Lack of confidence in ABC model choice
Approximate Bayesian computation (ABC) have become a essential tool for the analysis of complex stochastic models. Earlier, Grelaud et al. (2009) advocated the use of ABC for Bayesian model choice in the specific case of Gibbs random fields, relying …
Authors: Christian P. Robert (University Paris-Dauphine), Jean-Marie Cornuet (INRA, Montpellier)
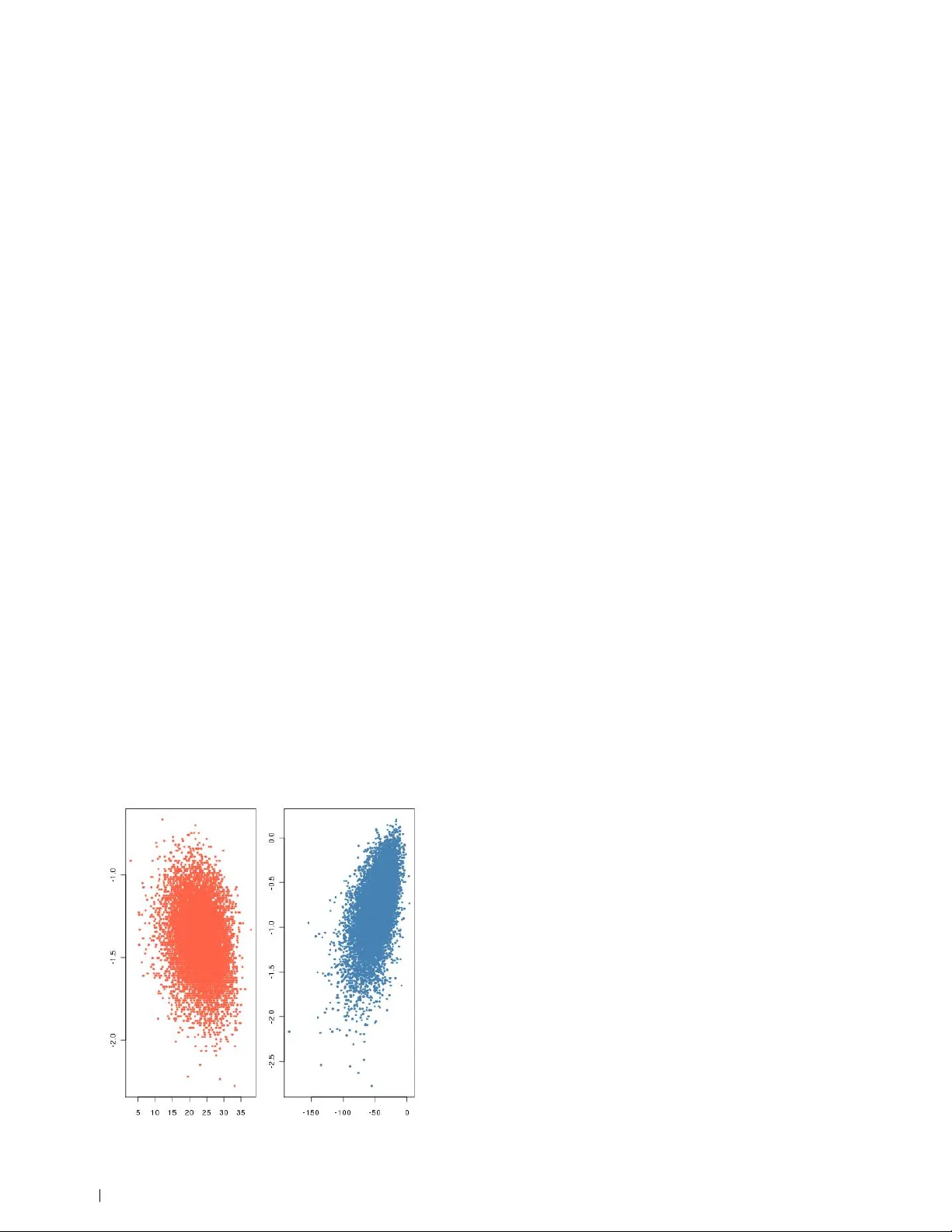
Lack of confidence in ABC mo del choice Christian P . Rob ert, ∗ † ‡ Jean-Ma rie Co rnuet, § Jean-Michel Ma rin, ¶ and Natesh S. Pillai k ∗ Universit´ e Pa ris Dauphine, † Institut Universitaire de France, ‡ CREST, Pa ris, § CBGP , INRA, Montp ellier, France, ¶ I3M, UMR CNRS 5149, Universit´ e Montpellier 2, France, and k Department of Statistics, Harva rd University , Cambridge, MA Submitted to Proceedings of the National Academy of Sciences of the United States of America Appro ximate Bayesian computation (ABC) have b ecome an essen- tial to ol for the analysis of complex sto chastic mo dels. Grelaud et al. (2009, Ba y esian Ana 3:427–442) advo cated the use of ABC fo r model choice in the sp ecific case of Gibbs random fields, relying on a inter-mo del sufficiency property to show that the appro ximation was legitimate. We implemented ABC mo del choice in a wide range of phylogenetic models in the DIY-ABC soft w a re (Cornuet et al. (2008) Bioinfo 24:2713–2719). We now present arguments as to why the theoretical arguments for ABC model choice are missing, since the algorithm involves an unknown loss of information induced by the use of insufficient summary statistics. The appro ximation error of the p osterior probabilities of the models under compa rison may thus b e unrelated with the computational effort spent in run- ning an ABC algo rithm. We then conclude that additional empirical verifications of the p erformances of the ABC p rocedure as those available in DIY ABC a re necessa ry to conduct model choice. 1 likelihood-free metho ds | Bay es factor | Bayesian mo del choice | sufficiency Abbreviations: ABC, appro ximate Bayesian computation; ABC-MC, ABC model choice; DIY-ABC, Do-it-yourself ABC; IS, imp ortance sampling; SMC, sequential Monte Carlo I nference on p opulation genetic mo dels such as coalescent trees is one representativ e example of cases when statistical analyses lik e Ba y esian inference cannot easily op erate b ecause the likelihoo d function asso ciated with the data cannot b e computed in a manageable time (T a v ar´ e et al., 1997; Beau- mon t et al., 2002; Cornuet et al., 2008). The fundamental reason for this imp ossibility is that the mo del asso ciated with coalescen t data has to integrate ov er trees of high complexity . In such settings, traditional approximation to ols like Mon te Carlo sim ulation (Robert and Casella, 2004) from the p osterior distribution are unav ailable for practical purp oses. Indeed, due to the complexity of the latent structures defin- ing the likelihoo d (lik e the coalescent tree), their simulation is to o unstable to bring a reliable approximation in a man- ageable time. Suc h complex mo dels call for a practical if cruder approximation metho d, the ABC metho dology (T a v ar´ e et al., 1997; Pritchard et al., 1999). This rejection tec hnique b ypasses the computation of the lik eliho o d via simulations from the corresp onding distribution (see Beaumont, 2010 and Lop es and Beaumont, 2010 for recent surveys, and Csill` ery et al., 2010a for the wide and successful arra y of applications based on implementations of ABC in genomics and ecology). W e argue here that ABC is a generally v alid approxima- tion metho d for doing Ba yesian inference in complex models. Ho wev er, without further justification, ABC methods cannot b e trusted to discriminate b etw een tw o comp eting mo dels when based on insufficient summary statistics. W e exhibit simple examples in whic h the information loss due to insuffi- ciency leads to inconsistency , i.e. when the ABC mo del selec- tion fails to reco ver the true mo del, even with infinite amounts of observ ation and computation. On the one hand, ABC using the entire data leads to a consisten t mo del choice decision but it is clearly infeasible in most settings. On the other hand, to o muc h information loss due to insuffiency leads to a statisti- cally inv alid decision pro cedure. The c hallenge is in achieving a balance b et ween information loss and consistency . Theo- retical results that mathematically v alidate mo del choice for insufficien t statistics are curren tly lac king on a general basis. Our conclusion at this stage is to opt for a cautionary approac h in ABC mo del c hoice, handling it as an exploratory to ol rather than trusting the Ba yes factor appro ximation. The corresp onding degree of approximation cannot b e ev aluated, except via Monte Carlo ev aluations of the mo del selection p erformances of ABC. More empirical measures such as those prop osed in the DIY-ABC soft ware (Corn uet et al., 2008) and in Ratmann et al. (2009) thus seem to b e the only av ailable solution at the curren t time for conducting model comparison. W e stress that, while T empleton (2008, 2010) rep eatedly expressed reserv ations ab out the formal v alidity of the ABC approac h in statistical testing, those criticisms were rebutted in Beaumon t et al. (2010); Csill` ery et al. (2010b); Berger et al. (2010) and are not relev ant for the current pap er. Statistical Metho ds The ABC algorithm. The setting in whic h ABC operates is the appro ximation of a simulation from the p osterior distribution π ( θ | y ) ∝ π ( θ ) f ( y | θ ) when distributions associated with b oth the prior π and the likelihoo d f can b e simulated (the later b eing unav ailable in closed form). The first ABC algorithm w as introduced by Pritchard et al. (1999) as follows: giv en a sample y from a sample space D , a sample ( θ 1 , . . . , θ M ) is pro duced by Algorithm 1: ABC sampler for i = 1 to N do rep eat Generate θ 0 from the prior distribution π ( · ) Generate z from the likelihoo d f ( ·| θ 0 ) un til ρ { η ( z ) , η ( y ) } ≤ set θ i = θ 0 , end for The parameters of the ABC algorithm are the so-called summary statistic η ( · ), the distance ρ {· , ·} , and the tolerance lev el > 0. The appro ximation of the p osterior distribution π ( θ | y ) pro vided by the ABC sampler is to instead sample from the marginal in θ of the joint distribution π ( θ , z | y ) = π ( θ ) f ( z | θ ) I A , y ( z ) R A , y × Θ π ( θ ) f ( z | θ )d z d θ , where I B ( · ) denotes the indicator function of B and A , y = { z ∈ D| ρ { η ( z ) , η ( y ) } ≤ } . The basic justification of the ABC approximation is that, when using a sufficien t statistic η and a small (enough) toler- ance , we hav e π ( θ | y ) = Z π ( θ , z | y )d z ≈ π ( θ | y ) . 1 CPR, JMM and NSP designed and p erformed resea rch, JMC and JMM analysed data, and CPR, JMC and JMM wrote the pap er. www.pnas.org/cgi/doi/10.1073/pnas.xxx PNAS Issue Date Volume Issue Number 1 – 10 In practice, the statistic η is necessarily insufficien t (since only exponential families enjo y sufficient statistics with fixed dimension, see Lehmann and Casella, 1998) and the approxi- mation then conv erges to the less informative π ( θ | η ( y )) when go es to zero. This loss of information is a necessary price to pa y for the access to computable quan tities and π ( θ | η ( y )) pro vides a conv ergent inference on θ when θ is identifiable in the distribution of η ( y ) (F earnhead and Prangle, 2010). While ac knowledging the gain brought by ABC in handling Ba yesian inference in complex mo dels, and the existence of in- v olved summary selection mechanisms (Joyce and Marjoram, 2008; Nunes and Balding, 2010), we demonstrate here that the loss due to the ABC approximation may b e arbitrary in the sp ecific setting of Ba y esian mo del c hoice via p osterior mo del probabilities. ABC mo del choice. The standard Bay esian to ol for mo del comparison is the marginal likelihoo d (Jeffreys, 1939) w ( y ) = Z Θ π ( θ ) f ( y | θ ) d θ , whic h leads to the Bay es factor for comparing the evidences of models with likelihoo ds f 1 ( y | θ 1 ) and f 2 ( y | θ 2 ), B 12 ( y ) = w 1 ( y ) w 2 ( y ) = R Θ 1 π 1 ( θ 1 ) f 1 ( y | θ 1 ) d θ 1 R Θ 2 π 2 ( θ 2 ) f 2 ( y | θ 2 ) d θ 2 . As detailed in Beaumont et al. (2010), it pro vides a v alid cri- terion for mo del comparison that is naturally p enalised for mo del complexity . Ba yesian mo del c hoice proceeds b y creating a probabilit y structure across M mo dels (or likelihoo ds). It in tro duces the mo del index M as an extra unknown parameter, associated with its prior distribution, π ( M = m ) ( m = 1 , . . . , M ), while the prior distribution on the parameter is conditional on the v alue m of the M index, denoted by π m ( θ m ) and defined on the parameter space Θ m . The choice b etw een those mo dels is then driv en b y the p osterior distribution of M , P ( M = m | y ) = π ( M = m ) w m ( y ) X k π ( M = k ) w k ( y ) where w k ( y ) denotes the marginal likelihoo d for mo del k . While this posterior distribution is straigh tforw ard to in- terpret, it offers a c hallenging computational conundrum in Ba yesian analysis. When the likelihoo d is not av ailable, ABC represen ts the almost unique solution. Pritchard et al. (1999) describ e the use of mo del choice based on ABC for distin- guishing betw een different m utation mo dels. The justification b ehind the metho d is that the a verage ABC acceptance rate asso ciated with a given mo del is prop ortional to the p osterior probabilit y corresp onding to this approximativ e model, when iden tical summary statistics, distance, and tolerance level are used ov er all mo dels. In practice, an estimate of the ratio of marginal likelihoo ds is given by the ratio of observed accep- tance rates. Using Bay es formula, estimates of the p osterior probabilities are straightforw ard to derive. This approac h has b een widely implemented in the literature (see, e.g., Estoup et al., 2004, Miller et al., 2005, Pascual et al., 2007, and Sain- udiin et al., 2011). A represen tativ e illustration of the use of an ABC mo del c hoice approac h is given b y Miller et al. (2005) whic h analyses the Europ ean inv asion of the w estern corn ro otw orm, North America’s most destructiv e corn p est. Because this p est was initially introduced in Central Europ e, it was b elieved that subsequen t outbreaks in W estern Europ e originated from this area. Based on an ABC mo del c hoice analysis of the genetic v ariability of the ro otw orm, the authors conclude that this b elief is false: There ha ve b een at least three independent in- tro ductions from North America during the past tw o decades. The ab ov e estimate is improv ed by regression regularisa- tion (F agundes et al., 2007), where model indices are pro- cessed as categorical v ariables in a p olychotomous regression. When comparing tw o mo dels, this inv olves a standard logis- tic regression. Rejection-based approac hes w ere lately intro- duced by Cornuet et al. (2008), Grelaud et al. (2009) and T oni et al. (2009), in a Monte Carlo simulation of mo del in- dices as w ell as model parameters. Those recen t extensions are already widely used in p opulation genetics, as exempli- fied by Belle et al. (2008); Cornuet et al. (2010); Excoffier et al. (2009); Ghirotto et al. (2010); Guillemaud et al. (2009); Leuen b erger and W egmann (2010); Patin et al. (2009); Ra- makrishnan and Hadly (2009); V erdu et al. (2009); W egmann and Excoffier (2010). Another illustration of the p opularit y of this approach is giv en by the a v ailability of four soft wares implemen ting ABC mo del c hoice methodologies: • ABC-SysBio, whic h relies on a SMC-based ABC for infer- ence in system biology , including mo del-choice (T oni et al., 2009). • ABCT o olb ox whic h prop oses SMC and MCMC implemen- tations, as well as Ba yes factor approximation (W egmann et al., 2011). • DIY ABC, whic h relies on a regularised ABC-MC algorithm on p opulation history using molecular mark ers (Cornuet et al., 2008). • P opABC, which relies on a regular ABC-MC algorithm for genealogical simulation (Lop es et al., 2009). As exp osed in e.g. Grelaud et al. (2009), T oni and Stumpf (2010), or Didelot et al. (2011), once M is incorporated within the parameters, the ABC approximation to its p osterior fol- lo ws from the same principles as in regular ABC. The corre- sp onding implementation is as follo ws, using for the summary statistic a statistic η ( z ) = { η 1 ( z ) , . . . , η M ( z ) } that is the con- catenation of the summary statistics used for all models (with an ob vious elimination of duplicates). Algorithm 2: ABC-MC for i = 1 to N do rep eat Generate m from the prior π ( M = m ) Generate θ m from the prior π m ( θ m ) Generate z from the mo del f m ( z | θ m ) un til ρ { η ( z ) , η ( y ) } ≤ Set m ( i ) = m and θ ( i ) = θ m end for The ABC estimate of the p osterior probability π ( M = m | y ) is then the frequency of acceptances from mo del m in the ab ov e sim ulation ˆ π ( M = m | y ) = N − 1 P N i =1 I m ( i ) = m . This also corresp onds to the frequency of simulated pseudo- datasets from mo del m that are closer to the data y than the tolerance . In order to improv e the estimation b y smo oth- ing, Cornuet et al. (2008) follo w the rationale that motiv ated the use of a local linear regression in Beaumon t et al. (2002) and rely on a weigh ted polychotomous regression to estimate π ( M = m | y ) based on the ABC output. This mo delling is implemen ted in the DIY ABC softw are. The difficult y with ABC-MC. There is a fundamental discrep- ancy b etw een the genuine Bay es factors/posterior probabili- ties) and the appro ximations resulting from ABC-MC. 2 www.pnas.org/cgi/doi/10.1073/pnas.xxx Footline Author The ABC approximation to a Bay es factor, B 12 sa y , re- sulting from Algorithm 2 is d B 12 ( y ) = π ( M = 2) N X i =1 I m ( i ) =1 π ( M = 1) N X i =1 I m ( i ) =2 . An alternativ e represen tation is given by d B 12 ( y ) = π ( M = 2) π ( M = 1) P T t =1 I m t =1 I ρ { η ( z t ) , η ( y ) }≤ P T t =1 I m t =2 I ρ { η ( z t ) , η ( y ) }≤ , where the pairs ( m t , z t ) are simulated from the joint prior and T is the num b er of simulations necessary for N acceptances in Algorithm 2. In order to study the limit of this approxima- tion, we first let T go to infinit y . (F or simplification purp oses and without loss of generality , we c ho ose a uniform prior on the model index.) The limit of d B 12 ( y ) is then B 12 ( y ) = P [ M = 1 , ρ { η ( z ) , η ( y ) } ≤ ] P [ M = 2 , ρ { η ( z ) , η ( y ) } ≤ ] = R R I ρ { η ( z ) , η ( y ) }≤ π 1 ( θ 1 ) f 1 ( z | θ 1 ) d z d θ 1 R R I ρ { η ( z ) , η ( y ) }≤ π 2 ( θ 2 ) f 2 ( z | θ 2 ) d z d θ 2 = R R I ρ { η , η ( y ) }≤ π 1 ( θ 1 ) f η 1 ( η | θ 1 ) d η d θ 1 R R I ρ { η , η ( y ) }≤ π 2 ( θ 2 ) f η 2 ( η | θ 2 ) d η d θ 2 , where f η 1 ( η | θ 1 ) and f η 2 ( η | θ 2 ) denote the densities of η ( z ) when z ∼ f 1 ( z | θ 1 ) and z ∼ f 2 ( z | θ 2 ), respectively . By L’Hospital formula, if goes to zero, the ab ov e con verges to B η 12 ( y ) = Z π 1 ( θ 1 ) f η 1 ( η ( y ) | θ 1 ) d θ 1 Z π 2 ( θ 2 ) f η 2 ( η ( y ) | θ 2 ) d θ 2 , namely the Bay es factor for testing mo del 1 versus mo del 2 based on the sole observ ation of η ( y ). This result reflects the curren t p ersp ective on ABC: the inference derived from the ideal ABC output when = 0 only uses the information con tained in η ( y ). Thus, in the limiting case, i.e. when the al- gorithm uses an infinite computational p ow er, the ABC o dds ratio do es not accoun t for features of the data other than the v alue of η ( y ), whic h is why the limiting Bay es factor only dep ends on the distribution of η under b oth models. When running ABC for p oin t estimation, the use of an insufficien t statistic do es not usually jeopardise conv ergence of the metho d. As sho wn, e.g., in (F earnhead and Prangle, 2010, Theorem 2), the noisy version of ABC as an inference metho d is conv ergent under usual regularit y conditions for mo del-based Bay esian inference (Bernardo and Smith, 1994), including identifiabilit y of the parameter for the insufficient statistic η . In contrast, the loss of information induced by η may seriously impact mo del-choice Bay esian inference. In- deed, the information contained in η ( y ) is lesser than the information con tained in y and this ev en in most cases when η ( y ) is a sufficient statistic for b oth mo dels . In other words, η ( y ) b eing sufficient for b oth f 1 ( y | θ 1 ) and f 2 ( y | θ 2 ) do es not usual ly imply that η ( y ) is sufficient for { m, f m ( y | θ m ) } . T o see why this is the case, consider the most fav ourable case, namely when η ( y ) is a sufficient statistic for b oth mo dels. W e then ha ve by the factorisation theorem (Lehmann and Casella, 1998) that f i ( y | θ i ) = g i ( y ) f η i ( η ( y ) | θ i ) ( i = 1 , 2), i.e. B 12 ( y ) = w 1 ( y ) w 2 ( y ) = R Θ 1 π ( θ 1 ) g 1 ( y ) f η 1 ( η ( y ) | θ 1 ) d θ 1 R Θ 2 π ( θ 2 ) g 2 ( y ) f η 2 ( η ( y ) | θ 2 ) d θ 2 = g 1 ( y ) R π 1 ( θ 1 ) f η 1 ( η ( y ) | θ 1 ) d θ 1 g 2 ( y ) R π 2 ( θ 2 ) f η 2 ( η ( y ) | θ 2 ) d θ 2 = g 1 ( y ) g 2 ( y ) B η 12 ( y ) . [ 1 ] Th us, unless g 1 ( y ) = g 2 ( y ), as in the sp ecial case of Gibbs ran- dom fields detailed b elow, the t wo Bay es factors differ by the ratio g 1 ( y ) /g 2 ( y ), which is only equal to one in a very small n umber of known cases. This decomp osition is a straightfor- w ard pro of that a mo del-wise sufficient statistic is usually not sufficien t across mo dels, hence for mo del comparison. An im- mediate corollary is that the ABC-MC approximation do es not alw ays conv erge to the exact Bay es factor. The discrepancy b etw een limiting ABC and genuine Ba yesian inferences does not come as a surprise, b ecause ABC is indeed an approximation metho d. Users of ABC algorithms are therefore prepared for some degree of imprecision in their final answer, a p oint stressed by Wilkinson (2008) and F earn- head and Prangle (2010) when they qualify ABC as exact inference on a wrong mo del. Ho wev er, the magnitude of the difference b etw een B 12 ( y ) and B η 12 ( y ) expressed b y [ 1 ] is suc h that there is no direct connection b etw een b oth answers. In a general setting, if η has the same dimension as one comp onent of the n components of y , the ratio g 1 ( y ) /g 2 ( y ) is equiv alent to a density ratio for a sample of size O( n ), hence it can b e arbitrarily small or arbitrarily large when n gro ws. Contrast- ingly , the Bay es factor B η 12 ( y ) is based on an equiv alen t to a single observ ation, hence do es not necessarily conv erge with n to the correct limit, as sho wn by the Poisson and normal examples b elo w and in SI. The conclusion derived from the ABC-based Bay es factor may therefore completely differ from the conclusion derived from the exact Ba yes factor and there is no p ossibility of a generic agreement b etw een b oth, or even of a manageable correction factor. This discrepancy means that a theoretical v alidation of the ABC-based mo del choice pro cedure is curren tly missing and that, due to this absence, p oten tialy costly simulation-based assessments are required when calling for this pro cedure. Therefore, users must b e warned that ABC approxima- tions to Bay es factors do not p erform as standard numerical or Mon te Carlo approximations, with the exception of Gibbs random fields detailed in the next section. In all cases when g 1 ( y ) /g 2 ( y ) differs from one, no inference on the true Bay es factor can b e derived from the ABC-MC approximation with- out further information on the ratio g 1 ( y ) /g 2 ( y ), most often una v ailable in settings where ABC is necessary . Didelot et al. (2011) also derived this relation betw een b oth Bay es factors in their form ula [18] . While they still ad- v o cate the use of ABC mo del c hoice in the absence of sufficien t statistic, we stress that no theoretical guarantee can b e giv en on the v alidity of the ABC approximation to the Bay es factor and hence of its use as a model c hoice procedure. Note that Sousa et al. (2009) resort to full allelic distri- butions in an ABC framework, instead of chosing summary statistics. They show how to apply ABC using allele frequen- cies to draw inferences in cases where selecting suitable sum- mary statistics is difficult (and where the complexity of the mo del or the size of dataset prohibits to use full-likelihoo d metho ds). In such settings, ABC-MC do es not suffer from the div ergence exhibited here because the measure of distance do es not inv olve a reduction of the sample. The same com- men t applies to the ABC-SysBio soft ware of T oni et al. (2009), whic h relies on the whole dataset. The theoretical v alidation of ABC inference in hidden Marko v mo dels by Dean et al. (2011) should also extend to the mo del choice setting b ecause the approach do es not rely on summary statistics but instead on the whole sequence of observ ations. Results Footline Author PNAS Issue Date Volume Issue Number 3 The sp ecific case of Gibbs random fields. In an apparen t con- tradiction with the abov e, Grelaud et al. (2009) show ed that the computation of the p osterior probabilities of Gibbs ran- dom fields under competition can b e done via ABC tech- niques, which provide a con verging appro ximation to the true Ba yes factor. The reason for this result is that, for these mo dels in the ab o ve ratio [ 1 ] , g 1 ( y ) = g 2 ( y ). The v alidation of an ABC comparison of Gibbs random fields is thus that their sp ecific structure allows for a sufficient statistic vector that runs across models and therefore leads to an exact (when = 0) simulation from the p osterior probabilities of the mo d- els. Eac h Gibbs random field mo del has its o wn sufficient statistic η m ( · ) and Grelaud et al. (2009) exp osed the fact that the vector of statistics η ( · ) = ( η 1 ( · ) , . . . , η M ( · )) is also suffi- cien t for the join t parameter ( M , θ 1 , . . . , θ M ). Didelot et al. (2011) p oint out that this sp ecific property of Gibbs random fields can b e extended to any exp onential family (hence to any setting with fixed-dimension sufficient statistics, see Lehmann and Casella, 1998). Their argument is that, by including all sufficient statistics and all dominating measure statistics in an encompassing mo del, mo dels under comparison are submo dels of the encompassing mo del. The concatenation of those statistics is then jointly sufficien t across mo dels. While this encompassing principle holds in full gen- eralit y , in particular when comparing mo dels that are already em b edded, we think it leads to an o verly optimistic persp ec- tiv e ab out the merits of ABC for mo del c hoice: in practice, most complex mo dels do not enjoy sufficien t statistics (if only b ecause they are b eyond exp onen tial families). The Gibbs case processed by Grelaud et al. (2009) therefore happens to b e one of the v ery few realistic coun terexamples. As demon- strated in the next section and in the normal example in SI, using insufficient statistics is more than a mere loss of infor- mation. Lo oking at what happens in the limiting case when one relies on a common mo del-wise sufficient statistic is a for- mal but useful study since it brings light on the potentially h uge discrepancy b etw een the ABC-based and the true Bay es factors. T o develop a solution to the problem in the formal case of the exponential families do es not help in understanding the discrepancy for non-exponential mo dels. Fig. 1. Comparison between the true log-Bay es factor (first axis) for the com- parison of a P oisson mo del versus a negative binomial mo del and of the log-Bayes factor based on the sufficient statistic P i y i (se c ond axis) , for P oisson (left) and negative binomial (left) samples of size n = 50 , based on T = 10 4 replications Arbitra ry ratios. The difficulty with the discrepancy b etw een B 12 ( y ) and B η 12 ( y ) is that this discrepancy is imp ossible to ev aluate in a general setting, while there is no reason to ex- p ect a reasonable agreement b etw een both quan tities. A first illustration was pro duced by Marin et al. (2011) in the case of MA( q ) models. A simple illustration of the discrepancy due to the use of a mo del-wise sufficient statistic is a a sample y = ( y 1 , . . . , y n ) that could come either from a Poisson P ( λ ) distribution or from a geometric G ( p ) distribution, already in tro duced in Grelaud et al. (2009) as a coun ter-example to Gibbs ran- dom fields and later repro cessed in Didelot et al. (2011) to supp ort their sufficiency argument. In this case, the sum S = P n i =1 y i = η ( y ) is a sufficient statistic for b oth mo dels but not across mo dels. The distribution of the sample given S is a multinomial M ( S, 1 /n, . . . , 1 /n ) distribution when the data is Poisson, while it is the uniform distribution ov er the y ’s such that P i y i = S in the geometric case, since S is then a negativ e binomial N eg ( n, p ) v ariable. The discrepancy ratio is therefore g 1 ( y ) /g 2 ( y ) = n + S − 1 S ! S ! n − S Y i y i ! . When simulating n P oisson or geometric v ariables and us- ing prior distributions as exp onential λ ∼ E (1) and uniform p ∼ U (0 , 1) on the parameters of the resp ective mo dels, the exact Bay es factor is av ailable and the distribution of the dis- crepancy is therefore av ailable. Fig 1 giv es the range of B 12 ( y ) v ersus B η 12 ( y ), sho wing that B η 12 ( y ) is then unrelated with B 12 ( y ): the v alues pro duced by b oth approac hes hav e noth- ing in common. As noted ab ov e, the approximation B η 12 ( y ) based on the sufficient statistic S is pro ducing figures of the magnitude of a single observ ation, while the true Bay es factor is of the order of the sample size. The discrepancy betw een both Ba yes factors is in fact in- creasing with the sample size, as sho wn by the following result: Lemma 1. Consider mo del sele ction b etwe en mo del 1: P ( λ ) with prior distribution π 1 ( λ ) e qual to an exponential E (1) dis- tribution and mo del 2: G ( p ) with a uniform prior distribution π 2 when the observe d data y c onsists of iid observations with exp e ctation E [ y i ] = θ 0 > 0 . Then S ( y ) = P n i =1 y i is the min- imal sufficient statistic for b oth mo dels and the Bayes factor b ase d on the sufficient statistic S ( y ) , B η 12 ( y ) , satisfies lim n →∞ B η 12 ( y ) = θ − 1 0 ( θ 0 + 1) 2 e − θ 0 a.s. Therefore, the Bay es factor based on the statistic S ( y ) is not consistent; it con verges to a non-zero, finite v alue. In this sp ecific setting, Didelot et al. (2011) show that adding P = Q i y i ! to the S creates a statistic ( S, P ) that is sufficien t across b oth mo dels. While this is mathematically correct, it do es not provide further understanding of the b e- ha viour of ABC-model choice in realistic settings: outside for- mal examples as the one ab ov e and well-structured if complex exp onen tial families lik e Gibbs random fields, it is not p ossi- ble to devise completion mechanisms that ensure sufficiency across mo dels, or even select well-discriminating statistics. It is therefore more fruitful to study and detect the diverging b eha viour of the ABC approximation as given, rather than attempting at solving the problem in a sp ecific and formal case. P opulation genetics. W e recall that ABC has first b een in- tro duced by p opulation geneticists (Beaumont et al., 2002; Pritc hard et al., 1999) for statistical inference about the evo- lutionary history of sp ecies, b ecause no lik eliho o d-based ap- proac h existed apart from very simple and hence unrealistic 4 www.pnas.org/cgi/doi/10.1073/pnas.xxx Footline Author situations. This approac h has since b een used in an increasing n umber of biological studies (Estoup et al., 2004; Estoup and Clegg, 2003; F agundes et al., 2007), most of them including mo del choice. It is therefore crucial to get insights in the v alid- it y of suc h studies, particularly when they deal with species of economical or ecological imp ortance (see, e.g., Lom baert et al., 2010). T o this end, we need to compare ABC estimates of p osterior probabilities to reliable likelihoo d-based estimates. Com bining differen t mo dules based on Stephens and Donnelly (2000), it is p ossible to approximate the likelihoo d of p opu- lation genetic data through imp ortance sampling (IS) even in complex scenarios. In order to ev aluate the p otential dis- crepancy b etw een ABC-based and likelihoo d-based p osterior ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● 0.0 0.2 0.4 0.6 0.8 1.0 0.0 0.2 0.4 0.6 0.8 1.0 Importance Sampling estimates of P(M=1|y) ABC estimates of P(M=1|y) Fig. 2. Comparison of IS and ABC estimates of the p osterior probabilit y of scena rio 1 in the first p opulation genetic exp eriment, using 24 summary statistics ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● 0.0 0.2 0.4 0.6 0.8 1.0 0.0 0.2 0.4 0.6 0.8 1.0 Importance Sampling estimates of P(M=1|y) ABC estimates of P(M=1|y) Fig. 3. Same caption as Fig 2 when using 15 summary statistics ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● 0.0 0.2 0.4 0.6 0.8 1.0 0.0 0.2 0.4 0.6 0.8 1.0 Importance Sampling estimates of P(M=1|y) ABC estimates of P(M=1|y) Fig. 4. Comparison of IS and ABC estimates of the p osterior probabilit y of scena rio 1 in the second p opulation genetic exp eriment probabilities of evolutionary scenarios, we designed tw o ex- p erimen ts based on simulated data with limited information con tent, so that the c hoice b etw een scenarios is problematic. This setting th us provides a wide enough set of in termediate v alues of model posterior probabilities, in order to b etter ev al- uate the divergence b etw een ABC and lik eliho o d estimates. In the first exp erimen t, we consider tw o p opulations (1 and 2) having diverged at a fixed time in the past and a third p opulation (3) ha ving diverged from one of those tw o p opu- lations (scenarios 1 and 2 resp ectively). Times are set to 60 generations for the first divergence and to 30 generations for the second div ergence. One h undred pseudo observed datasets ha ve b een simulated, represen ted b y 15 diploid individuals p er p opulation genotyped at five independent microsatellite loci. These lo ci are assumed to evolv e according to the strict Step- wise Mutation mo del (SMM), i.e. when a mutation occurs, the n umber of repeats of the m utated gene increases or decreases ● ● ● 1 2 3 4 5 0.4 0.5 0.6 0.7 0.8 0.9 Fig. 5. Boxplots of the p osterior p robabilities evaluated over 10 independent Monte Carlo evaluations, for five indep endent simulated datasets in the first p opula- tion genetic experiment ● ● 1 2 3 4 5 0.4 0.5 0.6 0.7 0.8 0.9 Fig. 6. Boxplots of the p osterior p robabilities evaluated over 10 independent Monte Carlo evaluations, for five independent simulated datasets in the second p op- ulation genetic experiment Footline Author PNAS Issue Date Volume Issue Number 5 T ab. S1 Summary statistics used in the p opulation genetic exp eriments, the Subset column corresponding to the ABC op erated with 15 summary statistics and the last three statistics being only used in this reduced collection Name Subset Definition NAL1 yes average numb er of alleles in population 1 NAL2 yes average numb er of alleles in population 2 NAL3 yes average numb er of alleles in population 3 HET1 y es average heterozygothy n p opulation 1 HET2 y es average heterozygothy n p opulation 2 HET3 y es average heterozygothy n p opulation 3 V AR1 y es average va riance of the allele size in p opulation 1 V AR2 y es average va riance of the allele size in p opulation 2 V AR3 y es average va riance of the allele size in p opulation 3 MGW1 no Ga rza-Williamson M in p opulation 1 MGW2 no Ga rza-Williamson M in p opulation 2 MGW3 no Ga rza-Williamson M in p opulation 3 FST1 no average FST in p opulation 1 FST2 no average FST in p opulation 2 FST3 no average FST in p opulation 3 LIK12 no probab ility that sample 1 is from population 1 LIK13 no probab ility that sample 1 is from population 3 LIK21 no probab ility that sample 2 is from population 1 LIK23 no probab ility that sample 2 is from population 3 LIK31 no probab ility that sample 3 is from population 1 LIK32 no probab ility that sample 3 is from population 2 D AS12 y es shared allele distance betw een p opulations 1 and 2 D AS13 y es shared allele distance betw een p opulations 1 and 3 D AS23 y es shared allele distance betw een p opulations 2 and 3 DM212 y es distance ( δµ ) 2 b et ween populations 1 and 2 DM213 y es distance ( δµ ) 2 b et ween populations 1 and 3 DM223 y es distance ( δµ ) 2 b et ween populations 2 and 2 b y one unit with equal probabilit y . The m utation rate, com- mon to all five lo ci, has b een set to 0 . 005 and effective p op- ulation sizes to 30. In this exp eriment, b oth scenarios hav e a single parameter: the effective population size, assumed to b e identical for all three p opulations. W e chose a uniform prior U [2 , 150] for this parameter (the true v alue b eing 30). The IS algorithm w as p erformed using 100 coalescent trees p er particle. The marginal likelihoo d of b oth scenarios has b een computed for the same set of 1000 particles and they provide the p osterior probabilit y of each scenario. The ABC compu- tations hav e b een performed with DIY ABC (Corn uet et al., 2008). A reference table of 2 million datasets has b een sim- ulated using 24 usual summary statistics (provided in T able S1) and the p osterior probability of each scenario has b een estimated as their prop ortion in the 500 simulated datasets closest to the pseudo observed one. This p opulation genetic ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ●● ● ● ● ● ● ● ● ● ● ● ● ● ●● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● 0.0 0.2 0.4 0.6 0.8 1.0 0.0 0.2 0.4 0.6 0.8 1.0 ABC estimates of P(M=1|y) Importance Sampling estimates of P(M=1|y) with 50,000 par ticles Fig. 7. Comparison b etw een tw o appro ximations of the p osterior probabilities of scenario 1 based on imp ortance sampling with 50,000 particles (first axis) and ABC (se c ond axis) for the larger p opulation genetic exp eriment setting do es not allow for a choice of sufficient statistics, even at the mo del lev el. The second exp eriment also opp oses tw o scenarios includ- ing three p opulations, tw o of them having div erged 100 gen- erations ago and the third one resulting of a recen t admixture b et w een the first tw o p opulations (scenario 1) or simply di- v erging from population 1 (scenario 2) at the same time of 5 generations in the past. In scenario 1, the admixture rate is 0 . 7 from p opulation 1. Pseudo observed datasets (100) of the same size as in exp eriment 1 (15 diploid individuals per p opu- lation, 5 indep endent microsatellite lo ci) ha ve been generated for an effective p opulation size of 1000 and mutation rates of 0 . 0005. In contrast with exp eriment 1, analyses included the following 6 parameters (provided with corresponding pri- ors): admixture rate ( U [0 . 1 , 0 . 9]), three effective p opulation sizes ( U [200 , 2000]), the time of admixture/second div ergence ( U [1 , 10]) and the time of the first divergence ( U [50 , 500]). T o accoun t for an higher complexity in the scenarios, the IS algo- rithm w as performed with 10,000 coalescen t trees p er particle. Apart from this change, b oth ABC and likelihoo d analyses ha ve b een performed in the same wa y as exp eriment 1. Fig 2 shows a reasonable fit betw een the exact p osterior probabilit y of mo del 1 (ev aluated b y IS) and the ABC approx- imation in the first exp eriment on most of the 100 sim ulated datasets, even though the ABC approximation is biased to- w ards 0 . 5. When using 0 . 5 as the decision b oundary betw een mo del 1 and mo del 2, there is hardly any discrepancy b etw een b oth approaches, demonstrating that mo del choice based on ABC can be trusted in this case. Fig 3 considers the same set- ting when mo ving from 24 to 15 summary statistics (given in T able S1): the fit somehow degrades. In particular, the num- b er of opp osite conclusions in the mo del choice mo ves to 12%. In the more complex setting of the second experiment, the dis- crepancy w orsens, as shown on Fig 4. The num b er of opposite conclusions reac hes 26% and the fit betw een both v ersions of the p osterior probabilities is considerably degraded, with a correlation co efficient of 0 . 643. 6 www.pnas.org/cgi/doi/10.1073/pnas.xxx Footline Author The v alidity of the imp ortance sampling approximation can ob viously be questioned in b oth experiments, how ever Fig 5 and Fig 6 displa y a strong stability of the posterior probabilit y IS approximation across 10 indep endent runs for 5 different datasets and gives prop er confidence in this ap- proac h. Increasing the num b er of lo ci to 50 and the sample size to 100 individuals p er p opulation (see SI) leads to p os- terior probabilities of the true scenario ov erwhelmingly close to one (Fig 7), thus bluring the distinction b etw een ABC and lik eliho o d based estimates but also reassuring on the ability of ABC to provide the right choice of mo del with a higher information conten t of the data. Actually , we note that, for this exp eriment, all ABC-based decisions conclude in fav our of the correct model. Discussion Since its introduction b y T av ar´ e et al. (1997) and Pritchard et al. (1999), ABC has been extensiv ely used in several areas in volving complex lik eliho o ds, primarily in population genet- ics, b oth for p oint estimation and testing of hypotheses. In realistic settings, with the exception of Gibbs random fields, whic h satisfy a resilience prop ert y with resp ect to their suf- ficien t statistics, the conclusions dra wn on mo del comparison cannot b e trusted p er se but require further simulations anal- yses as to the p ertinence of the (ABC) Bay es factor based on the summary statistics. This pap er has examined in de- tails only the case when the summary statistics are sufficient for b oth mo dels, while practical situations imply the use of insufficien t statistics. The rapidly increasing num ber of appli- cations estimating p osterior probabilities b y ABC indicates a clear need for further ev aluations of the w orth of those esti- mations. F urther research is needed for producing trustw orthy ap- pro ximations to the p osterior probabilities of mo dels. At this stage, unless the whole data is inv olv ed in the ABC approx- imation as in Sousa et al. (2009), our conclusion on ABC- based mo del c hoice is to exploit the approximations in an exploratory manner as measures of discrepancies rather than gen uine p osterior probabilities. This direction relates with the analyses found in Ratmann et al. (2009). F urthermore, a version of this exploratory analysis is already provided in the DIY-ABC softw are of (Cornuet et al., 2008). An option in this soft ware allows for the computation of a Monte Carlo ev aluation of false allo cation rates resulting from using the ABC posterior probabilities in selecting a model as the most lik ely . F or instance, in the setting of both our population ge- netic experiments, DIY-ABC gives false allocation rates equal to 20% (under scenarios 1 and 2) and 14 . 5% and 12 . 5% (un- der scenarios 1 and 2), resp ectively . This ev aluation obviously shifts a wa y from the performances of ABC as an appro xima- tion to the posterior probabilit y to wards the p erformances of the whole Ba yesian apparatus for selecting a mo del, but this nonetheless represents a useful and manageable quality assess- men t for practitioners. ACKNO WLEDGMENTS. The first three authors’ wo rk has been pa rtly supported by Agence Nationale de la Recherche via the 2009–2013 project EMILE. They are grateful to the reviewers and to Michael Stumpf for their comments. Computations were p erformed on the INRA CBGP and MIGALE clusters. References Beaumont, M. (2010). Approximate Ba yesian computation in evo- lution and ecology . Annual R eview of Ec olo gy, Evolution, and Systematics , 41 379–406. Beaumont, M. , Nielsen, R. , Rober t, C. , Hey, J. , Gaggiotti, O. , Knowles, L. , Estoup, A. , Mahesh, P. , Coranders, J. , Hickerson, M. , Sisson, S. , F agundes, N. , Chikhi, L. , Beerli, P. , Vit alis, R. , Cornuet, J.-M. , Huelsenbeck, J. , F oll, M. , Y ang, Z. , Rousset, F. , Balding, D. and Ex coffier, L. (2010). In defense of mo del-based inference in ph ylogeography . Mole cu- lar Ecolo gy , 19(3) 436–446. Beaumont, M. , Zhang, W. and Balding, D. (2002). Approxi- mate Bay esian computation in population genetics. Genetics , 162 2025–2035. Belle, E. , Benazzo, A. , Ghirotto, S. , Colonna, V. and Barbu- jani, G. (2008). Comparing mo dels on the genealogical relation- ships among Neandertal, Cro-Magnoid and mo dern Europ eans by serial coalescent simulations. Here dity , 102 218–225. Berger, J. , Fienberg, S. , Rafter y, A. and Rober t, C. (2010). Incoherent phylogeographic inference. Pro c. Nat. A c ad. Sci. USA , 107 E57. Bernardo, J. and Smith, A. (1994). Bayesian The ory . John Wi- ley , New Y ork. Cornuet, J.-M. , Ra vign ´ e, V. and Estoup, A. (2010). Inference on p opulation history and model c hecking using DNA sequence and microsatellite data with the soft ware DIY ABC (v1.0). BMC Bioinformatics , 11 401. Cornuet, J.-M. , Santos, F. , Beaumont, M. A. , Rober t, C. P. , Marin, J.-M. , Balding, D. J. , Guillemaud, T. and Estoup, A. (2008). Inferring p opulation history with DIY ABC: a user- friendly approac h to Appro ximate Ba yesian Computation. Bioin- formatics , 24 2713–2719. Csill ` er y, K. , Blum, M. , Gaggiotti, O. and Franc ¸ ois, O. (2010a). Appro ximate Ba yesian computation (ABC) in practice. T r ends in Ec olo gy and Evolution , 25 410–418. Csill ` er y, K. , Blum, M. , Gaggiotti, O. and Franc ¸ ois, O. (2010b). Inv alid arguments against ABC: A reply to A.R. T em- pleton. T r ends in Ecolo gy and Evolution , 25 490–491. Dean, T. , Singh, S. , Jasra, A. and Peters, G. (2011). Parame- ter estimation for hidden mark ov mo dels with in tractable likeli- hoo ds. T ech. rep. Didelot, X. , Everitt, R. , Johansen, A. and La wson, D. (2011). Likelihood-free estimation of model evidence. Bayesian Analysis , 6 1–28. Estoup, A. , Bea umont, M. , Sennedot, F. , Moritz, C. and Cor- nuet, J. (2004). Genetic analysis of complex demographic sce- narios: spatially expanding populations of the cane toad, Bufo Marinus. Evolution , 58 2021–2036. Estoup, A. and Clegg, S. (2003). Bay esian inferences on the re- cent island colonization history by the bird Zosterops lateralis lateralis. Mol. Ec ol. , 12 657–674. Excoffier, C. , Leuenberger, D. and Wegmann, L. (2009). Bay esian computation and model selection in p opulation genet- ics. F agundes, N. , Ra y, N. , Beaumont, M. , Neuenschw ander, S. , Salzano, F. , Bona tto, S. and Ex coffier, L. (2007). Statisti- cal ev aluation of alternative models of human ev olution. Pr oc. Nat. Ac ad. Sci. USA , 104 17614–17619. Fearnhead, P. and Prangle, D. (2010). Semi-automatic approx- imate Bay esian computation. Ghirotto, S. , Mona, S. , Benazzo, A. , P ap arazzo, F. , Caramelli, D. and Barbujani, G. (2010). Inferring genealogical processes from patterns of bronze-age and mo dern DNA v aria- tion in Sardinia. Mol. Biol. Evol. , 27 875–886. Grelaud, A. , Marin, J.-M. , Rober t, C. , Rodolphe, F. and T all y, F. (2009). Lik elihoo d-free methods for mo del choice in Gibbs random fields. Bayesian Analysis , 3(2) 427–442. Guillemaud, T. , Beaumont, M. , Ciosi, M. , Cornuet, J.-M. and Estoup, A. (2009). Inferring introduction routes of inv asive species using appro ximate Bay esian computation on microsatel- lite data. Here dity , 104 88–99. Jeffreys, H. (1939). The ory of Pr ob ability . 1st ed. The Clarendon Press, Oxford. Joyce, P. and Marjoram, P. (2008). Appro ximately sufficient statistics and Bayesian computation. Statistic al Applic ations in Genetics and Mole cular Biolo gy , 7 article 26. Lehmann, E. and Casella, G. (1998). The ory of Point Estimation (r evise d e dition) . Springer-V erlag, New Y ork. Footline Author PNAS Issue Date Volume Issue Number 7 Leuenberger, C. and Wegmann, D. (2010). Bay esian compu- tation and mo del selection without likelihoo ds. Genetics , 184 243–252. Lombaer t, E. , Guillemaud, T. , Cornuet, J.-M. , Mala usa, T. , F acon, B. and Estoup, A. (2010). Bridgehead effect in the worldwide inv asion of the biocontrol Harlequin Ladybird. PloS ONE , 5 e9743. Lopes, J. and Beaumont, M. (2010). ABC: a useful Bay esian tool for the analysis of p opulation data. Infe ction, Genetics and Evolution , 10 825–832. Lopes, J. S. , Balding, D. and Beaumont, M. A. (2009). PopABC: a program to infer historical demographic parameters. Bioinfor- matics , 25 2747–2749. Marin, J. , Pudlo, P. , Rober t, C. and R yder, R. (2011). Ap- proximate Bayesian computational metho ds. ArXiv:1011:0955. Miller, N. , Estoup, A. , Toepfer, S. , Bourguet, D. , Lapchin, L. , Derridj, S. , Kim, K. S. , Reynaud, P. , Furlan, L. and Guillemaud, T. (2005). Multiple transatlantic introductions of the W estern corn ro otw orm. Scienc e , 310 992. Nunes, M. A. and Balding, D. J. (2010). On optimal selection of summary statistics for approximate bay esian computation. Sta- tistic al Applic ations in Genetics and Mole cular Biolo gy , 9 34. P ascual, M. , Chapuis, M. , Balany ` a, J. , Huey, R. , Gilchrist, G. , Serra, L. and Estoup, A. (2007). In troduction history of Drosophila subobscura in the New World: a microsatellite-based survey using ABC metho ds. Mole cular Ec olo gy , 16 3069–3083. P a tin, E. , La v al, G. , Barreiro, L. , Salas, A. , Semino, O. , Sant achiara-Benerecetti, S. , Kidd, K. , Kidd, J. , V an Der Veen, L. , Homber t, J. et al. (2009). Inferring the demo- graphic history of African farmers and pygmy h unter-gatherers using a multilocus resequencing data set. PLoS Genetics , 5 e1000448. Pritchard, J. , Seielst ad, M. , Perez-Lezaun, A. and Feldman, M. (1999). Population growth of h uman Y c hromosomes: a study of Y chromosome microsatellites. Mole cular Biolo gy and Evolu- tion , 16 1791–1798. Ramakrishnan, U. and Hadl y, E. (2009). Using ph ylochronology to reveal cryptic p opulation histories: review and syn thesis of 29 ancient DNA studies. Mole cular Ecolo gy , 18 1310–1330. Ra tmann, O. , Andrieu, C. , Wiujf, C. and Richardson, S. (2009). Mo del criticism based on lik eliho o d-free inference, with an application to protein network evolution. Pro c. Nat. A c ad. Sci. USA , 106 1–6. Rober t, C. and Casella, G. (2004). Monte Carlo Statistic al Meth- o ds . 2nd ed. Springer-V erlag, New Y ork. Sainudiin, R. , Thornton, K. , Harlo w, J. , Booth, J. , Still- man, M. , Yoshida, R. , Griffiths, R. , McVean, G. and Don- nell y, P. (2011). Exp eriments with the site frequency sp ec- trum. Bul letin of Mathematic al Biolo gy . (T o app ear.), URL http://www.isibang.ac.in/ ~ statmath/eprints/2010/8.pdf . Sousa, V. , Fritz, M. , Beaumont, M. and Chikhi, L. (2009). Approximate Bay esian computation without summary statistics: the case of admixture. Genetics , 181 1507–1519. Stephens, D. and Donnell y, P. (2000). Inference in population genetics (with discussion). J. R oyal Statist. So ciety Series B , 62 602–655. T a v ar ´ e, S. , Balding, D. , Griffith, R. and Donnell y, P. (1997). Inferring coalescence times from DNA sequence data. Genetics , 145 505–518. Templeton, A. (2008). Statistical hypothesis testing in intraspe- cific ph ylogeography: nested clade phylogeographical analysis vs. approximate Bay esian computation. Mole cular Ec olo gy , 18(2) 319–331. Templeton, A. (2010). Coherent and incoherent inference in ph y- logeography and h uman evolution. Pr o c. Nat. A cad. Sci. USA , 107(14) 6376–6381. Toni, T. and Stumpf, M. (2010). Simulation-based mo del selec- tion for dynamical systems in systems and p opulation biology . Bioinformatics , 26 104–110. Toni, T. , Welch, D. , Strelko w a, N. , Ipsen, A. and Stumpf, M. (2009). Approximate Bay esian computation scheme for parame- ter inference and model selection in dynamical systems. Journal of the R oyal So ciety Interface , 6 187–202. Verdu, P. , Austerlitz, F. , Estoup, A. , Vit alis, R. , Georges, M. , Th ´ er y, S. , Froment, A. , Le Bomin, S. , Gessain, A. , Homber t, J.-M. , V an der Veen, L. , Quint ana-Murci, L. , Bahuchet, S. and Heyer, E. (2009). Origins and genetic di- versit y of pygm y hun ter-gatherers from western central africa. Curr ent Biolo gy , 19 312–318. Wegmann, D. and Excoffier, L. (2010). Bayesian inference of the demographic history of c himpanzees. Mole cular Biolo gy and Evolution , 27 1425–1435. Wegmann, D. , Leuenberger, C. , Neuenschw ander, S. and Ex- coffier, L. (2011). ABCto olb ox: a versatile to olkit for approx- imate Bay esian computations. BMC Bioinformatics . T o appear. Wilkinson, R. D. (2008). Approximate Bay esian computation (ABC) gives exact results under the assumption of model error. App endix: SI Results A normal illustration. The following repro duces the Poisson geometric illustration in a normal mo del. If we lo ok at a fully normal N ( µ, σ 2 ) setting, we hav e f ( y | µ ) ∝ exp ( − nσ − 2 ( ¯ y − µ ) 2 / 2 − σ − 2 n X i =1 ( y i − ¯ y ) 2 / 2 ) σ − n hence f ( y | ¯ y ) ∝ exp ( − σ − 2 n X i =1 ( y i − ¯ y ) 2 / 2 ) σ − n I P y i = n ¯ y . If w e reparameterise the observ ations in to u = ( y 1 − ¯ y , . . . , y n − 1 − ¯ y, ¯ y ), w e do get f ( u | µ ) ∝ σ − n exp − nσ − 2 ( ¯ y − µ ) 2 / 2 × exp − σ − 2 n − 1 X i =1 u 2 i / 2 − σ − 2 " n − 1 X i =1 u i # 2 2 since the Jacobian is 1. Hence f ( u | ¯ y ) ∝ exp − σ − 2 n − 1 X i =1 u 2 i / 2 − σ − 2 " n − 1 X i =1 u i # 2 / 2 σ − n Fig. 8. Empirical distributions of the log discrepancy log g 1 ( y ) /g 2 ( y ) for datasets of size n = 15 simulated from N ( µ, σ 2 1 ) (left) and N ( µ, σ 2 2 ) (right) distributions when σ 1 = 0 . 1 and σ 2 = 10 , based on 10 4 replications and a flat prio r 8 www.pnas.org/cgi/doi/10.1073/pnas.xxx Footline Author Considering b oth mo dels y 1 , . . . , y n iid ∼ N ( µ, σ 2 1 ) and y 1 , . . . , y n iid ∼ N ( µ, σ 2 2 ) , the discrepancy ratio is then giv en b y σ n − 1 2 σ n − 1 1 exp σ − 2 2 − σ − 2 1 2 n − 1 X i =1 ( y i − ¯ y ) 2 + " n − 1 X i =1 ( y i − ¯ y ) # 2 and is connected with the lack of consistency of the Bay es factor: Lemma 2. Consider mo del sele ction b etwe en mo del 1: N ( µ, σ 2 1 ) and mo del 2: N ( µ, σ 2 2 ) , σ 1 and σ 2 b eing given, with prior dis- tributions π 1 ( µ ) = π 2 ( µ ) e qual to a N (0 , a 2 ) distribution and when the observe d data y c onsists of iid observations with fi- nite me an and varianc e. Then S ( y ) = P n i =1 y i is the minimal sufficient statistic for b oth mo dels and the Bayes factor b ase d on the sufficient statistic S ( y ) , B η 12 ( y ) , satisfies lim n →∞ B η 12 ( y ) = 1 a.s. Fig 8 illustrates the b ehaviour of the discrepancy ratio when σ 1 = 0 . 1 and σ 2 = 10, for datasets of size n = 15 sim u- lated according to b oth mo dels. The discrepancy (expressed on a log scale) is once again dramatic, in concordance with the abov e lemma. If we now turn to an alternative choice of sufficient statis- tic, using the pair ( ¯ y , S 2 ) with S 2 = n X i =1 ( y i − ¯ y ) 2 , w e follow the solution of Didelot et al. (2011). Using a conju- gate prior µ ∼ N (0 , a 2 ), the true Ba yes factor is equal to the Ba yes factor based on the corresp onding distributions of the pair ( ¯ y , S 2 ) in the respective mo dels. Therefore, with suffi- cien t computing p ow er, the ABC appro ximation to the Bay es factor can be brought arbitrarily close to the true Bay es fac- tor. How ever, this coincidence does not bring any intuition on the behaviour of the ABC approximations in realistic settings. La rger experiment. W e also considered a more informative p opulation genetic experiment with the same scenarios (1 and 2) as in the second exp erimen t. One hundred datasets were sim ulated under scenario 1 with 3 p opulations, i.e. 6 param- eters. W e take 100 diploid individuals p er population, 50 lo ci p er individual. This thus corresp onds to 300 genot yp es p er dataset. The IS algorithm w as performed using 100 coalescent trees p er particle. The marginal likelihoo d of b oth scenarios has b een computed for the same set for b oth 1000 particles (IS1) and 50,000 particles (IS2). A national cluster of 376 pro cessors (including 336 Quad Core pro cessors) was used for this massive exp eriment (which required more than 12 calen- dar da ys for the importance sampling part). The confidence about the IS appro ximation can b e as- sessed on Fig 7, which sho ws that b oth runs most alwa ys pro- vide the same numerical v alue, which almost uniformly is v ery close to one. This makes the fit of the ABC approximation to the true v alue harder to assess, even though we can sp ot a trend to wards under-estimation. F urthermore, they almost all lead to correctly select model 1. Footline Author PNAS Issue Date Volume Issue Number 9
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment