Predicting protein contact map using evolutionary and physical constraints by integer programming (extended version)
Motivation. Protein contact map describes the pairwise spatial and functional relationship of residues in a protein and contains key information for protein 3D structure prediction. Although studied extensively, it remains very challenging to predict…
Authors: Zhiyong Wang, Jinbo Xu
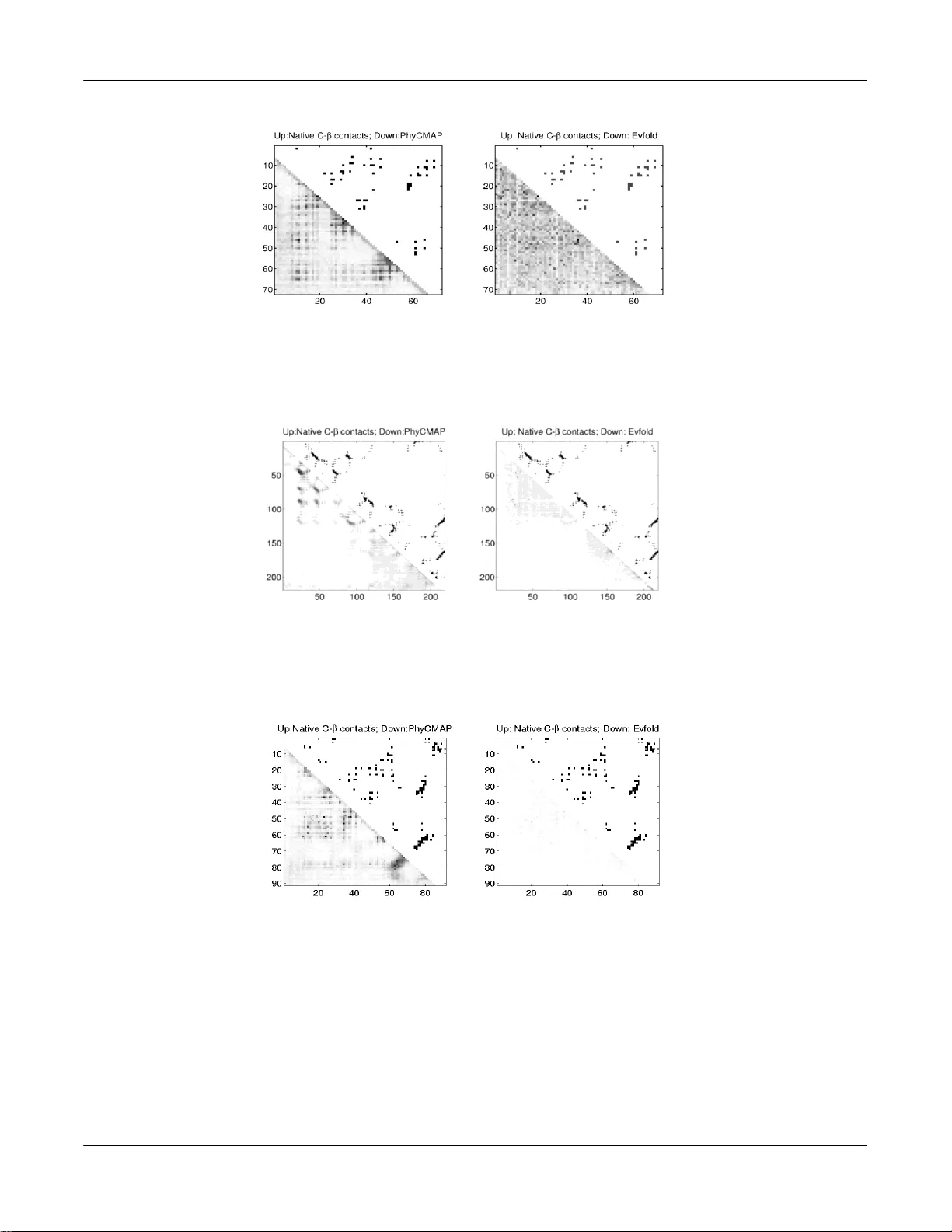
An earl y vers ion of this paper app ears at ISMB 2013 (see htt p://bioinform atic s.oxfo rdj ournals.org/c ontent/29/13/i26 6.full ) 1 Predict ing protei n contac t map using evol utionar y and ph y sical constra ints b y integer progra mming (extended versi on) Zhiyong Wang 1 and Jinbo Xu 1,* 1 Toy o ta Technolo gical Institute at Chic ago 6045 S Ke n w oo d, IL, 60637, U SA * Correspo n ding au thor (email : jinbo xu@gmail.co m) ABSTR ACT M otivation. Prot ein contact m ap d e sc rib es the pairwis e spatial and function al relationsh ip of r e s idu es in a prot ein an d c ontains key info r- mation f or pr o tei n 3D s tructur e predict ion. Alt hough stu died extens i vely, it remai ns very ch all enging to pr ed ict c on t a ct map us ing only s e- quence i nf o r mation. M ost existin g methods pr edict th e c onta c t map m atrix elemen t - by - element, ignori ng c orrelation among c ontacts an d phy s- ic a l f ea sib ility o f th e wh ole c ontact map. A c o uple o f r ec ent m ethods pr edict c ontact map ba s ed upon r esid ue co - evolution, t akin g i nto c on si d- eration c ontact c orr elation and enforcing a sp arsity restraint, but these methods r equire a very la rg e nu mber o f s equenc e homologs for t he pr o t ein un der c on si der ation and th e resultant c o nt a ct map m ay be s til l ph y sic ally unf avo r able. Results. This paper presents a n o vel meth od P h y C MAP f or c ontact m ap predict ion, integr ating b oth evol uti onar y and physic al r e str aint s b y machine l earning and i nteger lin ear program ming (I LP). Th e ev olution ary restr aints includ e s equence profile, r e si due co -evoluti on and c ontext- sp e cif ic s tatistical p otenti al. T h e phy s ic a l r estr aints sp ecify mor e c onc rete rel atio nship amon g c ontacts th an the sp a rsi t y r estr aint. As such, our method gr eatly r educ es the s olution sp a c e of the c ontact map m atrix and t hus, sig nific antly improv e s pred iction accuracy. Exp er im ental r esults confirm th at Ph yC MAP outperforms c urrentl y p opular m ethods n o matte r how m any s equence homologs are available f or the protein under co n sid eration. P hyCMAP c an predict c ontacts within min utes a ft er PSIBLAST s earch f o r s equenc e homol ogs is done , much f aster t han th e two recent methods P SICOV and EvF old. Availability : s ee http://r aptor x.uchic ago.edu f or the web s erver. 1 IN T RODUC TION I n this pape r we say two residue s of a prote in are in contac t i f thei r Eucl idean distanc e is less than 8Å . T he distanc e of two residue s ca n be calc ulated on C α o r C β at om s, c orrespo nding to C α o r C β co ntacts . A pro tein contac t map is a b inary L× L ma trix where L is the pro tein length . I n this m atrix, an eleme nt with v alue 1 i ndic ates the co rres ponding t wo residu es are i n contac t; othe r w ise, not i n co ntact. Pro tein co ntact map de scrib es t he pai rwise sp atia l and func tional relatio nship of residue s in a p rotei n. Predic ting c ontact map from se quenc e has been an activ e researc h t opic in rec ent y ears partial ly because co ntact map is helpf ul fo r pro tein 3D struc ture predic tion ( Ort iz, et al., 1999 ; Vassura , et al. , 2008 ; Vend rusco lo, et al ., 1997 ; Wu, et a l., 2011) and pro tein mo del qual it y assessm ent ( Z hou and Skolnic k, 2007) . Pr otein co ntact map has also been used fo r protein structu re alignm ent and cl assific ation (Cap rara, et al., 200 4 ; Wang , et al., 2013 ; Xu , et al. , 2006). Many mac hine learning metho ds have been develo ped fo r pr ot ein co ntact pred iction in the past d ecades (Farise lli a nd Cas adio , 1999 ; Gö bel, et al., 2004 ; O lmea and Valenci a, 1 997 ; Pun ta a nd Rost, 2005 ; Vendrusc o lo and Do many , 1998 ; Vu llo, et al., 2006) . For example , SVMSEQ (Wu and Z hang, 2 008) and SV Mc on ( Ch eng and Bald i, 2007) use suppo rt vec tor machin es; NNco n ( Tegg e, et al., 2009 ) uses a recursiv e ne ural netwo rk; Dis till (Baú , et al ., 2006) use s a 2-dime nsiona l recurs ive neural netwo rk; and C MAPpro (Di Lena, et al., 2012) uses a mu lti -lay er neural netwo rk. A lt houg h very different, these methods are com mon in that they predict the co ntact map mat rix ele ment - by- element, igno ring the c orrela tion amo ng contac ts a nd also physical fea sibility of the w hole co ntact map (phy sical co nstr ain ts ar e not totally independe nt of contac t correlation) . A specia l ty pe of phy sical constrain t is t hat a c ontac t map matrix must be s pars e, i.e., the nu m- ber o f co ntacts in a pro tein is only linear i n its length . Two recen t m ethods (PSIC OV (Jone s, et al., 2012) and Ev fo ld (M orco s, et al ., 2011 )) p redic t c ontac ts u sing o nl y residue co -evo lut io n i n- fo rmation deriv ed from seque nce homo log s and e nforc ing th e abov e-mentione d sparsi ty co nstraint. Ho wev er, both of them r equire a large number (at l eas t severa l hundred s) of sequence homo lo gs f or t he protein under predic tion. This ma kes the pred icted co ntacts not very us e- ful in real-wo rld prote in mode ling si nce a ( g lobular) pro tein with many s equence hom olo gs usually has v ery similar temp lates in PDB and thus, templ ate-based mode ls are of goo d quality and cannot be further impro ved by pr edic ted co ntacts. Conv ersely , a prote in wit hout close temp lates in PDB , wh ich may benefi t fro m co ntact predic tion, usu ally has very few sequenc e homo logs. Further , these two me thods e n- fo rce o nl y a v er y simpl e spa rsity co nstraint (i.e., the tota l num ber of co ntacts in a p rotein is sma ll), ig nori ng m any mo re co ncrete co n- straint s. To name a f ew, one residue can have only a small number of contacts, depending on i ts s eco ndary stru ctu re and neighbo ring res i- dues. The numb er o f co ntacts betw een two b eta st rands is bo unded by the strand length. A stro-Fo ld ( Klepei s and Flouda s, 2003) possibly i s t he firs t co ntact predict ion metho d that makes use of phy sical co nstraints , wh ich imply the s pars ity co nstraint u sed by PSIC OV and Ev fo ld. Howev er, s om e o f the phy s ical co nstraints are too re str ictive and po ssib ly unreali stic. Fo r examp le, it requires that a residu e i n one beta s tran d c an only be in c ontac t wi th o ne residu e in anoth er beta strand. More i mpo rtantly , A stro-Fo ld does not make use of evo lutionary info rmation an d thu s, significantly r educes its pred ict ion acc uracy . I n this paper , we pre sent a nov el m etho d Phy CMA P fo r c ontac t m a p pred ictio n by integra ting bo th evo lutionary and phy s ical c o nstraints using machine learning (i.e., Rando m Fo rests ) a nd integ er linea r pro g ramming (I L P). Phy CMA P first p redic ts the p robab ility o f any two Z. Wang et al. 2 residues fo rming a co ntact using both ev olutio nary and non- evo lutionary informatio n . Th en , Phy CMAP infers a given number o f t op co n- tacts based upo n pred icted c ontac t pro babil ities by enfo rcing a set of reali stic phy sical co nstr ain ts , which specify more co ncret e rel atio nship amo ng co ntacts and also imply the sparsity restraint used by PS ICOV and Ev fo ld. By c om bining both e vo lutio nary and phy sical c o n- straint s, our metho d grea tl y reduc es t he solut ion s pac e of a co ntact map and l eads to muc h better predic tion accura cy . Exper imen tal r esul ts co nfirm that Phy CMA P outperfo rms cur rently popular metho ds no m atter how many sequence homo logs are av ailable for the p rotein u nder predictio n . 2 METH ODS Overv iew . As shown in Fig ure 1, our me thod co nsists of s eve ral key co mponents . First, we emp loy Rando m Fo rests (RF) t o predic t the co ntact pro bability o f any t wo residu es f rom bo th ev olutiona ry and non-evolutiona ry inform ation . The n we emplo y an integ er l inea r pr o- gram ming ( IL P) metho d to s elect a set o f to p co ntacts by maximizing their ac cum ulative p robab ilitie s subj ect to a se t of phy sical c o n- straint s, wh ich a re repr esente d as linear co nstraints. The resulta nt top co ntacts fo rm a phy sically f avo rable c ontac t map for the protein un- der co nsidera ti on. Figure 1. The o verv iew of o ur co ntact predic tion appro ach. 2.1 Predicting contact probab ili ty by Rando m Forest s We use R andom Forests ( RF) to predic t the pro bability of an y tw o residues fo rming a c o ntact from s evera l ty pes of info rmat ion: 1) evo lutionary informatio n f rom a s ingle p rotei n f amily , e.g ., PSI - BLA ST sequenc e pro file (Altschul and Ko onin , 1998) and t w o ty pes o f mutu al info rmatio n; 2) non-ev o lut iona ry info rmation d e- rived fro m the s olv ed protein s tructu res, e. g. , residue c ontac t pote n- tial; 3) m ixed info rmatio n from the abov e two sourc es, e.g., EPA D (a co ntext-sp ecif ic distanc e-based s tatist ical pote ntial) ( Zhao and Xu , 2012) , s eco ndary stru ct ure predic ted by PSIPR ED (Jones , 1999 ) , an d homo logous pairwis e contac t score; and 4 ) a mi no acid phy s ic - chem ical propert ies . Some f eatu res are c alc ulated o n the residues in a l oc al windo w of size 5 c entered a t th e residu es un der c ons ide ration. I n total there are ~300 features fo r each r esidue pair. We trained our RF m odel using t he Rando mFo rest p ackag e in R ( Breima n, 2001 ; L iaw and Wi ener, 20 02) and selected the mo del par ameters by 5-fold cro ss valid ation. Be low we br iefly explain som e i mpo rtan t featur es. Sequen ce pro file. The posit ion-spec ific mut ation sco res a t residues i and j and their neighbo ring residues are u sed. Co ntrastive mu tual inf orm atio n (CMI). Let m i,j deno te the mu tual info rmation ( MI ) betw een two residu es i and j , w hich can be ca lc ula t- ed from the multipl e s equenc e alignm ent (MSA ) of all t he s equenc e ho mo log s. W e define the co ntras ti ve mu tual info rmation (CM I) as the MI difference betwee n one residue pair and all o f its neighboring pa irs, wh ich c an be calculate d as follow s. 2 1 , , 2 1 , , 2 , 1 , 2 , 1 , , ) ( ) ( ) ( ) ( j i j i j i j i j i j i j i j i j i m m m m m m m m C MI The C MI m easur es how the c o-mutatio n streng th of o ne residue pair di ff ers from its ne ighbo ring pairs. By us ing CM I i nste ad of MI, w e can remo ve the backg round bias of MI in a l oc al regio n, as sho wn in Figure 2 . When on ly a smal l numb er of sequenc e hom olog s are av ai l- able ( or at som e co nserved positions w ith en tropy <0.3) , we may have very low MI, wh ich may r esul t in artif icially hig h CMI . To a v oid this, we direc tl y set the CMI in these s ituations t o 0 . Mutual info rmat ion fo r t he chaining effect of residue co upling. When r esidue A has stro ng inte r actio n w ith B and B h as stro n g inte r actio n wit h r esidue C, it is likely that r esidue A also has inte r actio n w ith C. T o a cc ount fo r th is ki n d of chaini n g eff ect o f residue coupling, w e use th e pow er s of th e MI m atrix . In particula r , w e use MI k w her e k ranges from 2 to 11 t o capture thi s kind of chai n ing ef fec t. Wh e n there are ma ny s equence h omolo gs (i.e., MI is ac curate) , t his k ind o f MI powe r s great ly i m- prov es medium- and long-range co ntact p redictio n (see R esults for detai ls). Figure 2. Th e co ntr astive m utua l info rmatio n (lower triang le) and mutua l inf orm ation (upper t riang le) of protein 1j8bA .The nativ e co ntacts are represen ted by boxes. T he b ig y el low and red box es co ntain a g roup of med ium-rang e and long- range c ontac ts, respe c- tively . Integer Pro gramming Model Predict contact probability by RF Prior kno wledge on contact map Objectiv e function Linear phys ical constraints Physically feasible c ontact map Predicting protein contact map usin g evolut ionary and phy sical constra ints by i nteger progr a mming (e xtend ed ver sion) 3 Residue contac t po tential . We use res idue co ntact-ba sed pote ntial describ ed in (Tan , et al ., 2006) EPAD . I t is a co ntext-specif ic dis tance-dep enden t statistic al potent ial (Zhao and Xu, 2012), der ived from pro tein evo lutionary informat ion and solved p rotein str uctures i n PDB . The C α an d C β at om ic inter action po tential at a ll the dis tance bin s is us ed as fea tures . Th e atom ic distance is di scretiz ed into bins by 1 Å and all the distanc e>15Å is gro uped in to a sing le b in. Hom olo g ou s p airwi se contact sco re (HPS) . T his sc ore quan tify the pro bability of a residue pair f orming a contac t betwe en t wo se co n dary structur es. Let PS beta (a , b) denot e the probab ility of two amino acids a and b fo rming a bet a contac t . L et PS helix (a , b) deno te the probabil it y of two amino acids a and b form ing a he lix co ntact . Bo th PS beta (a, b) and PS helix (a, b) are der ived fro m a s imple st atist ics on PDB2 5. L et H denote t he set of sequence hom olog s of the pro tein unde r consid eratio n. Let i and j deno te two posit ions in th e multip le seque nc e align ment of the hom olo gs . W e calcu late t he ho mo logous pairw ise co ntact sc ore HP S for the two positions i and j as fo ll ow s. Where h i and h j de note the am ino acids of the homo log h at pos it ions i and j , respe ctiv ely. 2.2 The integ er linear p rogram ming (ILP ) metho d The var iables . Let i and j deno te res idue pos itio ns and L the protei n length . Let u a nd v ind ex seco ndary structu re seg ments of a pro tei n. L et Begin( u) and End(u) d enote the fir st and last residu es of the seg ment u and SS eg(u) the set { | } i B e g i n u i E n d u . Let SSt ype( u) denote the seco ndary structu re ty pe of o ne r esidue or o ne segmen t u . Let Len( u) denot e the leng th o f the seg ment u . W e u se the bin ary var i- ables i n Ta ble 1 . Table 1 . The b inary variables used in the I L P formulatio n. Variab le Explana tion X i,j equal to 1 if th ere is a co ntact betw een t wo resi dues i and j . AP u,v equal to 1 if two beta-s trands u and v form an anti-para llel b eta- sh e et. P u,v equal to 1 if two beta-s trands u and v form a par allel beta-shee t. S u,v equal to 1 if two beta-s trands u and v form a be ta-sheet . T u,v equal to 1 if th ere is an alph a -br idge betw een t wo helic es u and v . R r a non-neg ativ e integ ral relaxa tion variab le of t he r th so ft co nstraint. The ob jectiv e functio n. I ntuitiv ely we shall cho o se those co ntacts with the highest probab ility predicted by our Ran dom Forest mo del, i.e ., the obj ective functio n shal l be t he sum of predicted pro babilities o f the sel ect ed co ntacts. Ho wever , the selected co ntacts sha ll also m in i- miz e t he viola tion of the phy sical co nstraint s. To enforc e this, we use a vecto r of rel axa tion var iable s R t o measur e t he degree t o whic h all the sof t const raints ar e vio lated. Thus, o ur objectiv e function is as f o ll ow s. 6 , , , ) ( ma x i j j i j i R X R g P X Where , ij P is the co ntact pro bability predic ted by our Random Forest mo del fo r two residu es and r r R R g ) ( is a linear penalty f unction where r run o ver all t he sof t constraint s. Th e relax ation v ariable s wi ll be fur ther exp lained in the follo wing se ctio n. The con straints . We use both soft and hard co nstraints . T here is a s ingle r elaxa tion variab le for e ach gro up of soft constrain t, but the hard co nstraints are strictly en fo rced. We pena li ze the vio lation o f sof t constrain ts by inco rporating the relaxa tio n variables to the object ive f un c- tion. The co nstraints in groups 1, 2, and 6 are s of t co nstr ain ts. Thos e i n gro ups 3, 4, 5 and 7 are hard co nstraints , some of wh ich are similar to wha t are u sed by A str o-F old (Klepe is and F loudas, 2003). Group 1. This group of sof t co nstraints bound from abov e t he t ota l num ber of co ntacts tha t can be fo rmed by a single r esi due i (in a se c- ondary s tru ctu re ty pe s1 ) w ith all the other r esi dues in a se co ndary structure ty pe s2 . 2 ) ( : 2 , 1 1 , , 1 ) ( , s j SStype j s s j i b R X s i SS t y p e i Where 1 , 2 ss b is a co nstant emp iric ally determin ed from our tra i ning da ta (se e Tab le 2) and 1 R is th e rela xation v ariable . Table 2. Th e empirica l v alues of 1 , 2 ss b calcul ated from the training data. Th e first co lumn indic ates the co mbinat ion of two seco ndary structur e ty pes : H( α -helix), E( β -stran d) or C(co il). Each row contai ns two empi ric al v alues for a pair of se co ndary s tru ct ure ty pes. Co lum n “95%”: 95% of the sec o ndary s tructure pai rs have the number o f contact s sma ller than the v alue in this co lumn ; Co lumn “Max”: the lar gest number of c ontac ts . Z. Wang et al. 4 s1,s2 95% Max H,H 5 12 H,E 3 10 H,C 4 11 E,H 4 12 E,E 9 13 E,C 6 15 C,H 3 12 C,E 5 12 C,C 6 20 Group 2. Th is g roup o f c onstrai nts bound t he t ota l number of c ont acts betwe en two strands sh aring at least o ne co ntact. L et u and v d e- note tw o beta strands . We have ) ( ), ( , 2 , )) ( ), ( min ( 3 u SSeg j v SS eg i v u j i v L e n u L e n S R X 3 ) ( ), ( , )) ( ), ( ma x ( 3 . 3 R v Len u L en X u SSeg j v SSeg i j i The two c onstrain ts are explain ed i n Figure 3 as f ollo ws. Figure 3(A) shows that th e t ot al n umbe r o f co ntacts b etween two b eta st rands diverg es into two groups when m i n , 9 L e n u L e n v . One gro up is due to beta strand pairs fo rming a beta sheet . The ot her may be due to inco rrectly predicted beta str ands or beta strand pa irs not in a beta shee t . Figure 3(B) shows that the tot al numbe r of c ontac ts betwe en a pair of beta strands has an uppe r bound pro portional t o the length of the long er bet a strand. Since the re are points belo w the skew line in Fig ure 3(A ), which indic ate th at a bet a strand pair may hav e fewer than 3 m i n , L e n u L e n v co ntacts, we add a relaxa tion va ri abl e 2 R to the lower bound co nstrai nts in Group 2. Simi larly , we use a relax ation variable 3 R fo r the uppe r bo und co nstrain ts. (A ) (B) Figure 3 . The skew lines indic ate t he bounds fo r the to tal numbe r o f co ntacts between two b eta st rands. (A ) lower bo und; (B) uppe r bo und. Group 3. W hen two strands fo rm an anti -pa rallel beta-sh eet, the c ontacts of neighbo ring r esid ue p airs shall satisf y the fo llowing con- straint s. 1 1 , 1 1 , 1 , j i j i j i X X X Where ,1 ii are resid ues in one strand and ,1 jj are r esidues in the o ther st ran d. Group 4. When there are par allel co ntacts betwee n two strands, the co ntacts of neighbo ri ng residu e pair s should satisfy t he fo llowing co n- straint s. 1 1 , 1 1 , 1 , j i j i j i X X X Where ,1 ii are resid ues in one strand and ,1 jj are r esidues in the o ther st ran d. Predicting protein contact map usin g evolut ionary and phy sical constra ints by i nteger progr a mming (e xtend ed version) 5 Group 5. One be ta-stra nd u can form beta-sheet s wi th up to 2 other beta-strand s. 2 ) ( : , be ta v SSt ype v v u S Group 6. There is no c ontac t betwee n the s tart and end residues of a loo p segmen t u . ) ( ), ( , 0 4 , u E nd j u B egin i R X j i I n our training set, there are tot ally ~8000 loo p segment s, and only 3.4% of t hem hav e a co ntact betw een the start and end r esid ues. To allo w the ra re ca ses, we use a relax ation v ariable 4 R in the constrain ts. Group 7. One re sidue i canno t hav e co ntacts with bo th j and j+2 w h en j and j+2 are in the same alph a helix . 1 2 , , j i j i X X Group 8. This g roup o f co nstraints model s the rel ationsh ip amo ng di ff erent gro ups of variable s. v u v u v u S P AP , , , ) ( ), ( , , , v SSeg j u SSeg i S X v u j i 6 , 1 , i j L j i j i k X Where k is the num ber o f top co ntacts we want to pre dict. Integer pro gram solv er: IB M CPLE X acad emic v ersion V 12.1 (C PL EX, 2009). Training data . It c onsists of 900 non- redund ant protein str ucture s, any two o f w hic h share no mo re t han 25% sequence iden tity . Sinc e there a re far fewer co ntacts than no n-c ontacts, we use all the c o ntacts and ra ndo mly sampl e only 20% o f the non- contacts a s the training data. A ll the training prote ins are s elec ted befo re CA SP10 (th e 10 th Cri ti ca l Asse ss me nt o f Structur e Predic tio n) started in May 2012 . Test data I: CAS P10 . Th is set co ntain s 123 CA SP10 targe ts with publ icly available nativ e struc tures. Meanwh ile, 36 of th em are c lassi fied as ha rd targets becaus e the t op half of the ir subm itted mo dels have averag e TMsco re<0.5 . When they were j ust relea sed, mo st o f t he CA SP10 targ ets share low s equenc e identity (<25%) wit h the prote ins in PDB . BL A ST indicate s that there are only 5 short CA SP10 tar gets (~50 re sidues) which have se quence iden tity sli ght ly abov e 30% wit h our tra ining pr ote ins. Test data II: Set600. This set contains 601 p roteins r andom ly ext racted fro m PDB 25 (B renner , et al., 2000) a nd wa s co nstructed bef ore CA SP10 s tar ted . They have the follo wing properties: 1) less than 25% seque nce identi ty with the training prote ins; 2) each pr ote in h as at least 50 r esidu es a nd an X-ray struct ure w ith res ol utio n better than 1.9Å ; 3) eac h pro tein has at least 5 resid ues wi th pred icted s eco ndary structur e being alpha- helix o r bet a-stran d. Bo th t h e t raining set a nd Set6 00 a re s amp led from PDB25 (Wang and D unb rack, 2003) , i n w hic h any two proteins sha re less than 25% sequenc e ident ity . Se que nce iden tity is calcula ted usi ng the me thod in (Brenn er, et al., 20 00) . Program s t o be com pared . W e c om pare our met hod, denoted as Phy CMAP, with four state - of -the-art me thods: CMA Ppro (Di L en a, et al., 2 012 ), NNc o n (Teg ge, e t a l., 2009), PS I COV (Jones , et al., 2012) and Ev fo ld (Mo rco s, et al., 2 011). We run NNCo n, PSI COV a nd Ev fold loc all y a nd CMA Ppr o by its web server . We do not co mpare our me thod with Astro-F old bec ause it is not publicly a vail able. Since A stro-Fo ld does not make u se of any evolutiona ry i nfo rmation, it do es not perfo rm wel l fo r many pr otein s . Ev aluation methods . Dependi ng on the chain dis tance o f the two residues, t here are three ki nds of c ontacts: s hort-rang e, medium- rang e and long- range. Sho rt-rang e co ntacts are close ly related to loc al co nfo rmation and are r ela tively easy to pre dict . Med ium -range and lo ng- range contac ts dete rmine the ov eral l s hape of a pr o tein and are more challeng ing to predict . We evalu ate pred i ction acc uracy usi ng t he top 5, L /10, L/5 pr edicted co ntacts wher e L is the prote in leng th. M eff : the numbe r of non-redundan t s equence ho mo logs. Given a target pro tein and the mu ltip le sequenc e alig nment of al l of its ho m o- log s, we mea sure th e numbe r of no n-redundan t sequenc e ho mo logs by M eff as fo llows. e f f , 1 M i ij j S (1 ) Where bo th i and j go ov er all the s equenc e homo logs and S i,j is a bina ry simil ar ity value between two pro teins . Follo wing Ev fo ld (M orc o s, et al., 2011) , we co mpute the similar ity of two s equenc e ho mo logs using t hei r hamm ing distanc e. Tha t is , S i,j is 1 if the no rmalized h a m- ming distance is < 0. 3; 0 othe rwise. Z. Wang et al. 6 3 RESUL TS Perform ance on the CAS P10 set. As sho wn in Ta ble 3, tes ted on the who le CAS P10 set, ou r Phy CMA P si gnif icantly exceeds the 2 nd best metho d C MAPpro in terms o f the a cc uracy of the top 5 , L /10 and L /5 pr edic ted c ontac ts. T he adv an tage of Phy CM AP ov er CMAP pro beco mes smal ler but still subs tantial when shor t -range c ontac ts are exc luded from co nsideratio n. Phy CMA P signific antly outperform s NNco n, PSIC OV and Evf o ld no matter ho w the perf ormanc e is evalua ted. Figur e 4 s hows the head- to -head co mpar ison betwe en Ph y- CMA P and C MAP pro and PSI COV fo r long -rang e contac t predicti on. Ta ble 3 . This table lists th e predic tion ac curac y of Ph y CMA P, PSICO V, NNCON , CMA Ppro and Ev fo ld fo r s ho rt - , medium- and long - range co ntacts , tested on 123 CA SP10 target s. Short-ran ge, seq uenc e di s- tance fr om 6 to 12 Medium and l ong- ra nge, s e- quence distanc e > 12 Medium range, sequ ence d i s- tance > 12 and ≤24 Long rang e, se quence distance > 24 Top 5 L/10 L/5 Top 5 L/10 L/5 Top 5 L/10 L/5 Top 5 L/10 L/5 PhyCMA P(C α ) 0.623 0.555 0.459 0.650 0.584 0.523 0.585 0.518 0.448 0.421 0.363 0.320 PhyCMA P(C β ) 0.667 0.580 0.482 0.655 0.604 0.539 0.621 0.550 0.465 0.514 0.425 0.373 PSICO V(C α ) 0.294 0.225 0.179 0.397 0.345 0.306 0.384 0.303 0.255 0.350 0.277 0.226 PSICO V(C β ) 0.379 0.281 0.223 0.522 0.458 0.405 0.515 0.387 0.316 0.457 0.372 0.315 NNCO N(C α ) 0.595 0.499 0.399 0.472 0.409 0.358 0.463 0.393 0.334 0.286 0.239 0.188 CMAPpro (C α ) 0.506 0.437 0.368 0.517 0.466 0.424 0.485 0.414 0.363 0.380 0.336 0.297 CMAPpro (C β ) 0.543 0.477 0.395 0.519 0.466 0.415 0.491 0.419 0.370 0.395 0.347 0.313 Evfold (C α ) 0.236 0.193 0.165 0.380 0.326 0.295 0.351 0.294 0.249 0.328 0.257 0.225 Figure 4. H ead- to -head c om paris on o f Phy CMAP, CMA Ppro and PSI CO V on t he 123 CA SP10 targets . Each po int represe nts the ac c uracy of two metho ds under com parison on a single test p rotein. Top L /5 pred icted long -rang e C β contac ts ar e co nsider ed. The averag e a c- curac ies of Phy CMA P, CMA Ppro, and PSI COV are 0.373, 0 .313, and 0.3 15, respec tively . Perform ance on the 36 hard CΑS P10 target s. A s shown i n Table 4 , tested on the 36 hard CA SP10 t arg ets our Phy CMAP exceed s t he 2 nd best metho d CMA Ppro by 5-7% in terms o f the ac curac y of the top 5, L /10 and L/5 pred icted contac ts. The advant age of Phy CMAP ov er CMAP pro bec om es sm a lle r but s till substan tial wh en sho rt -ra nge co ntacts a re exc luded from co nsidera tio n. Phy CMA P si gnif ica ntl y outperfo rms NNco n, PSI COV and Evfo ld no mat ter how many pr e dicted co ntacts are evalua ted. PSIC OV and Ev fo ld almost fai l o n these hard CAS P10 targe ts. By co ntrast, CMAPpro , NNc o n and Phy CMA P still can co rrectly pr edic t som e contacts . Figure 5 sho ws the he ad- to - head c om paris on of Phy CMAP , CMA Ppro a nd PSI CO V for mediu m- and lo ng- range co ntact pred iction. Table 4 . A cc uracy of PhyCMA P, PSI COV, NNc on, Ev fo ld a nd CMA Ppr o on th e 36 ha rd CA SP10 targets . The C β resu lts are in gray r ow s. Short-ran ge, seq uenc e distanc e from 6 to 12 Medium and l ong-rang e, sequ ence d istance > 12 Top 5 L/10 L/5 Top 5 L/10 L/5 PhyCMA P (C α ) 0.456 0.439 0.378 0.394 0.378 0.325 PhyCMA P (C β ) 0.478 0.469 0.414 0.444 0.409 0.363 PSICO V (C α ) 0.100 0.083 0.082 0.183 0.156 0.150 PSICO V (C β ) 0.144 0.113 0.103 0.239 0.196 0.180 CMAPpro PSICOV PhyCMAP Predicting protein contact map usin g evolut ionary and phy sical constra ints by i nteger progr a mming (e xtend ed version) 7 NNCO N (C α ) 0.400 0.372 0.320 0.367 0.317 0.289 CMAPpro (C α ) 0.383 0.347 0.314 0.328 0.322 0.292 CMAPpro (C β ) 0.433 0.398 0.344 0.394 0.362 0.308 Evfold ( C α ) 0.100 0.095 0.094 0.194 0.179 0.156 No te that both PSI COV and Ev fo ld, two r ecen t metho ds receiving a l ot of attentio ns from the co mmunity , do not perfo rm very well o n the CA SP10 set. Th is is partia lly be cause t hey require a large numbe r of sequence homo logs fo r the pro tein under predic tion. Neve rthel ess, mo st o f the CASP targets, especi ally t he har d o ne s, do not s atisfy this requir emen t bec ause a prote in w ith so m any homo log s v er y likely has high ly similar templ ates in PDB and th us, w as not used as a CASP targe t. Pred iction accuracy vs. t he number o f sequenc e hom olo gs. We div ide the 123 CA SP10 targets in to 5 g roups acco rding t o thei r log M eff values: [0,2 ), [2 ,4), [4,6 ), [6,8 ), [8 ,10] , w hich co ntai n 1 9, 17, 25, 36 and 2 6 t arget s, r espec tiv ely. Meanwh ile M eff i s the numbe r o f non- redundan t se quenc e hom olo gs for the pro tein unde r co nsideration (see Metho ds fo r def in i ti on) . Only medium- and l ong -rang e co ntacts ar e co nsidered here . Figure 6 clea rly show s t hat the pred iction ac curac y increases wi th r espec t to M eff . The m ore non- redundan t homo logs are availabl e, t he better predic tion acc uracy c an be ac hiev ed by Phy C MAP, Evfo ld a nd PSI COV. Ho wev er, C MA Ppro and NNCO N have d e- crease d acc uracy when log M eff >8. Figur e 6 a lso s ho ws t hat Phy CMAP outpe rfo rms Evf old, CMA Ppro an d N NCON acro ss all M eff . Ph y- CMA P g reatly outperfo rms PSI COV in predic ting C α contac ts reg ardless of M eff an d al so in p redicting C β co ntacts when log M eff ≤ 6 . Ph y- CMA P is c om parable to PSI COV i n p redicting C β co ntacts wh en lo g M eff >6 . Figure 6 . T he r elatio nship b etween p redict ion a ccu- racy and the num ber of non-redunda nt sequence hom olog s ( M eff ). X- axis is l og M eff an d Y-ax is i s t he mean acc uracy of top L /10 pr edicted co ntacts in the co rresponding CASP 10 target gro up. O nly med i- um- and long- range c ontacts are c onsidered . Perform ance on Set600 . To fai rl y co mpare our me thod with Ev fo ld (Morc os, et al., 2011) and PSI CO V ( Jones , et al. , 2012) , both of which require a la rge numbe r o f se quenc e hom olog s, we divide S et600 into two subse ts ba sed upo n M eff , the numb er of no n-redun dant sequenc e homo logs . The first subset co ntains 471 protein s with M eff >100 (see M etho ds for de fi nition) . Each pro tein in this subs et ha s >500 sequenc e ho mo logs, wh ich satisfies the requ iremen t o f PSI COV. The 2 nd subset is mo re cha lleng ing, co ntai ning 130 p roteins w ith M eff ≤100 . As sho wn in T able 5, even o n the 1 st subse t, P hy CMA P still excee ds PSI COV and Ev fo ld, althoug h t he advantag e o ver PSI COV is not su bstantia l fo r C β co ntact predict ion wh en shor t -rang e co ntacts a re exc luded f rom co nsideration. Phy CMA P al so outpe rfo rms N Nco n and CMA Ppr o on t his set . A s shown in Table 6, on the 2 nd subs et Phy CMAP si gnific antly o utperforms PSI COV and is slig htly better than CMA Ppro and N Nco n . Fig ures 7 and 8 show t he h ead- to -head c o mpariso n of Phy CMA P, PSIC OV and CMA Ppro. As s how n in the fi g- ures, when M eff >10 0, Phy CM AP outperf orms CMA Ppro on most test pro teins ; when M eff ≤100 , Phy CMA P outperfo rms bo th CMA Ppro and Figure 5 . Head- to -head co mpariso n of Phy CMAP, C MAP pro and PSI CO V on the 36 hard CA SP10 target s. Each poin t repre sents the acc uracy of two metho ds un der co mpari son on a sing le tes t protein . To p L/5 pred icted med ium- and long -rang e C β co ntacts are c onsid- ered. The ave rage acc uraci es of Phy CMAP , CMA Ppro, and PSI CO V are 0.3 63, 0.30 8, and 0.180 , respect ively . PhyC MAP CMAPpro PSICO V Z. Wang et al. 8 PSI CO V on most te st prote ins. These results again conf irm that our me thod applies t o a pr ote in w it hou t many sequenc e ho mo logs, on which PSI COV and Ev fo ld usual ly fail. Figure 7 . He ad- to -head co mparison o f Phy CMA P, CMA Ppro and PSI COV on the Se t600 pro teins w ith M eff >100 . Eac h point represen ts the a cc uracy of two metho ds under co mparison on a singl e test p ro tein. To p L /5 pred icted medium- and long- range C β co n- tacts are co nsidered. The avera g e acc uracies o f Ph y- CMA P, C MAP pro, and PSI CO V ar e 0.611 , 0.515 , and 0.569 , respec tiv ely Figure 8 . Head- to -head co m parison of Ph y CMA P, CMA Ppro and PSI COV on the Se t600 proteins with M eff ≤100 . Each point represent s the accur acy of t wo metho ds under com parison o n a si ngl e test p ro tein. To p L /5 pred icted medium- and long- range C β co n- tacts are co nsidered. The average a cc uracies of Ph y- CMA P, CMA Ppro , and PSI COV are 0.3 45, 0.28 7, and 0.059, r espec tively . Table 5 . A cc uracy on the 471 p roteins with M eff > 100 . Short-ran ge, seq uenc e distance from 6 to 12 Medium and l ong-range, sequence dista nce > 12 Top 5 L/10 L/5 Top 5 L/10 L/5 PhyCMA P (C α ) 0.761 0.653 0.539 0.716 0.675 0.611 PhyCMA P (C β ) 0.746 0.637 0.531 0.731 0.656 0.608 PSICO V (C α ) 0.457 0.341 0.257 0.528 0.465 0.411 PSICO V (C β ) 0.584 0.425 0.316 0.732 0.646 0.565 NNCO N (C α ) 0.512 0.432 0.355 0.432 0.361 0.308 CMAPpro (C α ) 0.682 0.551 0.431 0.710 0.642 0.574 CMAPpro (C β ) 0.671 0.542 0.436 0.674 0.601 0.532 Evfold ( C α ) 0.379 0.297 0.234 0.473 0.438 0.400 Table 6 . A cc uracy on the 130 p roteins with M eff ≤ 1 00. Ev fold is not lis ted sinc e it do es no t pro duce me aning ful p redi ctio ns fo r some p roteins . Short-ran ge, seq uenc e distance from 6 to 12 Medium and l ong-range, sequence dista nce > 12 Top 5 L/10 L/5 Top 5 L/10 L/5 PhyCMA P (C α ) 0.534 0.451 0.377 0.432 0.372 0.319 PhyCMA P (C β ) 0.505 0.435 0.365 0.418 0.364 0.314 PSICO V (C α ) 0.060 0.061 0.064 0.049 0.039 0.035 PSICO V (C β ) 0.077 0.070 0.073 0.069 0.050 0.045 NNCO N (C α ) 0.442 0.363 0.309 0.368 0.339 0.301 CMAPpro (C α ) 0.435 0.365 0.314 0.368 0.331 0.300 CMAPpro PSICOV Ph yCMAP Ph yCMAP CMAPpro PSICOV Ph yCMAP Ph yCMAP Predicting protein contact map usin g evolut ionary and phy sical constra ints by i nteger progr a mming (e xtend ed version) 9 CMAPpro (C β ) 0.532 0.434 0.353 0.358 0.331 0.280 No te t hat CMAPpro used Astra l 1.73 (Bren ner, et al., 2000 ; Di Le na, et al., 2012) as its training set, w hich shares >90% sequenc e identity with 22 6 prote ins in Set600 (180 wi th M eff >100 and 46 w ith M eff ≤100 ). To mo re f airly co mpare the pr edictio n metho ds, w e exclud e the 22 6 prote ins fro m Set60 0 that shar e >90% sequenc e ident ity wit h th e CMA Ppro train ing set . He re the seque nc e i den tity is calculated using CD- HI T (Li and Godz ik, 2006 ; Li, et al., 2001). Thi s results in a set of 291 pro teins with M eff > 100 and 84 proteins M eff ≤ 100 . Table 7 sho ws that Phy CMA P greatly outperfo rms C MAPpro and Ev fo ld on the reduced dataset . Phy CMA P also outp erfo rms PSI CO V in predict ing C α co ntacts, but is slightly wo rse in predict ing lo ng-rang e C β co ntacts. Table 7 . T his tabl e lists the pre dictio n acc uracy o f Phy CMA P, PSICOV , NNCO N, C MA Ppro and Evf old for short- , medium- and long - range co ntacts , tested on a reduc ed Set6 00. Accuracy on th e 291 Set60 0 pro teins with M ef f >100 and ≤90% sequ ence id entify w ith Astral 1.73 Short-ran ge, seq uenc e di s- tance fr om 6 to 12 Medium and l ong-rang e, s e- quence distanc e > 12 Medium range, sequ ence d i s- tance > 12 and ≤24 Long rang e, se quence distance > 24 Top 5 L/10 L/5 Top 5 L/10 L/5 Top 5 L/10 L/5 Top 5 L/10 L/5 PhyCMA P(C α ) 0.759 0.653 0.536 0.713 0.680 0.622 0.639 0.570 0.496 0.582 0.528 0.461 PhyCMA P(C β ) 0.741 0.641 0.534 0.746 0.653 0.611 0.655 0.571 0.500 0.636 0.550 0.477 PSICO V(C α ) 0.459 0.343 0.258 0.528 0.469 0.417 0.462 0.363 0.282 0.483 0.418 0.358 PSICO V(C β ) 0.582 0.422 0.314 0.733 0.650 0.569 0.647 0.496 0.371 0.674 0.584 0.495 NNCO N(C α ) 0.475 0.390 0.318 0.377 0.313 0.267 0.342 0.284 0.236 0.224 0.182 0.152 CMAPpro (C α ) 0.643 0.519 0.412 0.689 0.618 0.554 0.580 0.511 0.439 0.527 0.469 0.416 CMAPpro (C β ) 0.642 0.520 0.422 0.653 0.580 0.515 0.573 0.494 0.421 0.504 0.444 0.396 Evfold (C α ) 0.382 0.297 0.235 0.488 0.442 0.398 0.451 0.366 0.289 0.442 0.389 0.342 Accuracy on th e 84 Set600 protei ns with M eff ≤1 00 and ≤90% seq uence iden tity w ith A stral 1.73 PhyCMA P(C α ) 0.580 0.488 0.404 0.481 0.430 0.357 0.476 0.417 0.335 0.204 0.179 0.159 PhyCMA P(C β ) 0.548 0.468 0.392 0.454 0.408 0.345 0.452 0.399 0.331 0.220 0.214 0.187 PSICO V(C α ) 0.070 0.071 0.072 0.065 0.050 0.044 0.074 0.055 0.049 0.063 0.043 0.035 PSICO V(C β ) 0.081 0.078 0.083 0.088 0.068 0.059 0.092 0.066 0.059 0.076 0.058 0.046 NNCO N(C α ) 0.535 0.421 0.342 0.324 0.298 0.248 0.348 0.321 0.271 0.162 0.132 0.114 CMAPpro (C α ) 0.465 0.370 0.316 0.346 0.328 0.285 0.360 0.332 0.286 0.173 0.169 0.158 CMAPpro (C β ) 0.447 0.367 0.321 0.346 0.320 0.287 0.366 0.331 0.290 0.191 0.189 0.176 Evfold (C α ) 0.074 0.068 0.066 0.079 0.058 0.039 0.074 0.053 0.045 0.063 0.042 0.032 3.1 Co ntribution o f co ntrastive mu tua l info rmation and pairwi se co ntact sco res The co ntrastive mutual i nfo rmatio n ( C MI ) and ho mo logo us pai rw ise contac t sc ores (HPS) help impro v e the perf ormanc e of our ran dom fo rest mo del. Tab le 8 show s their c ontribu tion to the pr edictio n accuracy on the 471 pr ot eins ( w ith M eff >100 ) in Se t600 . Table 8 . T he co ntributio n of c ontrastive mutual info rmatio n and ho mo logous pa ir co ntact sco res to C β co ntact predic tion . Simi lar resu lts are obse rved fo r C α co ntact predic ti on. Short-ran ge con tacts Medium and l ong-rang e Top 5 L/10 L/5 Top 5 L/10 L/5 with CMI and HPS 0.754 0.632 0.521 0.720 0.649 0.589 no CM I and H PS 0.600 0.570 0.487 0.538 0.560 0.506 3.2 Co ntribution o f the pow ers of the MI matri x The pow ers of the M I matrix great ly impro ve l ong -rang e co ntact predic tion when many sequenc e homo log s are a va ilabl e, as show n i n T a- ble 9 . The av erage accuracy is co mputed on the Se t600 protein s w it h M eff >100 exclu ding those with > 90 % sequ ence ident ify with A s- tral1.73 . Figur e 9 shows gain from using the powers of the MI matrix for ea ch test prote in . Thi s r esul t indic ates that it is impo rtant to c o n- Z. Wang et al. 10 sider the chai ning effec t of residue c ouplings for medium- and long- range contacts and the powers of the MI matrix are goo d measu re of the chain ing effec t . It is also worth to mention that we can calc ulate t he power s of the MI matrix quickly , muc h faster than calc ulatin g the matrix inverse (as done by PSI COV ). Table 9 . T he co ntributio n of the po wers of the MI matrix to C β co ntact predic tion, tes ted on the Set 600 prot eins with M eff > 100 . Sho rt -rang e Medium and L ong range Medium- rang e Lo ng-range L /10 L /5 L /10 L /5 L /10 L /5 L /10 L /5 w/ o the MI pow er s 0.548 0.484 0.589 0.533 0.510 0.443 0.459 0.395 with the MI po wers 0.556 0.484 0.651 0.592 0.564 0.491 0.550 0.477 Figure 9 . Thi s fig ure sho ws the g ain fro m the pow ers o f the MI matrix. The first and seco nd panels show the medi um-rang e and long- range co ntact p redic tion, respec tively . Each point represen ts the acc uracy of the top L /5 predic ted co ntacts by two Random Fo rest mo dels, o ne with t he power s of the MI matrix and the oth er without the MI po wers. 3.3 Co ntribution o f phy sical cons traints Table 10 sho ws the impro vem ent resu lting from t he phy sical co ns tr ain ts (i.e., the I LP method) o ver the Rando m Forests (RF ) metho d on Set600 . On t he 471 proteins with M eff >100 , I LP improv es medium - a nd long -range contact p redic ti on, bu t not sho rt - rang e co ntact p redi c- tion. This result co nfirms that the phy sical co nstrai nts u sed by our ILP method indeed cap ture some glo bal prope rtie s of a protein co nt act map. The imp rov ement resu lting fro m the phy si cal c onstrai nts is la rger on t he 130 p rote ins w ith M eff ≤1 00 . I n particu lar, t he i mp rov e ment on short-ra nge co ntacts is subs tantial . Th ese results may imply t hat when homo logo us inform ati on is suffic ient , w e can pred ict short-r ange co ntacts v ery accurat ely and thus, cannot furth er imp rov e the pred ictio n by the phy sical co nstraints. W hen homo logo us info rmatio n is in- suff icient fo r a cc urate co ntact pred iction, we can impro ve the pred ic tion using t he phy sical co nstra ints, w hich are co mplement ary to ev olu- tiona ry information. Table 10 . The contr ibution of phy sical co nstrain ts for C β co ntact predic tio n. Short-ran ge con tacts Medium and l ong-rang e Top 5 L/10 L/5 Top 5 L/10 L/5 471 Set600 protei ns with M eff 100 RF+ILP 0.746 0.637 0.531 0.731 0.656 0.608 RF 0.754 0.632 0.521 0.720 0.649 0.589 130 Set600 proteins with M eff ≤ 100 RF+ILP 0.505 0.435 0.365 0.418 0.364 0.314 RF 0.445 0.368 0.299 0.414 0.342 0.281 3.4 Specif ic Examp les We show the co ntact predic tio n re sults of four CA SP10 targets T0 677D2, T0693D 2 , T0 756D1 and T0701D1 in Figure 10 , Fig ure 10 1 , Figure 12 , and Fig ure 13 , r espec tive ly . T067 7D2 does not hav e many se quence hom olo gs ( M eff =3 1.53) . A s s how n in Figure 10, our pr edi c- tion matc hes well w ith the nat ive co ntacts. Phy CMAP has top L/5 predictio n accurac y 0.357 on m edium- and long -rang e contac ts w hile Ev fold cannot co rre ctly predict any co ntacts. T0693D 2 has many sequ ence ho mo logs with M ef f =2208 .39. A s show n in Fig ure 11 , Ph y- CMA P does we ll in pred icting the long-distanc e co ntacts a round th e residue pai r (20,100 ). For th is target , Phy CM AP and Ev fo ld obta in top L/ 5 pred iction acc uracy o f 0.628 and 0 .419, respec tively , when both medium- and long -rang e co ntacts are co nsidered . I n Figure 12 , T0756D1 has m any sequenc e homo logs with M eff = 18 24.46 . The accuracy of the top L /5 pr edic ted co ntacts by our Phy CMA P i s 0. 944 , while the accurac y by Evfold is 0.500 . I n Figure 13, T0701D1 has a large numbe r of sequenc e hom olog s with M eff = 3300 .02, the acc uracy of the top L /5 predicted co ntacts by our Ph y CMAP is 0.794 , while the acc uracy by Evfo ld i s 0.44 4. Predicting protein contact map usin g evolut ionary and phy sical constra ints by i nteger progr a mming (e xtend ed version) 11 Figure 10 . The p redic ted me dium and long- range co ntacts fo r T0677D2. T he upper tr iangles display the na tive C β co ntacts. The lower triang les of t he l eft and rig ht plo ts displ ay the contac t pro babilitie s pred icted by PhyC MAP an d Ev fold, respect ively . Figure 11. The predic ted med ium and lo ng-rang e contac ts fo r T069 3D2. The upper triang les display the n ative C β co ntacts. The lower triang les o f the left and r ight plots display the co ntact pr o b abili ties pred icted by Phy CMAP and Ev fold, respec tively . Figure 12. The predic ted med ium and lo ng-rang e contac ts fo r T0756D 1. The upper triang les disp lay t he n ative C β co ntacts. The lower triang les o f the left and r ight plots display the co ntact pr o babili ties pred icted by Ph y C MA P and Ev fold, respec tively . Z. Wang et al. 12 Figure 13. The predic ted med ium and long -rang e contac ts fo r T070 1D1. The upper triang les disp lay t he nativ e C β co ntacts. The lower triang les o f the left and r ight plots display the co ntact pr o babili ties pred icted by Ph y C MA P and Ev fold, respec tively . 4 CONCLUSI ON AN D DISCUSS IONS This pape r has presen ted a novel metho d fo r pro tein c ontac t m ap predic tion by integra ting evo luti ona ry info rmation, non-ev olutionar y in- fo rmation and phy si cal co nstraints using machin e learning and integ er linear prog ramm ing. Our me thod dif fers fro m current ly popul ar co ntact predictio n m ethods in tha t w e enforc e a few phy si cal co nstraints , which imply the sparsity constrain t (used by PSI COV an d Ev fold), to the who le co ntact map and take into co nsider ation co ntact c orrela tion. Our metho d also differs fro m the first -princ iple metho d (e.g., A str o- Fold) in that w e exploit evo luti ona ry info rmation fro m sev eral aspects (e.g., t wo ty pes of mutual informat ion, co ntext -spe c ific distance pot ential and sequenc e prof ile) t o signif i cant ly improv e predic tio n accurac y . Exper imental re sult s co nfirm that our metho d outpe r- fo rms ex isting popula r mac hine learning m ethods (e.g., CMA Ppro and NN CON) and two r ecent residue co -evo lution me thods PSIC OV and Ev fo ld r egard less of the numb er o f sequence hom olog s availabl e fo r the protein unde r co nsider ati on . The study of our me thod i n dicates t hat the p hy sical co nstraint s are hel pfu l fo r co ntact predic tion , espec ially when the pro tein unde r consid- eration does not h av e many sequence hom o logs . Nev erthe less, t he phy sical c o nstr ain ts us ed by our c urrent method do not cov er a ll t he proper ties o f a pro tein contac t m ap. To further imp rov e pred iction acc uracy on medium and long-range co ntact pr edic tion, we may take into co nsidera ti on glo bal pr ope rties of a pr otein dis tanc e matrix. Fo r exa mp le, the pai rwise distances o f any three residues shall satisf y the triangle inequality . Som e resid ues a lso h ave co rrelated pairw ise d istanc es. To enforc e this kind of distanc e co nstraints , we shall introduc e distance v ariable s and also d efine their relatio nship with co ntact varia b les. By introduc ing distance v ariab les, w e m ay also optimiz e the distance probab ility , as oppo sed to the co ntact pro bability used by our cur rent I L P method. Fur ther, we can also introduc e va ri a ble s of b eta- sheet ( gro up of beta-str ands) to captur e more global p roperties of a co ntact map. One may ask ho w our approac h c om pares to a mo del -b ased filte ring strateg y i n wh ich 3D mode ls are built base d on ini tial predic ted c o n- tacts and then used to fi lter incorrec t p redictio ns. Ou r m ethod di ff ers f rom this general “mo del - based fi ltering ” str ategy i n a co uple of a s- pects . First , it is ti me-co nsuming to build thousa nds or at l eas t hund reds of 3D models wi th initial pr edicted co ntacts. I n co ntrast, our m et h- od c an do contact pred iction (us ing phy sical constrain ts) withi n minu tes after PSI- BLA ST search is done. Seco nd, the quality of the 3D mo dels is also determ ined by other facto rs such as energy functio n and energ y optimization (or co nfo rmation sam pling) metho ds w hi le our metho d i s i ndepen den t of t hese fac tors . Even if the energy func ti on is v ery acc urate, the energ y o ptimization alg orithm o ften is trapped t o loc al m i nima becau se the energ y functio n i s not r ugg ed. Tha t is, the 3D model s resul ting from e nergy minim izat ion are biased towa rds a specific region of the co nfo r mation space, unless an extreme ly large sca le of co nform ati on sampl ing is co nduc ted. Therefo re, the pred icte d co ntacts deriv ed fro m these mo de ls may also suffer from t his “loc al m inima” issue. By co ntrast, our integ er prog ramm ing metho d can searc h thro ugh the who le co nfo rmation spac e and f ind the globa l optimal solution and thus, is not b iased to any l oc al min ima re gio n. By using o ur pr edicted c ontacts as c onstraints, we may pinpoint t o the g o od r egio n o f a co nfo rmation space (w ithou t be ing b iased by l oc al minima ), reduc e the search spac e and sign ificantly speed up co nfo r mation se arch . 5 A CKNOW LEDGEMENTS The authors are gratef ul to anony m ous reviewe rs for their insig h tfu l co mm ents and to the Univ ersity o f Chicag o Beagle team and Tera Grid fo r their suppo rt of com putati ona l reso urces. F unding : Thi s work is suppo r ted by the Nationa l I nstitute s of Health gr ant R01G M0897532 , Nationa l Scie nce Founda tion D BI-0960390, the NSF CA REER award and the A lfred P. Slo an R ese arch Fel lows hip. 6 REFERENC E Altschul, S.F. an d Koo nin, E.V. (1998) Ite ra ted prof ile sea r c h es wit h PSI -BL AST -- a too l for disco ve r y in protei n database s, Trends in Biochem ical Scien ces , 23 , 444-447. Predicting protein contact map usin g evolut ionary and phy sical constra ints by i nteger progr a mming (e xtend ed version) 13 Baú , D. , et al. (2 006) Distill: a suite of web se rvers fo r the predictio n o f one -, t w o- an d three -dime nsional structural fe atur es of protei ns, BMC Bioin formatics , 7 , 402. Breiman, L. (2001) R andom fo rests, Mach L earn , 45 , 5 -32. Brenne r , S .E., Ko ehl, P . and Le vitt, M. (2000) The AS TR AL c ompe n dium fo r protei n structure and seque nce anal ys is, Nucleic Ac ids Res , 28 , 254- 256. Capra r a, A. , et al. (20 04) 1001 optima l PD B st r uctu r e align ments: i n teger prog rammi n g met h ods f o r fin ding the m aximum co nt act map o ve rl ap, J Compu t Biol , 11 , 27-52 . Cheng, J. and B aldi, P. (2007) Improv ed r esidue co nt act prediction using suppo rt v ecto r ma c hin es and a la rge f eature s et, BMC Bioin formatics , 8 , 113 . CPL EX, I.B .M.I. (2009 ) V12. 1: Use r’s Manual fo r CPL EX, Inte rna tional Bus iness Mac hines Corp oration . Di Lena, P., Nag ata, K. and B aldi, P. (20 12) De ep Architec tur es f or Protei n Co ntact Map P redictio n, B ioin formatics . Fariselli, P. and Casad io, R. (1999) A neu ral netw o r k based pr edicto r of r esidue co ntacts i n prote ins, Pr otein Eng , 12 , 15-21. Gö be l, U. , et al. (2004) Correl at ed mutatio n s and residue co ntacts i n proteins , Pr oteins , 18 , 309- 317. Jo n es, D.T. (1999) Protei n se co n dary structure prediction based on positio n -spec ific scoring matrice s, J M ol B iol , 292 , 1 95- 202. Jo n es, D.T . , et al. (2012) PSICOV: precise s tructu ral co n tac t prediction using sparse inverse co v ar iance estima tion o n large multiple sequence alignme nt s, Bioinform atics , 28 , 184 -190. Klepe is, J . a n d F loudas, C. (2003) ASTRO-FOL D: a comb inato r ial and glob al opt imizatio n framew ork fo r a b initi o predictio n of thr ee-dime n sio nal st r uctu r es o f protei ns from the amino acid sequence , Bi ophys J , 85 , 2119. Li, W . a nd Godzi k, A. (2006) Cd -hit: a fast prog ram fo r c luste r ing and co mparing la r ge sets of pr otei n or nucleotide seque nces, Bioinform atics , 22 , 1658-16 59. Li, W., Jaroszew ski, L. an d Godzik , A. (2001) Clus tering o f highly h omo logo us sequences to r educ e the size of lar ge pro tein datab ases, Bioinformat ics , 17 , 282-2 83. Liaw , A. and Wie ner, M. (200 2) Classific ation and R egressi o n by r ando mForest, R new s , 2 , 18-22. Mo r co s, F . , e t al. (2 011) Di rect-co upling analy sis o f r esidue c oevo lution cap tures native contacts a cross m any pr otei n families, Pr oc Natl Ac ad Sci U S A , 108 , E1293- E1301 . Olmea, O. and V ale n cia, A. (1997) Imp rov in g contact predic tions by the c omb in atio n of co rr elated muta tio n s and other so ur ce s of se quence in fo rmation, Fold ing and Desig n , 2 , S25-S 32. Ortiz, A.R . , et al. (19 99) Ab initio fo lding o f pr oteins using r estraints de ri ve d from evolutio nary i n fo rmatio n , Prot eins , 37 , 177-185. Punta, M . and Ro st, B. (2005) PRO Fcon: nove l pr ed iction of lo ng -range co nta cts , Bioinfor matics , 21 , 2960-2968. Tan, Y.H ., Huang, H . a nd Kihara, D. (20 06) S t atistic al potenti al -b ased amino acid s imilarity mat r ices for aligning dista ntly related prote in seque nces, Proteins , 64 , 587-600. Tegge, A.N. , et al. (2009) NNc on: imp rov ed pr otein co ntact map predictio n usi n g 2D-recursive n eural netw orks, Nu cleic Acids Res , 37 , W515-W 518 . Vassura, M. , et al. (2008) Rec onstructio n of 3D struct ur es from protei n contact maps, Compu tational Biology and Bioinforma tics, IE EE/ACM Transa ctions on , 5 , 357-367 . Vendrusco lo , M. and Doma n y , E. (1998) Pai rwise co nta ct potentials are unsuitable fo r pr ote in fo lding, The Jour nal of Chemical Physics , 109 , 11101 . Vendrusco lo , M., Kusse ll, E . and Domany , E. (1997) Re cove r y of protei n st r ucture f r o m c ontact maps, Folding and Des ign , 2 , 295-306. Vullo, A ., W als h , I. an d Pollastri, G . (20 06) A two-stage appr oac h fo r improv ed pr ediction o f residue c ontact maps, BMC Bioinforma tics , 7 , 180. Z. Wang et al. 14 Wang, G. and Dunb rack, R.L.J. (2003) PISCES: a p rotein se quence c ulling se r ve r , Bioinform atics , 19 , 158 9-1591. Wang, S. , et al. (2013) P rotei n st ructure alig nment bey o n d spatial proximi ty , Scientifi c Reports , 144 8. Wu, S., Szil agy i, A. and Z hang, Y. (2011) Imp r o ving p rotei n structure pred iction using multiple sequence -base d contact predictio n s, S tructure , 19 , 1182- 1191. Wu, S. a nd Zhang, Y. (2008) A comp r e hen s ive asse ssme nt of sequence-b as ed and template-base d m et h ods f o r p rotei n co nt act pred iction, Bioin formatics , 24 , 924-9 31. Xu, J., Jiao, F. and B erger, B. (2006) A para meterized algorit hm f or p r ote in structu r e a lig nment. Re s earch i n Computa tional Molecular Biology . Sp ringer, pp. 488-499 . Zhao, F. and Xu, J . (2012) A Po sition-Spe cific Distance-D epende nt Stat istical Po te nt ial fo r Prote in Structu r e and Functio nal Study , Structure (London , Eng land : 1993) , 20 , 1118-112 6. Zhou, H. and Skol n ick, J. (2007) Protei n model quality a sse ssment predict ion by comb in ing fragme nt compa r isons a n d a co nsensus Cα co nt act po tential, Prot eins , 71 , 1211-1218 .
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment