Dynamical Systems on Networks: A Tutorial
We give a tutorial for the study of dynamical systems on networks. We focus especially on “simple” situations that are tractable analytically, because they can be very insightful and provide useful springboards for the study of more complicated scenarios. We briefly motivate why examining dynamical systems on networks is interesting and important, and we then give several fascinating examples and discuss some theoretical results. We also briefly discuss dynamical systems on dynamical (i.e., time-dependent) networks, overview software implementations, and give an outlook on the field.
💡 Research Summary
The paper “Dynamical Systems on Networks: A Tutorial” serves as a comprehensive introductory guide for researchers interested in studying dynamical processes that unfold on complex network structures. Its primary aim is to present “simple” yet analytically tractable scenarios that illuminate how network topology influences dynamics, thereby providing a springboard for tackling more intricate problems.
The authors begin by motivating the study of dynamics on networks, highlighting applications ranging from epidemic spreading and synchronization of oscillators to meme propagation and opinion formation. They argue that the non‑trivial connectivity of real‑world networks—characterized by heterogeneous degree distributions, clustering, and degree‑degree correlations—fundamentally shapes the temporal evolution of any process defined on the nodes or edges.
A concise review of basic network concepts follows. The paper adopts the simplest setting: unweighted, undirected, static graphs represented by a symmetric adjacency matrix A with zero diagonal. Key descriptors such as the degree k, degree distribution P(k), mean degree z, and heavy‑tailed (power‑law) distributions are defined. The configuration model is introduced as a null ensemble that preserves P(k) while randomizing edge placement, allowing researchers to isolate the effects of degree heterogeneity from higher‑order structures like clustering. The authors note that real networks often display assortative or disassortative mixing and significant triangle motifs, which are absent in the pure configuration model, and they point out that studying dynamics on clustered graphs is an active research frontier.
The core of the tutorial is a series of illustrative dynamical systems, each chosen for analytical accessibility.
-
Percolation – The authors discuss site, bond, and K‑core percolation, explaining how the emergence of a giant connected component (GCC) signals a phase transition at a critical occupation probability pc. They describe the iterative pruning algorithm that defines K‑cores and connect it to cascade processes. “Explosive” percolation (Achlioptas processes) is presented as a case where the GCC appears abruptly, illustrating how rule‑based edge addition can dramatically alter critical behavior.
-
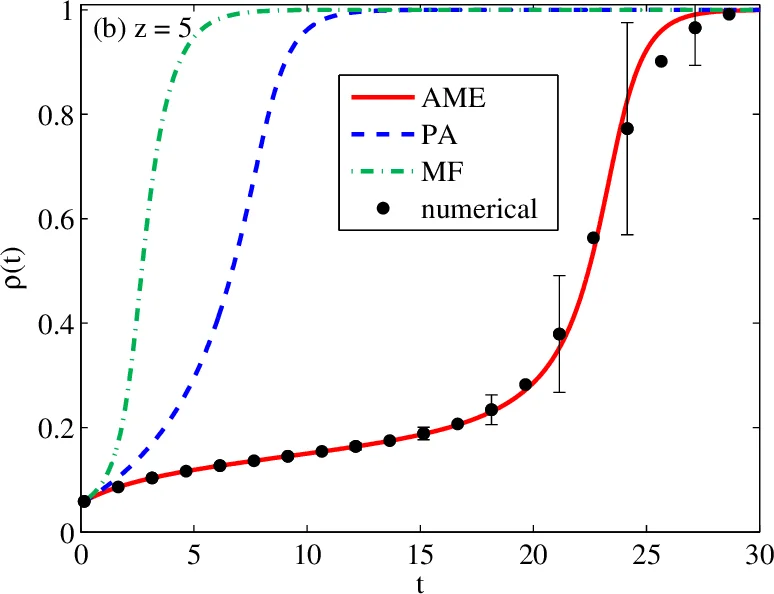
Binary‑state dynamics – Classic compartmental epidemic models (SIR, SIS, SEIR) are recast in a mean‑field framework, yielding low‑dimensional ODEs for the fractions of susceptible, infected, and recovered individuals. The tutorial emphasizes how degree heterogeneity modifies the basic reproduction number and the epidemic threshold. The authors also cover threshold models of social contagion (e.g., the Centola‑Macy model) and discuss synchronous versus asynchronous updating schemes, noting that update order can affect convergence and stability.
-
Continuous‑state dynamics – Synchronization of phase oscillators (Kuramoto model) is treated as a prototypical example. The authors show that the second smallest eigenvalue λ2 of the graph Laplacian governs the onset of global synchronization, linking spectral graph theory to dynamical stability. Diffusive processes described by the Laplacian (e.g., heat flow, opinion averaging) are also examined, with attention to how clustering and community structure can create metastable patterns.
Throughout these sections, the tutorial provides analytical results (e.g., percolation thresholds for random graphs, epidemic thresholds derived from heterogeneous mean‑field theory) and highlights where approximations (pairwise, moment‑closure) become necessary.
A brief software survey follows. The authors list widely used open‑source libraries such as Python’s NetworkX for graph handling, EoN (Epidemics on Networks) for stochastic epidemic simulations, and Julia’s Graphs.jl for high‑performance computations. Sample code snippets illustrate how to generate configuration‑model networks, integrate mean‑field ODEs, and run event‑driven Gillespie simulations on large graphs.
The tutorial then turns to dynamical (time‑dependent) networks. Adaptive networks are introduced, where node states feed back onto edge creation or deletion (e.g., individuals avoiding contacts when ill). The authors discuss the co‑evolution of dynamics and topology, noting that the relative timescales of the two processes determine whether a quasi‑static approximation is valid. Extensions to multilayer and interdependent networks are mentioned, with references to recent work on percolation across layers and cascading failures.
In the concluding section, the authors outline open problems: rigorous derivation of critical thresholds for clustered and directed graphs, understanding multistability in synchronization on heterogeneous networks, and integrating real‑time data streams into adaptive network models. They advocate for a synthesis of analytical theory, large‑scale simulation, and data‑driven inference to advance the field.
Overall, the paper delivers a well‑structured, pedagogical roadmap that equips newcomers with the essential mathematical tools, conceptual frameworks, and computational resources needed to explore dynamical systems on networks, while also pointing toward the most promising avenues for future research.
Comments & Academic Discussion
Loading comments...
Leave a Comment