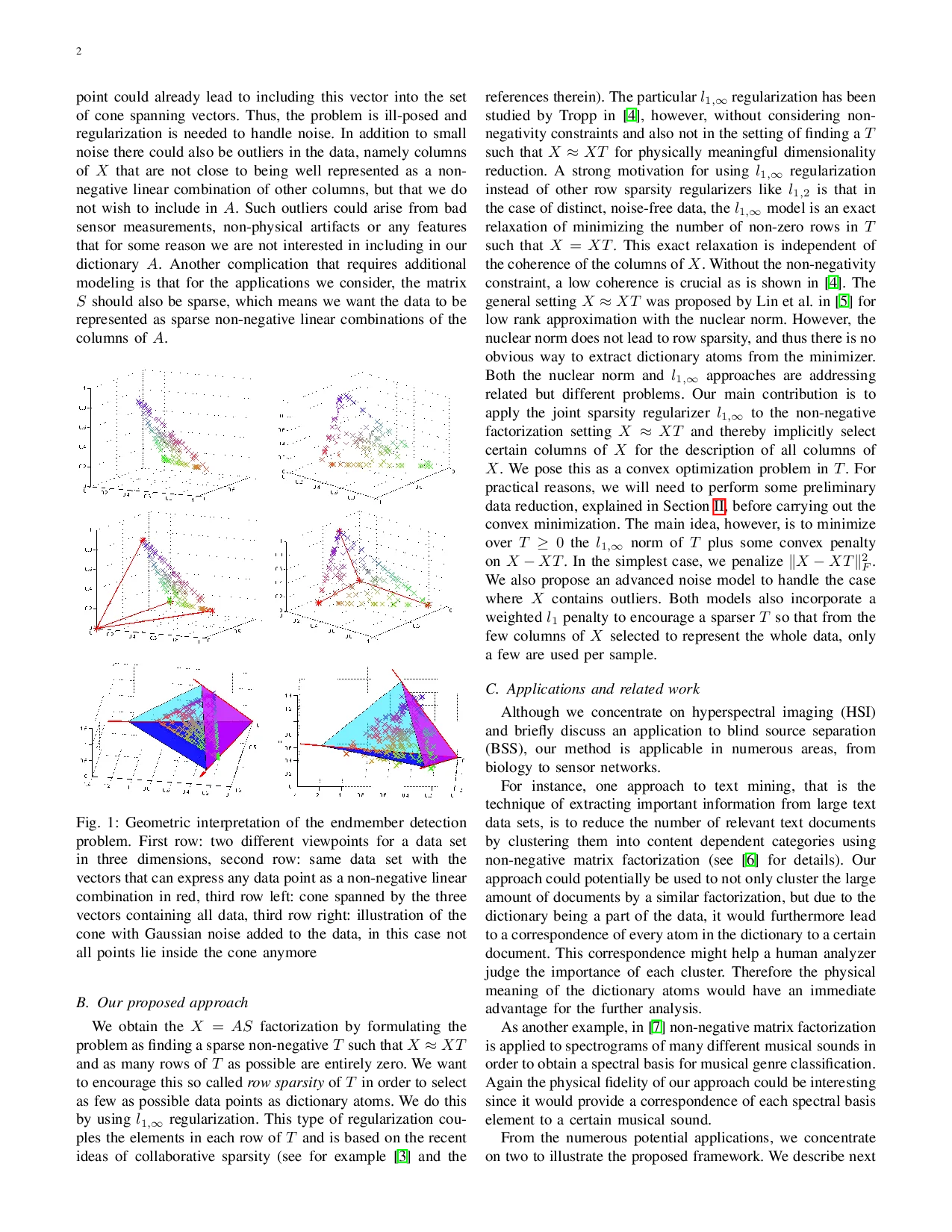
The paper introduces a novel convex framework for non‑negative matrix factorization (NMF) that simultaneously yields a physically meaningful dictionary and performs dimensionality reduction. The key idea is to enforce that the columns of the dictionary matrix A are selected directly from the data matrix X (or very close to them). Under the “pixel‑purity” assumption—each endmember (dictionary atom) appears as an exact column of X—the factorization X ≈ AS can be rewritten as a self‑representation X = X T with T ≥ 0. In this formulation, a zero row of T corresponds to a column of X that is not used as a dictionary atom; thus, minimizing the number of non‑zero rows of T (the row‑0 “norm”) directly selects the smallest possible subset of columns that span a cone containing all data points.
Because the row‑0 norm is combinatorial, the authors replace it with the convex (l_{1,\infty}) regularizer (|T|{1,\infty}= \sum_i \max_j |T{ij}|). They prove that for distinct, noise‑free data the (l_{1,\infty}) relaxation is exact: any optimal solution of the convex problem yields the same set of active rows as the combinatorial formulation, independent of column coherence. This is a significant advantage over other row‑sparsity penalties such as (l_{1,2}), which require low coherence for exactness.
Real data inevitably contain noise and outliers, so the authors augment the basic model with two additional terms: (i) a Frobenius‑norm data‑fidelity term (|X - X T|_F^2) that balances reconstruction error against sparsity, and (ii) a weighted (l_1) penalty on the entries of T to encourage sparsity within each active row (i.e., each pixel should be expressed by only a few selected endmembers). An outlier‑robust variant further down‑weights columns with large residuals, effectively removing them from the dictionary selection process.
The resulting optimization problem is convex in T and is solved efficiently using an ADMM‑based algorithm. Because T is a d × d matrix (d = number of pixels or spectra), the authors first apply a preprocessing dimensionality‑reduction step (e.g., clustering or random sampling) to reduce the problem size to a manageable k × k submatrix, where k ≪ d.
Two application domains are explored in depth:
-
Hyperspectral Imaging (HSI) – In HSI each pixel records a high‑dimensional spectrum that is often a non‑negative mixture of a few material signatures (endmembers). The pixel‑purity assumption is realistic for moderate spatial resolution images, allowing the dictionary atoms to be actual measured spectra. The proposed method automatically selects a minimal set of endmembers and computes sparse abundance maps S. Experiments on synthetic data and real AVIRIS scenes show that the convex model recovers the correct endmembers with fewer atoms and lower reconstruction error than classic algorithms such as VCA, N‑FINDR, and standard NMF. The sparsity‑enhanced version further improves abundance map interpretability.
-
Blind Source Separation (BSS) of NMR data – The same linear mixing model X = AS applies, with non‑negativity reflecting physical concentration constraints. By assuming each source dominates at least one measurement (analogous to pixel purity), the method extracts source spectra directly from the observed mixtures. Results demonstrate accurate source recovery even in the presence of moderate noise, outperforming ICA and conventional NMF approaches.
Theoretical analysis extends to the noisy case, providing stability guarantees: the (l_{1,\infty}) regularizer still promotes row sparsity, and the balance between the fidelity term and the regularization parameters can be tuned to achieve a desired trade‑off between reconstruction accuracy and dictionary size. Computational complexity is shown to scale linearly with the number of selected atoms after preprocessing, making the approach feasible for large‑scale hyperspectral cubes.
In summary, the paper makes three major contributions: (1) a convex formulation of NMF that enforces dictionary atoms to be actual data columns, guaranteeing physical interpretability; (2) proof that (l_{1,\infty}) regularization yields an exact relaxation of the row‑0 sparsity objective for distinct, noise‑free data; and (3) practical extensions for noisy and outlier‑contaminated data, together with an efficient ADMM solver and compelling demonstrations on hyperspectral unmixing and NMR source separation. This work bridges the gap between mathematically tractable convex optimization and the need for physically meaningful, sparse representations in real‑world signal processing applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment