vSMC: Parallel Sequential Monte Carlo in C++
Sequential Monte Carlo is a family of algorithms for sampling from a sequence of distributions. Some of these algorithms, such as particle filters, are widely used in the physics and signal processing researches. More recent developments have establi…
Authors: Yan Zhou
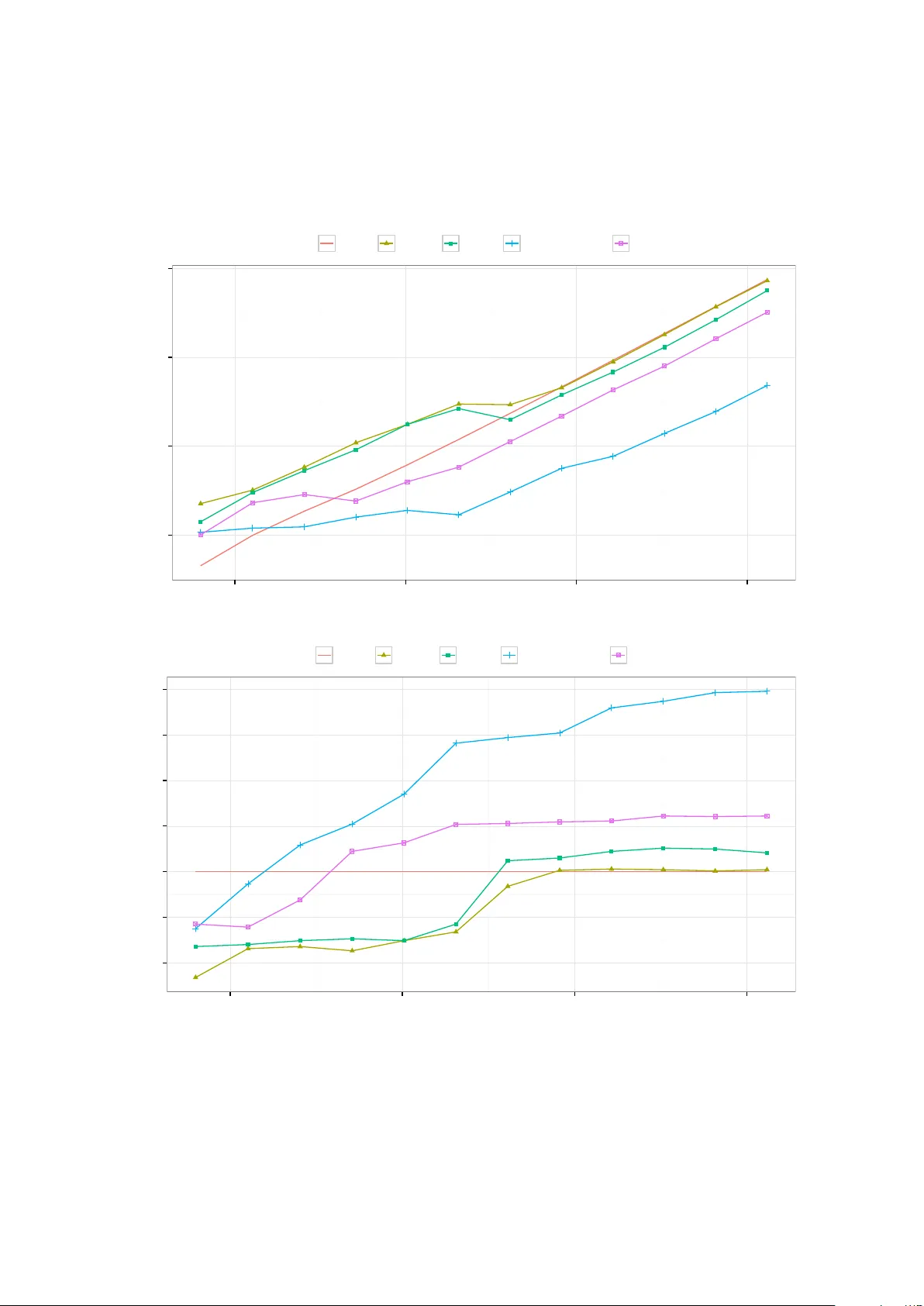
vSMC: P arallel Sequen tial Mon te Carlo in C++ Y an Zhou Univ ersity of W arwic k Abstract Sequen tial Mon te Carlo is a family of algorithms for sampling from a sequence of distributions. Some of these algorithms, suc h as particle filters, are widely used in the ph ysics and signal pro cessing researc hes. More recen t dev elopments ha ve established their application in more general inference problems such as Ba yesian mo deling. These algorithms ha ve attracted considerable atten tions in recen t y ears as they ad- mit natural and scalable parallelizations. Ho w ev er, these algorithms are p erceived to be difficult to implemen t. In addition, parallel programming is often unfamiliar t o man y researc hers though conceptually app ealing, esp ecially for sequential Mon te Carlo related fields. A C++ template library is presented for the purp ose of implementing general sequen- tial Mon te Carlo algorithms on parallel hardw are. Two examples are presen ted: a simple particle filter and a classic Bay esian mo deling problem. Keywor ds : Monte Carlo, particle filter, C++ template generic programming. 1. In tro duction Sequen tial Mon te Carlo (SMC) metho ds are a class of sampling algorithms that combine imp ortance sampling and resampling. They hav e b een primarily used as “particle filters” to solv e optimal filtering problems; see, for example, Capp ´ e, Go dsill, and Moulines ( 2007 ) and Doucet and Johansen ( 2011 ) for recen t reviews. They are also used in a static setting where a target distribution is of in terest, for example, for the purp ose of Bay esian mo deling. This w as prop osed b y Del Moral, Doucet, and Jasra ( 2006b ) and developed by Peters ( 2005 ) and Del Moral, Doucet, and Jasra ( 2006a ). This f ramew ork inv olves the construction of a sequence of artificial distributions on spaces of increasing dimensions whic h admit the distributions of in terest as particular marginals. SMC algorithms are p erceiv ed as b eing difficult to implemen t while general to ols were not a v ailable until the dev elopment by Johansen ( 2009 ), which provided a general framework for implemen ting SMC algorithms. SMC algorithms admit natural and scalable parallelization. Ho wev er, there are only parallel implemen tations of SMC algorithms for man y problem sp e- cific applications, usually asso ciated with sp ecific SMC related researc hes. Lee, Y au, Giles, Doucet, and Holmes ( 2010 ) studied the parallelization of SMC algorithms on GPUs with some generalit y . There are few general to ols to implemen t SMC algorithms on parallel hardware though multicore CPUs are very common to day and computing on sp ecialized hardware suc h as GPUs are more and more popular. The purp ose of the current w ork is to provide a general framework for implementing SMC al- 2 vSMC: Parallel Sequential Monte Carlo in C++ gorithms on both sequen tial and parallel h ardw are. There are t wo main goals of the presen ted framew ork. The first is reusabilit y . It will b e demonstrated that the same implemen tation source co de can b e used to build a serialized sampler, or using differen t programming mo d- els (for example, Op enMP ( OpenMP Arc hitecture Review Board 2011 ) and Intel TBB ( Intel Co op eration 2013c )) to build parallelized samplers for multicore CPUs. They can b e scaled for clusters using MPI ( Message Passing Interface F orum 2012 ) with few mo difications. And with a little effort they can also b e used to build parallelized samplers on sp ecialized mas- siv e parallel hardware suc h as GPUs using OpenCL ( Kronos Op enCL W orking Group 2012 ). The second is extensibility . It is p ossible to write a back end for vSMC to use new parallel programming mo dels while reusing existing implementations. It is also p ossible to enhance the library to impro ve p erformance for sp ecific applications. Almost all comp onen ts of the library can b e reimplemen ted by users and thus if the default implementation is not suitable for a sp ecific application, they can b e replaced while b eing int egrated with other comp onents seamlessly . 2. Sequen tial Mon te Carlo 2.1. Sequen tial imp ortance sampling and resampling Imp ortance sampling is a technique whic h allows the calculation of the exp ectation of a function ϕ with resp ect to a distribution π using samples from some other distribution η with resp ect to which π is absolutely contin uous, based on the identit y , E π [ ϕ ( X )] = Z ϕ ( x ) π ( x ) d x = Z ϕ ( x ) π ( x ) η ( x ) η ( x ) d x = E η h ϕ ( X ) π ( X ) η ( X ) i (1) And thus, let { X ( i ) } N i =1 b e samples from η , then E π [ ϕ ( X )] can b e approximated by ˆ ϕ 1 = 1 N N X i =1 ϕ ( X ( i ) ) π ( X ( i ) ) η ( X ( i ) ) (2) In practice π and η are often only kno wn up to some normalizing constants, which can b e estimated using the same samples. Let w ( i ) = π ( X ( i ) ) /η ( X ( i ) ), then we ha v e ˆ ϕ 2 = P N i =1 w ( i ) ϕ ( X ( i ) ) P N i =1 w ( i ) (3) or ˆ ϕ 3 = N X i =1 W ( i ) ϕ ( X ( i ) ) (4) where W ( i ) ∝ w ( i ) and are normalized suc h that P N i =1 W ( i ) = 1. Sequen tial imp ortance sampling (SIS) generalizes the imp ortance sampling technique for a sequence of distributions { π t } t ≥ 0 defined on spaces { Q t k =0 E k } t ≥ 0 . A t time t = 0, sample { X ( i ) 0 } N i =1 from η 0 and compute the weigh ts W ( i ) 0 ∝ π 0 ( X ( i ) 0 ) /η 0 ( X ( i ) 0 ). A t time t ≥ 1, each sample X ( i ) 0: t − 1 , usually termed p articles in the literature, is extended to X ( i ) 0: t b y a prop osal Zhou, Y an 3 distribution q t ( ·| X ( i ) 0: t − 1 ). And the weigh ts are recalculated by W ( i ) t ∝ π t ( X ( i ) 0: t ) /η t ( X ( i ) 0: t ) where η t ( X ( i ) 0: t ) = η t − 1 ( X ( i ) 0: t − 1 ) q t ( X ( i ) 0: t | X ( i ) 0: t − 1 ) (5) and thus W ( i ) t ∝ π t ( X ( i ) 0: t ) η t ( X ( i ) 0: t ) = π t ( X ( i ) 0: t ) π t − 1 ( X ( i ) 0: t − 1 ) η t − 1 ( X ( i ) 0: t − 1 ) q t ( X ( i ) 0: t | X ( i ) 0: t − 1 ) π t − 1 ( X ( i ) 0: t − 1 ) = π t ( X ( i ) 0: t ) q t ( X ( i ) 0: t | X ( i ) 0: t − 1 ) π t − 1 ( X ( i ) 0: t − 1 ) W ( i ) t − 1 (6) and imp ortance sampling estimate of E π t [ ϕ t ( X 0: t )] can b e obtained using { W ( i ) t , X ( i ) 0: t } N i =1 . Ho wev er this approach fails as t b ecomes large. The weigh ts tend to b ecome concentrated on a few particles as the discrepancy b etw een η t and π t b ecomes larger. Resampling techniques are applied such that, a new particle system { ¯ W ( i ) t , ¯ X ( i ) 0: t } M i =1 is obtained with the prop ert y , E h M X i =1 ¯ W ( i ) t ϕ t ( ¯ X ( i ) 0: t ) i = E h N X i =1 W ( i ) t ϕ t ( X ( i ) 0: t ) i (7) In practice, the resampling algorithm is usually chosen such that M = N and ¯ W ( i ) = 1 / N for i = 1 , . . . , N . Resampling can b e performed at each time t or adaptively based on some criteria of the discrepancy . One p opular quan tit y used to monitor the discrepancy is effe ctive sample size (ESS), in tro duced by Liu and Chen ( 1998 ), defined as ESS t = 1 P N i =1 ( W ( i ) t ) 2 (8) where { W ( i ) t } N i =1 are the normalized weigh ts. And resampling can b e p erformed when ESS ≤ αN where α ∈ [0 , 1]. The common practice of resampling is to replicate particles with large weigh ts and discard those with small w eights. In other words, instead of generating a random sample { ¯ X ( i ) 0: t } N i =1 directly , a random sample of integers { R ( i ) } N i =1 is generated, such that R ( i ) ≥ 0 for i = 1 , . . . , N and P N i =1 R ( i ) = N . And eac h particle v alue X ( i ) 0: t is replicated for R ( i ) times in the new particle system. The distribution of { R ( i ) } N i =1 shall fulfill the requirement of Equation 7 . One such distribution is a multinomial distribution of size N and weigh ts ( W ( i ) t , . . . , W ( N ) t ). See Douc, Capp ´ e, and Moulines ( 2005 ) for some commonly used resampling algorithms. 2.2. SMC samplers SMC samplers allo w us to obtain, iterativ ely , collections of weigh ted samples from a sequence of distributions { π t } t ≥ 0 o ver essentially any random v ariables on some spaces { E t } t ≥ 0 , by constructing a sequence of auxiliary distributions { ˜ π t } t ≥ 0 on spaces of increasing dimensions, ˜ π t ( x 0: t ) = π t ( x t ) Q t − 1 s =0 L s ( x s +1 , x s ), where the sequence of Marko v kernels { L s } t − 1 s =0 , termed bac kward kernels, is formally arbitrary but critically influences the estimator v ariance. See Del Moral et al. ( 2006b ) for further details and guidance on the selection of these ke rnels. 4 vSMC: Parallel Sequential Monte Carlo in C++ Standard sequential imp ortance sampling and resampling algorithms can then b e applied to the sequence of syn thetic distributions, { ˜ π t } t ≥ 0 . A t time t − 1, assume that a set of w eighted particles { W ( i ) t − 1 , X ( i ) 0: t − 1 } N i =1 appro ximating ˜ π t − 1 is av ailable, then at time t , the path of each particle is extended with a Marko v k ernel say , K t ( x t − 1 , x t ) and the set of particles { X ( i ) 0: t } N i =1 reac h the distribution η t ( X ( i ) 0: t ) = η 0 ( X ( i ) 0 ) Q t k =1 K t ( X ( i ) t − 1 , X ( i ) t ), where η 0 is the initial distribution of the particles. T o correct the discrepancy b etw een η t and ˜ π t , Equation 6 is applied and in this case, W ( i ) t ∝ ˜ π t ( X ( i ) 0: t ) η t ( X ( i ) 0: t ) = π t ( X ( i ) t ) Q t − 1 s =0 L s ( X ( i ) s +1 , X ( i ) s ) η 0 ( X ( i ) 0 ) Q t k =1 K t ( X ( i ) t − 1 , X ( i ) t ) ∝ ˜ w t ( X ( i ) t − 1 , X ( i ) t ) W ( i ) t − 1 (9) where ˜ w t , termed the incr emental weights , are calculated as, ˜ w t ( X ( i ) t − 1 , X ( i ) t ) = π t ( X ( i ) t ) L t − 1 ( X ( i ) t , X ( i ) t − 1 ) π t − 1 ( X ( i ) t − 1 ) K t ( X ( i ) t − 1 , X ( i ) t ) (10) If π t is only kno wn up to a normalizing constant, say π t ( x t ) = γ t ( x t ) / Z t , then we can use the unnormalize d incremen tal weigh ts w t ( X ( i ) t − 1 , X ( i ) t ) = γ t ( X ( i ) t ) L t − 1 ( X ( i ) t , X ( i ) t − 1 ) γ t − 1 ( X ( i ) t − 1 ) K t ( X ( i ) t − 1 , X ( i ) t ) (11) for imp ortance sampling. F urther, with the previously normalize d we igh ts { W ( i ) t − 1 } N i =1 , we can estimate the ratio of normalizing constan t Z t / Z t − 1 b y ˆ Z t Z t − 1 = N X i =1 W ( i ) t − 1 w t ( X ( i ) t − 1 , X ( i ) t ) (12) Sequen tially , the normalizing constan t betw een initial distribution π 0 and some target π T , T ≥ 1 can b e estimated. See Del Moral et al. ( 2006b ) for details on calculating the incremen tal w eights. In practice, when K t is inv ariant to π t , and an appro ximated suboptimal bac kward k ernel L t − 1 ( x t , x t − 1 ) = π ( x t − 1 ) K t ( x t − 1 , x t ) π t ( x t ) (13) is used, the unnormalized incremental weigh ts will b e w t ( X ( i ) t − 1 , X ( i ) t ) = γ t ( X ( i ) t − 1 ) γ t − 1 ( X ( i ) t − 1 ) . (14) 2.3. Other sequen tial Monte Carlo algorithms Some other commonly used sequential Mon te Carlo algorithms can b e viewed as sp ecial cases of algorithms introduced ab o v e. The annealed imp ortance sampling (AIS; Neal ( 2001 )) can b e viewed as SMC samplers without resampling. P article filters as seen in the ph ysics and signal pro cessing literature, can also b e interpret ed as the sequential imp ortance sampling and resampling algorithms. See Doucet and Johansen Zhou, Y an 5 ( 2011 ) for a review of this topic. A simple particle filter example is used in Section 5.1 to demonstrate basic features of the vSMC library . 3. Using the vSMC library 3.1. Ov erview The vSMC library makes use of C++ ’s template generic programming to implement general SMC algorithms. This library is formed by a few ma jor mo dules listed b elow. Some features not included in these mo dules are introduced later in con text. Core The highest level of abstraction of SMC samplers. Users int eract with classes defined within this module to create and manipulate general SMC samplers. Classes in this mo dule include Sampler , Particle and others. These classes use user defined callback functions or callable ob jects, such as functors, to p erform problem sp ecific op erations, suc h as updating particle v alues and weigh ts. This module is documented in Section 4.1 . Symmetric Multipro cessing (SMP) This is the form of computing most p eople use ev- eryda y , including multiprocessor workstations, multicore desktops and laptops. Classes within this mo dule make it p ossible to write generic op erations whic h manipulate a sin- gle particle that can be applied either sequen tially or in parallel through v arious parallel programming mo dels. A metho d defined through classes of this mo dule can b e used by Sampler as callback ob jects. This mo dule is documented in Section 4.2 . Message P assing In terface MPI is the de facto standard for parallel programming on dis- tributed memory architectur es. This mo dule enables users to adapt implementati ons of algorithms written for the SMP mo dule such that the same sampler can b e parallelized using MPI . In addition, when used with the SMP module, it allo ws easy implemen tation of h ybrid parallelization suc h as MPI / OpenMP . In Section 5.2 , an example is shown ho w to extend existing SMP parallelized samplers using this mo dule. Op enCL This mo dule is similar to the tw o ab o ve except it eases the parallelization through Op enCL , such as for the purp ose of General Purpose GPU Programming (GPGPU). Op enCL is a framework for writing programs that can b e execute across heterogeneous platforms. OpenCL programs can run on either CPUs or GPUs. It is beyond the scop e of this pap er to give a prop er introduction to GPGPU, Op enCL and their use in vSMC . Ho wev er, later we will demonstrate the relativ e performance of this programming model on b oth CPUs and GPUs in Section 5.2 . 3.2. Obtaining and installation vSMC is a header only library . There is practically no installation step. The library can b e do wnloaded from http://zhouyan.github.io/vSMC /vSMC.zip . After downloading and unpac king, one can start using vSMC by ensuring that the compiler can find the headers inside the include directory . T o p ermanently install the library in a system directory , one can simply copy the con tents of the include directory to an appropriate place. 6 vSMC: Parallel Sequential Monte Carlo in C++ Alternativ ely , one can use the CMake ( Martin and Hoffman 2010 ) (2.8 or later) configuration script and obtain the source by Git ( T orv alds et al. 2013 ). On a Unix-like system (suc h as Mac OS X, BSD, Lin ux and others), git cl one git:/ /github.com/zhouyan/vSMC.g it cd vSM C git su bmodule init git su bmodule update mkdir build cd bui ld cmake .. -DCMAKE_BUILD_TYPE=Releas e make install F or Unix-like systems, there is also a shell script build.sh that builds all examples in this pap er and pro duces the results and b ench marks as in Section 5 . See do cumentations in that script for details of how to change settings for the users’ platform. 3.3. Do cumenta tion T o build the reference man ual, one need Do xygen ( v an Heesch 2013 ), version 1.8.3 or later. Con tinuing the last step (still in the build directory), in voking make docs will create a doc directory inside build , whic h con tains the HTML references. Alternativ ely the reference man ual can also b e found on http://zhouyan.github.io/vSMC/doc/ html/index.html . It is b eyond the scop e of this pap er to do cumen t every feature of the vSMC library . In man y places we will refer to this reference manual for further information. 3.4. Third-part y dep endencies vSMC uses Random123 ( Salmon, Moraes, Dror, and Shaw 2011 ) count er-based RNG for random num b er generating. F or an SMC sampler with N particles, vSMC constructs N (sta- tistically) indep endent RNG streams. It is p ossible to use millions of suc h streams without a h uge memory fo otprint or other p erformance penalties. Since eac h particle has its o wn indep enden t RNG stream, it frees users from man y thread-safet y and statistical indep en- dence considerations. It is highly recommended that the users install this library . Within vSMC , these RNG streams are wrapp ed under C++11 RNG engines, and can b e replaced b y other compatible RNG engines seamlessly . Users only need to b e familiar with classes defined in C++11 or their Bo ost ( Da wes et al. 2013 ) equiv alents to use these RNG streams. See the do cumentation of the corresp onding libraries for details, as w ell as examples in Section 5 . The other third-part y dep endency is the Bo ost library . V ersion 1.49 or later is required. Ho wev er, this is actually optional provided that proper C++11 features are av ailable in the standard library , for example using clang ( The LL VM Developer Group 2013a ) with libc++ ( The LL VM Developer Group 2013b ). The C++11 headers of concern are and . T o instruct vSMC to use the standard library headers instead of falling back to the Bo ost library , one needs to define the configuration macros b efore including an y vSMC headers. F or example, clang++ -std=c++11 -stdlib=libc++ \ Zhou, Y an 7 -DVSMC_HAS_CXX11LIB_FUN CTIONAL=1 \ -DVSMC_HAS_CXX11LIB_RAN DOM=1 \ -o pro g prog.cp p tells the library to use C++11 and . The av ailabilit y of these headers are also chec ked by the CMake configuration script. 3.5. Compiler supp ort vSMC has b een tested with recent v ersions of clang , GCC ( GNU Pro ject 2013 ), Intel C++ Complier ( In tel Co op eration 2013a ) and Microsoft Visual C++ ( Microsoft Coop eration 2012 ). vSMC can optionally use some C++11 features to improv e performance and usability . In particular, as mentioned b efore, vSMC can use C++11 standard libraries instead of the Bo ost library . At the time of writing, clang with lib c++ has the most comprehensiv e supp ort of C++11 with resp ect to standard compliant and feature completion. GCC 4.8 , Microsoft Visual C++ 2012 and Intel C++ Complier 2013 also ha v e very goo d C++11 supp ort. Note that, pro vided the Bo ost library is a v ailable, all C++11 language and library features are optional. vSMC can b e used with an y C++98 conforming compilers. 4. The vSMC library 4.1. Core mo dule The core mo dule abstracts general SMC samplers. SMC samplers can b e viewed as formed b y a few concepts regardless of the sp ecific problems. The following is a list of the most commonly seen comp onents of SMC samplers and their corresp onding vSMC abstractions. • A collection of all particle state v alues, namely { X ( i ) t } N i =1 . In vSMC , users need to define a class, sa y T , to abstract this concept. W e refer to this as the value c ol le ction . W e will slightly abuse the generic name T in this pap er. Wheneve r a template parameter is men tioned with the name T , it alwa ys refers to such a v alue collection t yp e unless stated otherwise. • A collection of all particle state v alues and their asso ciated w eights. This is abstracted b y a Particle ob ject. W e refer to this as the p article c ol le ction . A Particle ob ject has three primary sub-ob jects. One is the ab ov e type T v alue collection ob ject. Another is an ob ject that abstracts weigh ts { W ( i ) t } N i =1 . By default this is a WeightSet ob ject. The last is a collection of RNG streams, one for each particle. By default this is an RngSet ob ject. • Op erations that p erform tasks common to all samplers to these particles, suc h as re- sampling. These are the member functions of Particle . • A sampler that up dates the particles (state v alues and w eights) using user defined callbac ks. This is a Sampler ob ject. • Monitors that record the imp ortance sampling estimates of certain functions defined for the v alues when the sampler iterates. These are Monitor ob jects. A monitor for 8 vSMC: Parallel Sequential Monte Carlo in C++ the estimates of E [ h ( X t )] computes h ( X ( i ) t ) for each i = 1 , . . . , N . The function v alue h ( X t ) is allow ed to b e a v ector. Note that within the core mo dule, all op erations are applied to Particle ob jects, that is { W ( i ) t , X ( i ) t } N i =1 , instead of a single particle. Later we will see how to write op erations that can b e applied to individual particles and can b e parallelized easily . Pr o gr am structur es A vSMC program usually consists of the follo wing tasks. • Define a v alue collection type T . • Constructing a Sampler ob ject. • Configure the b ehavior of initialization and up dating b y adding callable ob jects to the sampler ob ject. • Optionally add monitors. • Initialize and iterate the sampler. • Retriev e the outputs, estimates and other informations. In this section we do cumen t how to implement eac h of these tasks. Within the vSMC library , all public classes, namespaces and free functions, are declared in the namespace vsmc . The value c ol le ction The template parameter T is a user defined type that abstracts the v alue collection. vSMC do es not restrict how the v alues shall b e actually stored. They can b e stored in memory , spread among no des of a cluster, in GPU memory or even in a database. Ho wev er this kind of flexibility comes with a small price. The v alue collection do es need to fulfill tw o requiremen ts. W e will see later that for most common usage, vSMC pro vides readily usable implemen tations, on top of which users can create problem sp ecific classes. First, the v alue collection class T has to provide a constructor of the form T (Siz eType N) where SizeType is some integer type. Since vSMC allows one to allo cate the states in an y w ay suitable, one needs to pro vide this constructor whic h Particle can use to allo cate them. Second, the class has to pro vide a mem b er function named copy that copies each particle according to replication num b ers given by a resampling algorithm. F or the same reason as ab o v e vSMC has no wa y to know how it can extract and copy a single particle when it is doing the resampling. The signature of this member function ma y lo ok like template void copy (std::size_t N, const SizeType *copy_from); Zhou, Y an 9 The p oin ter copy_from p oints to an arra y that has N elemen ts, where N is the num b er of particles. After calling this member function, the v alue of particle i shall b e copied from the particle j = copy_from[i] . In other words, particle i is a child of particle copy_from[i] , or copy_from[i] is the paren t of particle i . If a particle j shall remain itself, then copy_from[j] == j . How the v alues are actually copied is user defined. Note that, the member function can tak e other forms, as usual when using C++ template generic programming. The actual t yp e of the p oin ter copy_from , SizeType , is Particle::size_type , whic h dep ends on the type T . F or example, define the member function as the following is also allow ed, void copy (int N, const std ::size_t *copy_from); pro vided that Particle::size_type is indeed std::size_t . Ho w ev er, writing it as a mem b er function template releases the users from finding the actual type of p oin ter copy_from and the sometimes troubling forw ard declaration issues. Will not elab orate suc h more tec h- nical issues further. Constructing a sampler Once the v alue collection class T is defined. One can start constructing SMC samplers. F or example, the following line creates a sampler with N particles Sampler sampler(N); The num b er of particles is the only mandatory argumen t of the constructor. There are t w o optional parameters. The complete signature of the constructor is, explicit Sampler::Sampler (Samp ler::size_type N, ResampleScheme scheme = Stratified, double threshold = 0.5); The scheme parameter is self-explanatory . vSMC provides a few built-in resampling schemes; see the reference manual for a list of them. User defined resampling algorithms can also b e used. See the reference man ual for details. The threshold is the threshold of ESS / N b elo w whic h a resampling will be performed. It is ob vious that if threshold ≥ 1 then resampling will b e p erformed at each iteration. If threshold ≤ 0 then resampling will never b e p erformed. Both parameters can b e c hanged later. Ho wev er the size of the sampler can never b e changed after the construction. Initialization and up dating All the callable ob jects that initi alize, mo v e and w eigh t the particle collection can b e added to a sampler through a set of mem b er functions. All these ob jects op erate on the Particle ob ject. Because vSMC allows one to manipulate the particle collect ion as a whole, in principle man y kinds of parallelization are possible. T o set an initialization metho d, one need to implement a function with t he following signature, std::size_t init_func (Particle &particle, void *param); or a class with operator() prop erly defined, such as 10 vSMC: Parallel Sequential Monte Carlo in C++ struct init_class { std: :size_t operator() (Particle &particle, void *param); }; They can b e added to the sampler through sampler.init(init_func) ; or sampler.init(init_class ()); resp ectiv ely . C++11 std::function or its Bo ost equiv alent boost::function can also b e used. F or example, std::function &, void *)> init_obj(init_func); sampler.init(init_obj); The addition of updating metho ds is more flexible. There are t wo kinds of updating metho ds. One is simply called move in vSMC , and is performed b efore the p ossible resampling at eac h iteration. These mov es usually p erform the updating of the weigh ts among other tasks. The other is called mcmc , and is performed after the p ossible resampling. They are often MCMC t yp e mo ves. Multiple move ’s or mcmc ’s are also allo wed. In fact a vSMC sampler consists of a queue of move ’s and a queue of mcmc ’s. The move ’s in the queue can b e changed through Sampler::move , Sampler &move (const Sampler::move_type &new_move, bool append); If append == true then new_move is app ended to the existing (p ossibly empt y) queue. Oth- erwise, the existing queue is cleared and new_move is added. The member function returns a reference to the updated sampler. F or example, the follo wing mo ve, std::size_t move_func (std::size_t iter, Par ticle &particle); can b e added to a sampler by sampler.move(move_func, false); This will clear the (p ossibly empty) existing queue of move ’s and set a new one. T o add m ultiple mo ves into the queue, sampler.move(move1, true).move(move2, true).move(move3, true) ; Ob jects of class t yp e with operator() properly defined can also b e used, similarly to the initialization metho d. The queue of mcmc ’s can b e used similarly . See the reference man ual for other metho ds that can b e used to manipulate these t wo queues. In principle, one can combine all mov es in to a single mo v e. How ever , sometimes it is more natural to think of a queue of mo ves. F or instance, if a multi-block Metrop olis random walk consists of kernels K 1 and K 2 , then one can implemen t eac h of them as functions, say mcmc_k1 and mcmc_k2 , and add them to the sampler sequentially , Zhou, Y an 11 sampler.mcmc(mcmc_k1, true).mcmc(mcmc_k2, true); Then at each iteration, they will b e applied to the particle collection sequen tially in the order in which they are added. R unning the algorithm, monitoring and outputs Ha ving set all the operations, one can initialize and iterate the sampler by calling sampler.initialize((voi d *)param ); sampler.iterate(iter_nu m); The param argumen t to initialize is optional, with NULL as its default. This parameter is passed to the user defined init_func . The iter_num argument to iterate is also optional; the default is 1. Before initializing the sampler or after a certain time p oint, one can add monitors to the sampler. The concept is similar to BUGS ( Spiegelhalter, Thomas, and Best 1996 )’s monitor statemen t, except it does not monitor the individual v alues but rather the importance sam- pling estimates. Consider approximating E [ h ( X t )], where h ( X t ) = ( h 1 ( X t ) , . . . , h m ( X t )) is an m -vector function. The importance sampling estimate can be obtained by AW where A is an N by m matrix where A ( i, j ) = h j ( X ( i ) t ) and W = ( W ( i ) t , . . . , W ( N ) t ) T is the N -vector of normalized weigh ts. T o compute this imp ortance sampling estimate, one need to define the follo wing ev aluation function (or a class with operator() prop erly defined), void monitor_eval (std::size_t iter, std::si ze_t m, const Particle &particle, double *res) and add it to the sampler by calling, sampler.monitor("some.u ser.chosen.variable.name", m, monito r_eval); When the function monitor_eval is called, iter is the iteration num b er of the sampler, m is the same v alue as the one the user passed to Sampler::monitor ; and th us one do es not need global v ariable or other similar techniques to access this v alue. The output p ointer res p oin ts to an N × m output arra y of row ma jor order. That is, after the calling of the function, res[i * dim + j] shall be h j ( X ( i ) t ). After each iteration of the sampler, the imp ortance sampling estimate will b e computed automatically . See the reference manual for v arious wa ys to retriev e the results. Usually it is sufficien t to output the sampler b y std::ofstream output("file.name"); output << sampler << std::endl; where the output file will contain the imp ortance sampling estimates among other things. Alternativ ely , one can use the Monitor::record member function to access sp ecific his- torical results. See the reference manual for details of v arious ov erloaded version of this mem b er function. 12 vSMC: Parallel Sequential Monte Carlo in C++ A refere nce to the v alue collecti on T ob ject can b e retriev ed through the Particle::value mem b er function. Particle ob jects manage the weigh ts through a weigh t set ob ject, whic h b y default is of type WeightSet . The Particle::weight_set mem b er function returns a reference to this w eigh set ob ject. A user defined wei gh t set class that abstracts { W ( i ) t } N i =1 can also b e used. The details in volv e some more adv anced C++ template tec hniques and are do cumented in the reference manual. One p ossible reason for replacing the default is to provide special memory managemen t of the weigh t set. F or example, the MPI mo dule pro vides a sp ecial w eigh t set class that manages w eights across multiple nodes and p erform prop er normalization, computation of ESS, and other tasks. The default WeightSet ob ject provides some w ays to retriev e weigh ts. Here we do cument some of the most commonly used. See the reference man ual for details of others. The weigh ts can b e accessed one b y one, for example, Particle &particle = sampler.particle(); double w_i = part icle.weight_set().weight(i ); double log_w_i = particle.weight_set().log_weight(i); One can also read all weigh ts into a con tainer, for example, std::vector w(particle.size()); particle.weight_set().r ead_weight(w.begin()); double *lw = new double[particle.size()]; particle.weight_set().r ead_log_weight(lw); Note that these mem b er functions accept general output iterators. Implementing initialization and up dating So far w e ha ve only discussed how to add initialization and up dating ob jects to a sampler. T o actually implement them, one writes callable ob jects that op erate on the Particle ob ject. F or example, a mov e can b e implemen ted through the follo wing function as mentioned b efore, std::size_t move_func (std::size_t iter, Par ticle &particle); Inside the bo dy of this function, one can change the v alue b y manipulating the ob ject through the reference returned by particle.value() . When using the default weigh t set class, the w eigh ts can b e up dated through a set of member functions of WeightSet . F or example, std::vector weight(particle.size()); particle.weight_set().s et_equal_weight(); particle.weight_set().s et_weight(weight.begin()); particle.weight_set().m ul_weight(weight.begin()); particle.weight_set().s et_log_weight(weight.begin ()); particle.weight_set().a dd_log_weight(weight.begin ()); Zhou, Y an 13 The set_equal_weight mem b er function sets all weigh ts to b e equal. The set_weight and set_log_weight member functions set the v alues of w eigh ts and logarithm weigh ts, resp ectiv ely . The mul_weight and add_log_weight member functions m ultiply the w eigh ts or add to the logarithm weigh ts by the given v alues, resp ectively . All these member functions accept general input iterators as their argumen ts. One imp ortan t thing to note is that, whenever one of these member functions is called, b oth the w eigh ts and logarithm w eigh ts will b e re-calculated, normalized, and the ESS will b e up dated. The reason for not allowing changing a single particle’s weigh t is that, in a m ulti- threading environmen t, it is p ossible for one to change one weigh t in one thread, and obtain another in another thread without prop er normalizing. Conceptually , changing one w eight actually changes all weigh ts. Gener ating r andom numb ers The Particle ob ject has a sub-ob ject, a collection of RNG engines that can b e used with C++11 or Bo ost distributions. F or each particle i , one can obtain an engine that pro duces an RNG stream indep enden t of others b y particle.rng(i); T o generate distribution random v ariates, one can use the C++11 library . F or example, std::normal_distributio n rnorm(mean, sd); double r = rnorm( particle.rng(i)); or use the Boost library , boost::random::normal_d istribution rnorm(mean, sd); double r = rnorm( particle.rng(i)); vSMC itself also makes use of C++11 or Bo ost dep ending on the v alue of the configuration macro VSMC_HAS_CXX11LIB_RAN DOM . Though the user is free to c ho ose whic h one to use in their own code, there is a conv enien t alternative. F or eac h class defined in C++11 , it is imported to the vsmc::cxx11 namespace. Therefore one can use cxx11::normal_distribut ion rnorm(mean, sd); while the underlying implemen tation of normal_distribut ion can be either C++11 standard library or Bo ost . The b enefit is that if one needs to develop on multiple platforms, and only some of them supp ort C++11 and some of them ha ve the Bo ost library , then only the configure macro VSMC_HAS_CXX11LIB_RANDOM needs to b e changed. This can b e configured through CMake and other build systems. Of course, one can also use an entirely differen t RNG system than those pro vided b y vSMC . 4.2. SMP mo dule 14 vSMC: Parallel Sequential Monte Carlo in C++ The value c ol le ction Man y typic al problems’ v alue collections can b e viewed as a matrix of certain t yp e. F or example, a simple particle filter whose state is a v ector of length Dim and type double can b e viewed as an N by Dim matrix where N is the num b er of particles. A trans-dimensional problem can use an N b y 1 matrix whose type is a user defined class, say StateType . F or this kind of problems, vSMC pro vide a class template template class StateMatrix; whic h pro vides the constructor and the copy member function required by the core mo dule in terface, as well as metho ds for accessing individual v alues. The first template parameter (p ossible v alue RowMajor or ColMajor ) sp ecifies how the v alues are ordered in memory . Usu- ally one shall choose RowMajor to optimize data access. The second template parameter is the n umber of v ariables, an integer v alue no less than 1 or the sp ecial v alue Dynamic , in whic h case StateMatrix provides a mem b er function resize_dim suc h that the num b er of v ariables can b e c hanged at run time. The third template parameter is the type of the state v alues. Eac h particle’s state is thus a vector of length Dim , indexed from 0 to Dim - 1 . T o obtain the v alue at p osition pos of the vector of particle i , one can use one of the followi ng member functions, StateBase value( N); StateType val = value.state(i, pos); StateType val = value.state(i, Position ()); StateType val = value.state(i); where Pos is a compile time constant expression whose v alue is the same as pos , assuming the p osition is kno wn at compile time. One can also read all v alues. T o read the v ariable at p osition pos , std::vector vec(value.size()); value.read_state(pos, vec.begin()); Or one can read all v alues through an iterator, std::vector mat(Dim * value.size()); value.read_state_matrix (ColMajor, mat.first()); Alternativ ely , one can also read all v alues through an iterator which p oints to iterators, std::vector > mat(Dim); for (s td::size_t i = 0; i != Dim; ++i) mat[i].resize(value.siz e()); std::vector::iterator> iter(Dim); for (s td::size_t i = 0; i != Dim; ++i) iter[i] = mat[i].begin(); value.read_state_matrix (iter.first()); Zhou, Y an 15 If the compiler support C++11 , vSMC also provides a StateTuple class template, whic h is similar to StateMatrix except that the types of v alues do not hav e to b e the same for eac h v ariable. This is similar to R ( R Core T eam 2013 )’s data.frame . F or example, supp ose eac h particle’s state is formed b y tw o double ’s, an int and a user defined type StateType , then the following constructs a v alue collection using StateTuple , StateTuple value(N); And there are a few wa ys to access the state v alues, similar to StateMatrix , double x0 = value .state(i, Position<0>()); int x2 = value.st ate<2>(i); std::vector vec(value.size()); state.read_state(Positi on<3>(), vec.begin()); See the reference man ual for details. A single p article F or a Particle ob ject, one can construct a SingleParticle ob ject that abstracts one of the particle from the collection. F or example, Particle particle(N); SingleParticle sp(i, &particle) ; create a SingleParticle ob ject corresp onding to the particle at p osition i . There are a few member functions of SingleParticle that makes access to individual particles easier than through the interface of Particle . Firt sp.id() returns the v alue of the argumen t i in the ab ov e co de that created this SingleParticle ob ject. In addition, sp.rng() is equiv alent to particle.rng(i) . Also sp.particle() returns a constant ref- erence to the Particle ob ject. And sp.particle_ptr() returns a p oin ter to such a constan t Particle ob ject. Note that, one cannot get write access to a Particle ob- ject through interface of SingleParticle . Instead, one can only get write access to a single particle. F or example, If T is a StateMatrix instan tiation or its deriv ed class, then sp.state(pos) is equiv alen t to particle.value().state(i, pos) and the reference it re- turns is m utable. See the reference man ual for more informations on the in terface of the SingleParticle class template. SingleParticle ob jects are usually not constructed by users, but rather b y the libraries’ other classes in the SMP module, and passed to user defined functions, as we will see ver y so on. Se quential and p ar al lel implementations Once we hav e the SingleParticle concept, we are ready to introduce how to write implemen tations of SMC algorithms that manipulate a single particle and can b e applied to all particles in parallel or sequentially . F or sequential implemen tations, this can b e done through five base classes, template class StateSEQ ; template class InitializeSEQ; 16 vSMC: Parallel Sequential Monte Carlo in C++ template class MoveSEQ; template class MonitorEvalSEQ; template class PathEvalSEQ; The template parameter BaseState needs to satisfy the general v alue collection requirements in addition to a copy_particle member function, for example, StateMatrix . Other base classes exp ect T to satisfy general v alue collection requirements. The details of all these class templates can b e found in the reference manual. Here w e use the MoveSEQ class as an example to illustrate their usage. Recall that Sampler exp ect a callable ob ject which has the following signature as a mov e, std::size_t move_func (std::size_t iter, Par ticle &particle); F or the purp ose of illustration, the t yp e T is defined as, typedef StateMatrix T; Here is a t ypical example of implemen tation of suc h a function, std::size_t move_func (std::size_t iter, Par ticle &particle) { std::vector inc_weight(particle.size()); for (P article::size_type i = 0; i != particle.size(); ++i) { cxx11::normal_distribut ion rnorm(0, 1); particle.value().state( i, 0) = rnorm(parti cle.rng(i)); inc_weight[i] = cal_inc_weight(par ticle.value().state(i, 0); } particle.weight_set().a dd_log_weight(inc_weight.b egin()); } where cal_inc_weight is some function that calculates the logarithm in cremen tal weigh ts. As w e see, there are three main parts of a t ypical mov e. First, we allo cate a vec tor inc_weight . Second, w e generate normal random v ariates for eac h particle and calculate the incremen tal w eights. This is done through a for loop. Third, w e add the logarithm incremental weigh ts. The first and the third are glob al op erations while the second is lo c al . The first and the third are often optional and absent. The lo cal op eration is also usually the most computational in tensive part of SMC algorithms and can b enefit the most from parallelizations. With MoveSEQ , one can derive from this class, whic h defines the operator() required b y the core mo dule in terface, std::size_t operator() (std::size_ t iter, Particle &particle); and customize what this op erator do es by defining one or more of the following three mem b er functions, corresp onding to the three parts, resp ectiv ely , void pre_processor (std::size_t iter, Partic le &pa rticle); std::size_t move_state (std::size_ t iter, SingleParticle sp); void post_processor (std::size_t iter, Parti cle &particle); Zhou, Y an 17 F or example, #include class move : publ ic MoveS EQ { public : void pre_processor (std::size_t iter, Partic le &pa rticle) {inc_weight_.resize(par ticle.size());} std::size_t move_state (std::size_ t iter, SingleParticle sp) { cxx11::normal_distribut ion rnorm(0, 1); sp.state(0) = rnorm(sp.rng()); inc_weight_[sp.id()] = cal_inc_wei ght(sp.state(0)); } void post_processor (std::size_t iter, Parti cle &particle) {particle.weight_set(). add_log_weight(inc_weight_ .begin());} private : std::vector inc_weight_; }; The operator() of MoveSEQ is equiv alent to the single function implementation as shown b efore. In the simplest case, MoveSEQ only tak es aw a y the lo op around Part 2. How ever if one implemen t the mov e in such a wa y , and then replace MoveSEQ with MoveOMP , the changi ng of the base class name causes vSMC to use Op enMP to parallelize the lo op. F or example, one can declare the class as #include class move : publ ic MoveO MP; and use exactly the same implemen tation as before. Now when move::operator () is called, it will be parallelized by OpenMP . Other bac ken ds are av ailable in case Op enMP is not a v ailable. Among them there are Cilk Plus ( In tel Co op eration 2011 ) and Intel TBB . In addition to these w ell kno wn parallelization programming models, vSMC also has its o wn implemen tation using C++11 . There are other back ends documented in the reference manual. T o use an y of these parallelization, all one need to do is to chan ge a few base class names. In practice, one can use conditional compilation, for example, to use a sequential implementation or a Op enMP parallelized one, w e can write, #ifdef USE_SEQ #include 18 vSMC: Parallel Sequential Monte Carlo in C++ #define BASE_MOVE MoveSEQ #endif #ifdef USE_OMP #include #define BASE_MOVE MoveOMP #endif class move : publ ic BASE_ MOVE; And we can compile the same source int o differen t samplers with Makefile rules suc h as the follo wing, prog-seq : prog.cpp $(CXX) $(CXXFLAGS) -DUSE_SEQ -o prog-seq prog.cpp prog-omp : prog.cpp $(CXX) $(CXXFLAGS) -DUSE_omp -o prog-omp prog.cpp Or one can configure the source file through a build system suc h as CMake , whic h can also determine whic h programming mo del is av ailable on the system. A dapter Sometimes, the cum b ersome task of writing a class t o implemen t a mo v e and other operations can out w eight the p ow er w e gain through ob ject orien ted programming. F or example, a simple move_state -lik e function is all that w e need to get the w ork done. In this case, one can create a mov e through the MoveAdapter . F or example, say we ha v e implemen ted std::size_t move_state (std::size_ t iter, SingleParticle sp); as a function. Then one can create a callabl e ob ject acceptable t o Sampler::move through MoveAdapter move_obj(move_state); MoveAdapter move_obj(move_state); MoveAdapter move_obj(move_state); MoveAdapter move_obj( move_state); MoveAdapter move_obj(move_state); sampler.move(move_obj, false); These are resp ectiv ely , sequen tial, C++11 , In tel TBB , Cilk Plus , and Op enMP implemen tations. The first template parameter is the type of v alue collection and the second is the name of the base class template. Actually , the MoveAdapter ’s constructor accepts tw o more optional argume n ts, the pre_processor and the post_processor , corresponding to the other t wo aforemen tioned member functions. Similar adapters for the other three base classes also exist. Another scenario where an adapter is desired is that whic h bac kend to use needs be decided at runtime. The ab ov e simple adapters can already b e used for this purp ose. In addition, another form of the adapter is as the follo wing, Zhou, Y an 19 class move; MoveAdapter move_obj; sampler.move(move_obj, false); where the class move has the same definition as b efore but it no longer deriv es from an y base class. The class move is required to ha v e a default constructor, a cop y constructor and an assignmen t operator. 4.3. Thread-safet y and scalabilit y considerations When implementing parallelized SMC algorithms, issues such as thread-safet y cannot b e a voided ev en though the vSMC library hides most parallel constructs from the user. Classes in the vSMC library usually guaran tee that their member functions are thread-safe in the sense that calling the same member function on different ob jects at the same time from differen t threads is safe. Ho wev er, calling mutabl e member functions on the same ob ject from differen t threads is usually not safe. Calling immutable mem b er functions is generally safe. There are a few exceptions, • The constructors of Particle and Sampler are not thread-safe. Therefore if one need to construct m ultiple Sampler from different threads, a mutex protection is needed. Ho wev er, subsequen t mem b er function calls on the constructed ob jects are thread-safe according to the abov e rules. • Mem b er functions that concern a single particle are generally thread-safe in the sense that one can call them on the same ob ject from different threads as long as they are called for differen t particles. F or example Particle::rng and StateMatrix::state are thread-safe. In general, one can safely manipulate differen t individual particles from differen t threads, whic h is the minimal requirement for scalable parallelization. But one cannot manipulate the whole particle collection from differen t threads, for example WeightSet::set_log_weight . User defined callbacks shall generally follow the same rules. F or example, for a MoveOMP sub- class, pre_proces sor and post_processor do es not hav e b e thread-safe, but move_state needs to be. In general, av oid write access to memory lo cations shared by all particles from move_state and other similar member functions. One needs to take some extra care when using third-par t y libraries. F or example, in our example implementation of the move class, the rnorm ob ject, whic h is used to generate Normal random v ariates, is defined within move_state instead of b eing a class member data even though it is created with the same parameters for eac h particle. This is b ecause the call rnorm(sp.rng()) is not thread-safe in some imple- men tations, for example, when using the Bo ost library . F or scalable p erformance, certain practices should b e av oided when implementing member functions such as move_state . F or example, dynamic memory allo cation is usually lo ck- based and thus should be a voided. Alternatively one can use a scalable memory allo cator suc h as the one pro vided b y In tel TBB . In general, in functions suc h as move_state , one should av oid using lo cks to guaran tee thread-safet y , which can b e a b ottleneck to parallel p erformance. 20 vSMC: Parallel Sequential Monte Carlo in C++ 5. Example 5.1. A simple particle filter The mo del and algorithm This is an example used in SMCTC ( Johansen 2009 ). Through this example, we will show ho w to re-implement a simple particle filter in vSMC . It shall walk one through the basic features of the library . SMCTC is the first general framew ork for implemen ting SMC algorithms in C++ and serves as one of the most imp ortant inspirations for the vSMC library . It is widely used b y man y researc hers. One of the goals of the current work is that users familiar with SMCTC shall find little difficulty in using the new library . The state space mo del, kno wn as the almost constan t v elo cit y model in the trac king literature, pro vides a simple scenario. The state v ector X t con tains the p osition and velocity of an ob ject moving in a plane. That is, X t = ( x t pos , y t pos , x t vel , y t vel ) T . Imp erfect observ ations Y t = ( x t obs , y t obs ) T of the positions are p ossible at eac h time instance. The state and observ ation equations are linear with additive noises, X t = AX t − 1 + V t Y t = B X t + αW t where A = 1 ∆ 0 0 0 1 0 0 0 0 1 0 0 0 0 1 B = 1 0 0 0 0 1 0 0 α = 0 . 1 and w e assume that the elemen ts of the noise vec tor V t are indep enden t Normal with v ariance 0 . 02 and 0 . 001 for p osition and v elo city , resp ectively . The observ ation noise, W t comprises indep enden t, iden tically distributed t -distributed random v ariables with degree of freedom ν = 10. The prior at time 0 corresp onds to an axis-aligned Gaussian with v ariance 4 for the p osition co ordinates and 1 for the velocity co ordinates. The particle filter algorithm is shown in Algorithm 1 . Implementations W e first introduce the b o dy of the main function, sho wing the t ypical w ork flo w of a vSMC program. #include #include #include #include const std::size_t DataNum = 100; const std::size_t ParticleNum = 1000; const std::size_t Dim = 4; Zhou, Y an 21 Initialization Set t ← 0. Sample x (0 ,i ) pos , y (0 ,i ) pos ∼ N (0 , 4) and x (0 ,i ) vel , y (0 ,i ) vel ∼ N (0 , 1). W eight W ( i ) 0 ∝ L ( X ( i ) 0 | Y 0 ) where L is the likelihoo d function. Iter ation Set t ← t + 1. Sample x ( t,i ) pos ∼ N ( x ( t − 1 ,i ) pos + ∆ x ( t − 1 ,i ) vel , 0 . 02) x ( t,i ) vel ∼ N ( x ( t − 1 ,i ) vel , 0 . 001) y ( t,i ) pos ∼ N ( y ( t − 1 ,i ) pos + ∆ y ( t − 1 ,i ) vel , 0 . 02) y ( t,i ) vel ∼ N ( y ( t − 1 ,i ) vel , 0 . 001) W eight W ( i ) t ∝ W ( i ) t − 1 L ( X ( i ) t | Y t ). R ep e at the Iteration step until al l data ar e pr o c esse d . Algorithm 1: P article filter algorithm for the almost constant velocity mo del. int ma in () { Sampler sampler(ParticleNum); sampler.init(cv_init); sampler.move(cv_move(), false); MonitorEvalAdapter cv_est(cv_monitor_state); sampler.monitor("pos", 2, cv_est); sampler.initialize((voi d *)"pf.d ata"); sampler.iterate(DataNum - 1); std::ofstream est("pf.est"); est << sampler << std::endl; est.close(); est.clear(); return 0; } In the main function, w e constructed a sampler with ParticleNum particles. And then we added the initialization function cv_init and the mov e ob ject of t yp e cv_move . And then w e added a monitor that will record the imp ortance sampling estimates of the t wo p osition parameters. Next, we initialized the sampler with data file pf.data and iterate the sampler un til all data are pro cessed. The last step is that w e output the results into a file called pf.est . The class cv will b e our v alue collection which is a subclass of StateMatrix . T o illustrate b oth the core mo dule and the SMP mo dule, the initialization cv_init will b e implement ed as a standalone function. The mov e cv_move will b e implemented as a deriv ed class of MoveSEQ . T o monitor the imp ortance sampling estimates of the tw o 22 vSMC: Parallel Sequential Monte Carlo in C++ p osition parameters, we will implemen t a simple function cv_monitor_st ate and use the adapter MonitorEvalAdapter . The v alue collection is an N b y Dim (in this case Dim = 4) matrix of type double . W e can simply use StateMatrix as our v alue collection. Ho wev er, w e w ould lik e to enhance its functionalit y through inheritance. First, since the data is shared by all particles, it is natural to bind it with the v alue collection. Second, both the initialization and mo ve will need to calculate the log-lik eliho o d. W e can implemen t it as a standalone function, but since the log-likelihoo d function will need to access the data, it is con venien t to implemen t it as a member function of the v alue collection cv . Here is the full implementation of this simple v alue collection class class cv : public StateMatrix { public : cv (si ze_type N) : StateMatrix (N), x_obs_(DataNum), y_obs_(DataNum) {} double log_likelihood (std::size_t iter, siz e_type id) const { const double scale = 10; const double nu = 10; double llh_x = scale * (sta te(id, 0) - x_obs_[ iter]); double llh_y = scale * (sta te(id, 1) - y_obs_[ iter]); llh_x = std::log(1 + llh_x * llh_x / nu); llh_y = std::log(1 + llh_y * llh_y / nu); return -0.5 * (nu + 1) * (llh_x + llh_y); } void read_data (const char *filename) { std::ifstream data(filename); for (s td::size_t i = 0; i != DataNum; ++i) data >> x_obs_[i] >> y_obs_[i]; data.close(); data.clear(); } private : std::vector x_obs_; std::vector y_obs_; }; The log_likelihood mem b er function accepts the iteration num b er and the particle’s id as Zhou, Y an 23 argumen ts. It returns the log-likelihoo d of the id ’th particle at iteration iter . The read_data mem b er function simply read the data from a file. The initialization is implemented through the cv_init function, std::size_t cv_init (Particle &particle, void *filename); suc h that, it first c hecks if filename is NULL . If it is not, then w e use it to read the data. So the first initialization may lo ok like sampler.initialize((voi d *)filen ame); And after that, if w e w ant to re-initialize the sampler, we can simply call, sampler.initialize(); This will reset the sampler and initialize it again but without reading the data. If the data set is large, rep eated IO can b e v ery exp ensiv e. After reading the data, we will initialize eac h particle’s v alue by Normal random v ariates, and calculate its log-lik eliho o d. The last step is to set the logarithm weigh ts of the particle collection. Since this is not a accept- reject t yp e algorithm, the returned acceptance coun t bares no meaning. Here is the complete implemen tation, std::size_t cv_init (Particle &particle, void *filename) { if (fi lename) particle.value().read_d ata(static_cast(filena me)); const double sd_pos0 = 2; const double sd_vel0 = 1; cxx11::normal_distribut ion norm_pos(0, sd_pos0); cxx11::normal_distribut ion norm_vel(0, sd_vel0); std::vector log_weight(particle.size()); for (P article::size_type i = 0; i != particle.size(); ++i) { particle.value().state( i, 0) = norm_pos(pa rticle.rng(i)); particle.value().state( i, 1) = norm_pos(pa rticle.rng(i)); particle.value().state( i, 2) = norm_vel(pa rticle.rng(i)); particle.value().state( i, 3) = norm_vel(pa rticle.rng(i)); log_weight[i] = particle.value().l og_likelihood(0,i); } particle.weight_set().s et_log_weight(log_weight.b egin()); return 0; } In this example, w e read all data from a single file for simpl icit y . In a realisti c application, the data is often pro cessed online – the filter is applied when new data b ecomes av ailable. In this 24 vSMC: Parallel Sequential Monte Carlo in C++ case, the user can use the optional argumen t of Sampler::initialize to pass necessary information to op en a data connection instead of a file name. In the ab ov e implemen tation we iterated ov er all particles. There are other wa ys to iterate o ver a particle collection. First we can use SinglePartic le ob jects, for (P article::size_type i = 0; i != particle.size(); ++i) { SingleParticle sp(i, &particle ); sp.state(0) = norm_pos(sp.rng()); } Second, vSMC pro vides STL style iterators, Particle::iterator , whose value_type is SingleParticle . Therefore one can write the following lo op for (P article::iterator iter = particle.begin(); iter != particle.end(); ++iter) iter->state(0) = norm_pos(iter->rn g()); Third, if one has a compiler supporting C++11 auto and range-based for , the following is also supp orted, for (a uto &sp : particle) sp.state(0) = norm_pos(sp.rng()); There are little or no p erformance difference among all these forms of lo ops. Ho wev er, one can c ho ose an appropriate form to work with the interfaces of other libraries. F or example, the iterator supp ort allows vSMC to be used with STL and other similar libraries. The up dating metho d cv_move is similar to cv_init . It will up date the particle’s v alue b y adding Normal random v ariates. How ev er, as we see ab ov e, eac h call to cv_init causes a log_weight vector b eing allo cated. Its size do es not change b etw een iterations. So it can b e view ed as some resource of cv_move and it is natural to use a class ob ject to manage it. Here is the implementation of cv_move , class cv_move : public MoveSEQ { public : void pre_processor (std::size_t iter, Partic le &particle) {incw_.resize(particle. size());} std::size_t move_state (std::size_ t iter, SingleParticle sp) { const double sd_pos = std::sqrt(0.02); const double sd_vel = std::sqrt(0.001); const double delta = 0.1; cxx11::normal_distribut ion norm_pos(0, sd_pos); cxx11::normal_distribut ion norm_vel(0, sd_vel); sp.state(0) += norm_pos(sp.rng()) + delta * sp.state(2 ); Zhou, Y an 25 sp.state(1) += norm_pos(sp.rng()) + delta * sp.state(3 ); sp.state(2) += norm_vel(sp.rng()); sp.state(3) += norm_vel(sp.rng()); incw_[sp.id()] = sp.particle().val ue().log_likelihood(iter, sp.id()); return 0; } void post_processor (std::size_t iter, Parti cle &particle) {particle.weight_set(). add_log_weight(&incw_[0]); } private : std::vector incw_; }; First before calling any move_state , the pre_processor will b e called, as describ ed in Sec- tion 4.2 . At this step, we will resize the v ector used for storing incremental w eigh ts. After the first resizing, subsequen t calls to resize will only cause reallo cation if the size changed. In our example, the size of the particle system is fixed, so we don’t need to worry about excessiv e dynamic memory allo cations. The move_state member function mov es each particle accord- ing to our mo del. And after move_state is called for eac h particle, the post_processor will b e called and w e simply add the logarithm incremen tal weigh ts. F or each particle, w e wan t to monitor the x pos and y pos parameters and get the imp ortance sampling esti mates. T o extract the t wo v alues from a particle, w e can implemen t the following function void cv_monitor_state (std::size_t iter, std ::size_t dim, ConstSingleParticle csp, doubl e *res) { assert(dim <= Dim); for (s td::size_t d = 0; d != dim; ++d) res[d] = csp.state(d); } and in the main function we construct a monitor b y MonitorEvalAdapter cv_est(cv_monitor_state); and add it to the sampler through sampler.monitor("pos", 2, cv_est); If later we decided to monitor all states, w e only need to c hange the 2 in the ab ov e line to Dim . After w e implement ed all the ab ov e, compiled and ran the program, a file called pf.est was written by the following statemen t in the main function, 26 vSMC: Parallel Sequential Monte Carlo in C++ ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● 4 6 8 10 12 −6 −5 −4 −3 −2 −1 X position Y position Data ● ● Estimates Observations Figure 1: Observ ations and estimates of a simple particle filter. est << sampler << std::endl; The output file con tains the ESS, resampling, imp ortance sampling estimates and other in- formations in a table. This file can b e read b y most statistical soft wares for further analysis. F or instance, we can pro cess this file with R , and get the plot of the estimates of p ositions against the observ ations, as sho wn in Figure 1 . 5.2. Ba yesian Mo deling of Gaussian mixture mo del The mo del and algorithm Since Ric hardson and Green ( 1997 ), the Gaussian mixture mo del (GMM) has pro vided a canonical example of a mo del-order-determination problem. W e use the same mo del as in Del Moral et al. ( 2006b ) to illustrate the implementation of this classical example in Mont e Carlo literature. This mo del is also used in Zhou, Johansen, and Aston ( 2013 ) for demon- stration of the use of SMC in the context of Ba yesian model comparison, whic h pro vides more details of the following setting. The mo del is as follows; data y = ( y 1 , . . . , y n ) are indep enden tly and iden tically distributed as y i | θ r ∼ r X j =1 ω j N ( µ j , λ − 1 j ) where N ( µ j , λ − 1 j ) denotes the Normal distribution with mean µ j and precision λ j ; θ r = Zhou, Y an 27 F or eac h model M r ∈ M perform the follo wing algorithm. Initialization Set t ← 0. Sample θ ( i,t ) r ∼ π ( θ ( i,t ) r | M r ). W eight W ( i ) 0 ∝ 1. Iter ation Set t ← t + 1. W eight W ( i ) t ∝ W ( i ) t − 1 p ( y | θ ( i,t − 1) r , M r ) α ( t/T r ) − α ([ t − 1] /T r ) . Apply resampling if necessary . Sample θ ( i,t ) r ∼ K t ( ·| θ ( i,t − 1) r ), a π t -in v ariant MCMC k ernel. R ep e at the Iteration step up to t = T r . Algorithm 2: SMC algorithm for Ba yesian mo deling of Gaussian mixture mo del. ( µ 1: r , λ 1: r , ω 1: r ) and r is the n umber of componen ts in each mo del. The parameter space is th us R r × R + r × S r where S r = { ω 1: r : 0 ≤ ω j ≤ 1; P r j =1 ω j = 1 } is the standard ( r − 1)-simplex. The priors whic h are the same for eac h comp onent are taken to b e µ j ∼ N ( ξ , κ − 1 ), λ j ∼ G ( ν, χ ) and ω 1: r ∼ D ( ρ ) where D ( ρ ) is the symmetric Diric hlet distribution with parameter ρ and G ( ν, χ ) is the Gamma distribution with shap e ν and scale χ . The prior parameters are set in the same manner as in Richardson and Green ( 1997 ); also see Zhou et al. ( 2013 ) for details. The data is simul ated from a four comp onents mo del with µ 1:4 = ( − 3 , 0 , 3 , 6), and λ j = 2, ω j = 0 . 25, j = 1 , . . . , 4. Our in terest is to simulate the posterior distribution of mo dels with r comp onents, denoted by M r and obtaining the normalizing constan t for the purp ose of Ba yesian mo del comparison ( Rob ert 2007 , chap. 7). Numerous strategies are p ossible to construct a sequence of distributions for the purp ose of SMC sampling. One option is to use for each mo del M r , r ∈ { 1 , 2 , . . . } , the sequence { π t } T r t =0 , defined by π t ( θ t r ) ∝ π ( θ t r | M r ) p ( y | θ t r , M r ) α ( t/T r ) . (15) where the n umber of distribution, T r , and the annealing sc hedule, α : [0 , 1] → [0 , 1] may be differen t for eac h mo del. This leads to Algorithm 2 . The MCMC kernel K t in Algorithm 2 is constructed as a three-blo cks Metrop olis random w alk, 1. Up date µ 1: r through a Normal random w alk. 2. Up date λ 1: r through a Normal random w alk on logarithm scale, that is, on log λ j , j = 1 , . . . , r . 3. Up date ω 1: r through a Normal random walk on logit scale, that is, on ω j /ω r , j = 1 , . . . , r − 1. The standard direct estimate of the normalizing constants ( Del Moral et al. 2006b ) can b e obtained from the output of this SMC algorithm, ˆ λ T r ,N ds = N X i =1 π ( θ ( i, 0) r | M r ) ν ( θ ( i, 0) 0 ) × T r Y t =1 N X i =1 W ( i ) t − 1 p ( y | θ ( i,t ) r M r ) α ( t/T r ) − α ([ t − 1] /T r ) (16) 28 vSMC: Parallel Sequential Monte Carlo in C++ where W ( i ) t − 1 is the imp ortance w eight of sample θ ( i ) t − 1 . Path sampling for estimation of normalizing c onstants As shown in Zhou et al. ( 2013 ) the estimation of the normalizing constant asso ciated with our sequence of distributions can also b e achiev ed b y a Monte Carlo approximation to the p ath sampling form ulation given by Gelman and Meng ( 1998 ), also known as thermo dynamic in tegration or Ogata’s metho d. Given a parameter α whic h defines a family of distributions, { p α = q α / Z α } α ∈ [0 , 1] that mo ve smo othly from p 0 = q 0 / Z 0 to p 1 = q 1 / Z 1 as α increases from zero to one, one can estimate the logarithm of the ratio of their normalizing constants via a simple integr al relationship, log Z 1 Z 0 = Z 1 0 E α d log q α ( · ) d α d α, (17) where E α denotes exp ectation under p α . The sequence of distributions in the SMC algorithm for this example can b e interpreted as b elonging to such a family of distributions, with α = α ( t/T r ). The SMC sampler pro vides us with a set of weigh ted samples obtained from a s equence of dis- tributions suitable for approximating this integral. At each time t we can obtain an estimate of the exp ectation within the integral via the usual imp ortance sampling estimator, and this in tegral can then b e approx imated via a trap ezoidal integration. In summary , the path sam- pling estimator of the ratio of normalizing constan ts λ T r = log ( Z 1 / Z 0 ) can b e appro ximated b y ˆ λ T r ,N ps = T r X t =1 1 2 ( α t − α t − 1 )( U N t + U N t − 1 ) (18) where U N t = N X i =1 W ( i ) t d log q α ( X ( i ) t ) d α α = α t (19) Implementations In this example w e will implement the following classes. • gmm_param is a class that abstracts the parameters of the mo del, θ r = ( µ 1: r , λ 1: r , ω 1: r ). • gmm_state is the v alue collection class. • gmm_init is a class that implement s op erations used to initialize the sampler. • gmm_move_smc is a class that implemen ts op erations used to up date the weigh ts as w ell as selecting the random w alk proposal scales and distribution parameter α ( t/T r ). • gmm_move_mu , gmm_move_lambda and gmm_move_weight are classes that implement the random walks, each for one of the three blo cks. • gmm_path is a class that implemen ts monitors for the path sampling estimator. This class is similar to the imp ortance sampling monitor in tro duced b efore. It is to b e used with Sampler::pat h_sampling . Its interface requiremen t will b e do cumented later. Zhou, Y an 29 • gmm_alpha_linear and gmm_alpha_prior are classes that implement tw o of the many p ossible annealing schemes, α ( t/T r ) = t/T r (linear) and α ( t/T r ) = ( t/T r ) p , p > 1 (prior). • And last, the main function, which configure, initialize and iterate the samplers. This example is considerably more complicated than the last one. Instead of do cumen ting all the implementatio n details, for man y classes we will only show the interfaces. In most cases, the implement ations are straigh tforward as they are either data member accessors or straigh t translation of mathematical formulations. F or member functions with more complex structures, detailed explanation will b e giv en. Int erested readers can see the source code for more details. Later w e will build b oth sequential and parallelized samplers. A few configuration macros will b e defined at compile time. F or example, the sequential sampler is compiled with the follo wing header and macros, #include #define BASE_SATE StateSEQ #define BASE_INIT InitializeSEQ #define BASE_MOVE MoveSEQ #define BASE_PATH PathEvalSEQ The definitions of these macros will b e change d at compi le time to build parallelized samplers. F or example, when using Op enMP parallelization, the header backend_omp.hpp will be used instead of backend_seq.hpp ; and StateSEQ will b e changed to StateOMP along with similar c hanges to the other macros. In the distributed source co de, this is configured by the CMake build system. Again, we first in tro duce the main function. The required headers are the same as the last particle filter example in addition to the SMP back end headers as describ ed ab ov e. The follo wing v ariables used in the main function will b e set by the user input. int Pa rticleNum; int An nealingScheme; int Pr iorPower; int Co mpNum; std::string DataFile; In the main function, we will create ob jects that set the distribution parameter α ( t/T r ) at eac h iteration according to the user input of AnnealingScheme . Belo w is the main function. Note that some source co de of IO op erations which set the parameters abov e are omitted. int ma in () { gmm_move_smc::alpha_set ter_type alpha_setter; if (An nealingScheme == 1) alpha_setter = gmm_alpha_linear(It erNum); if (An nealingScheme == 2) alpha_setter = gmm_alpha_prior(Ite rNum, Pr iorPower); 30 vSMC: Parallel Sequential Monte Carlo in C++ Sampler sampler(ParticleNum); sampler.particle().valu e().comp_num(CompNum); sampler .init(gmm_init()); .move(gmm_move_smc(alph a_setter), false); .mcmc(gmm_move_mu(), false); .mcmc(gmm_move_lambda() , true); .mcmc(gmm_move_weight() , true); .path_sampling(gmm_path ()); .initialize((void *) DataFile.c_st r()); .iterate(IterNum); double ds = sampl er.particle().value().nc( ).log_zconst(); double ps = sampl er.path().log_zconst(); std::cout << "Standard estimate : " << ds << std::endl; std::cout << "Path sampling estimate : " << ps << std::endl; return 0; } The sampler first sets the num b er of comp onents and allo cate memory through member function comp_num of gmm_state . Then it sets the initialization and up dating metho ds. Before p ossible resampling, a gmm_move_smc ob ject is added. After that, three Metrop olis random walks are app ended. In addition, we add a gmm_path ob ject to calculate the path sampling in tegration. Then we initialize and iterate the sampler and get the normalizing constan t estimates. It is ob vious that the parameter class g mm_param need to store the parameters ( µ 1: r , λ 1: r , ω 1: r ). W e also asso ciate with each particle its log-lik eliho o d and log-prior. Here is the definition of the gmm_param class. W e omitted definitions of some data access member functions. class gmm_param { public : void comp_num (std::size_t num); void save_old (); double log_prior () const {return log_prior_;} double &log_prior () {return log_prior_;} double log_likelihood () const {return log_likelihood_;} double &log_likelihood () {return log_likeli hood_;} int mh _reject_mu (double p, double u); int mh _reject_lambda (double p, double u); int mh _reject_weight (double p, double u); Zhou, Y an 31 int mh _reject_common (double p, double u); double log_lambda_diff () const; double logit_weight_diff () const; void update_log_lambda (); private : std::size_t comp_num_; double log_prior_, log_prior_old_, log_likelihood_, log_likelihood_old_; std::vector mu_, mu_old_; std::vector lambda_, lambd a_old_; std::vector weight_, weigh t_old_; std::vector log_lambda_; }; The comp_num member function allo cate the memory for a giv en num b er of comp onents. The save_old member function sav e the curren t particle states. It is used before the states are up dated with the random walk prop osals, as w e will see later when we implement the gmm_move_mu . The mh_reject_mu member function accept the Metrop olis acceptance prob- abilit y p and a uniform (0 , 1] random v ariate, sa y u ; it rejects the prop osed change if p < u , and restore the particle state of the parameters µ 1: r b y those v alues sav ed by save_old . The mem b er functions mh_reject_lambda and mh_reject_weight do the same for the other t wo set of parameters. All these three also call the mh_reject_common which restore the stored log-lik eliho o d and log-prior v alues. The use of these member functions will be seen in the implemen tation of gmm_move_mu , in the con text of which their own implementation b ecome ob vious. Other member functions provide some useful computations suc h as the logarithm of the λ 1: r . They are used when compute the log-lik eliho o d. The class gmm_state contains some prop erties common to all particles, such as the data and the distribution parameter α ( t/T r ). Also, we will ha v e it to record the logarithm of the ratio of normalizing constants, using the NormalizingConstant class. W e will see ho w to up date this v ariable at each iteration in the implemen tation of gmm_move_smc . The prior parameters are also stored in the v alue collection. Here is the definition of this v alue collection class. Again, we omitted some data access member functions, class gmm_state : public BASE_STATE > { public : NormalizingConstant &nc () {return nc_;} const NormalizingConstant &nc () const {return nc_;} void alpha (double a) { a = a < 1 ? a : 1; a = a > 0 ? a : 0; 32 vSMC: Parallel Sequential Monte Carlo in C++ if (a == 0) { alpha_inc_ = 0; alpha_ = 0; } else { alpha_inc_ = a - alpha_; alpha_ = a; } } void comp_num (std::size_t num) { comp_num_ = num; for (s ize_type i = 0; i != this->size(); ++i) this->state(i, 0).comp_num(num); } double update_log_prior (gmm_param ¶m) const; double update_log_likelihood (gmm_ param &p aram) con st { static const double log2pi = 1.8378770664093455; // log(2pi) double ll = -0.5 * obs_.siz e() * log2pi; param.update_log_lambda (); for (s td::size_t k = 0; k != obs_.size(); ++k) { double lli = 0; for (s td::size_t i = 0; i != param.comp_num(); ++i) { double resid = obs_[k] - pa ram.mu(i); lli += param.weight(i) * std::exp( 0.5 * param.log_lambda(i) - 0.5 * param.lambda(i) * resid * resid); } ll += std::log(lli); } return param.log_likelihood() = ll; } void read_data (const char *filename); private : NormalizingConstant nc_; std::size_t comp_num_; double alpha_, alpha_inc_; double mu0_, sd0_, shape0_, scale0_; double mu_sd_, lambda_sd_, weight_sd_; std::vector obs_; Zhou, Y an 33 }; The v ariable alpha_inc_ is ∆ α ( t/T r ) = α ( t/T r ) − α (( t − 1) /T r ), which will b e used when w e up date the weigh ts. The v ariable nc_ of type NormalizingConstant will b e up dated when the w eigh ts are c hanged by gmm_move_smc and it will compute the standard normalizing constan t estimate ˆ λ T r ,N ds . The v ariables mu0_ and sd0_ are the prior parameters of the means µ 1: r . The v ariables shape0_ and scale0_ are the prior parameters of the precisions λ 1: r . The v ariables mu_sd_ , lambda_sd_ , and weight_sd_ are the prop osal scales of the three random w alks, resp ectivel y . The data access mem b er functions of these v ariables are omitted in the ab o v e source co de snipp et. In the update_log_likelihood mem b er function, the calculation is a straightforw ard trans- lation of the mathematical form ulation. The gmm_param::up date_log_lambda mem b er func- tion is used b efore the lo op, which simply calculates log λ j for j = 1 , . . . , r , and stores their v alues. The purp ose is to av oid rep eated computation of these quan tities inside the lo op. When the function returns, it uses the mutable version of the gmm_param::log _likelihood mem b er function to up date the log-likelihoo d stored in the param ob ject. This is the rea- son that the function is named with a update prefix. As w e will see later, whenever the parameter v alues are up dated, it will b e follo w ed by a call to update_log_likelihood and update_log_prior , which is implemented in a similar fashion. Therefore the v alue w e get b y calling gmm_param::log_likelihood will alw ays b e “up-to-date” while no rep eated com- putation is in volv ed. Surely there are other and p ossibly better design choices. Ho wev er, for this simple example, this design serv es our purp ose w ell. The initialization is implemented using the gmm_init class, class gmm_init : public BASE_INIT { public : std::size_t initialize_state (vsmc ::SingleParticle sp) { const gmm_state &state = sp.particle().value (); gmm_param ¶m = sp.state(0); vsmc::cxx11::normal_dis tribution<> rmu( state.mu0(), state.sd0()); vsmc::cxx11::gamma_dist ribution<> rlambda( state.shape0(), state.scale0()); vsmc::cxx11::gamma_dist ribution<> rweight(1, 1); double sum = 0; for (s td::size_t i = 0; i != param.comp_num(); ++i) { param.mu(i) = rmu(sp.rng()); param.lambda(i) = rlambda(sp.rng() ); param.weight(i) = rweight(sp.rng() ); sum += param.weight(i); } for (s td::size_t i = 0; i != param.comp_num(); ++i) 34 vSMC: Parallel Sequential Monte Carlo in C++ param.weight(i) /= sum; state.update_log_prior( param); state.update_log_likeli hood(param); return 1; } void initialize_param (vsmc::Parti cle &particle, void *filename) { if (fi lename) particle.value().read_d ata(static_cast(filena me)); particle.value().alpha( 0); particle.set_equal_weig ht(); particle.value().nc().i nitialize(); } }; The initializ e_param mem b er function is called b efore the pre_processor , whic h is absent in this case and hav e the default implementation which do es nothing. And it pro cesses the op- tional parameter of Sampler::initialize , the file name of the data. The initialize_state mem b er function initialize the state v alues according to the prior and up date the log-prior and log-likeli ho o d. After initialization, at each iteration, gmm_move_smc class will implement the up dating of w eights as well as selecting of the prop osal scales and the distribution parameter. F or exampl e, when using the linear annealing scheme , w e can implement a gmm_alpha_linear class as the follo wing, class gmm_alpha_linear { public : gmm_alpha_linear (const std::size_ t iter_n um) : iter_num_(iter_num) {} void operator() (std::size_t iter, Particle< gmm_state> &particle) {particle.value().alpha (static_cast(iter) / iter_nu m_);} private : std::size_t iter_num_; }; It accepts the total num b er of iterations T r as an argument to its constructor. And it imple- men ts an operator() that up date the distribution parameter α ( t/T r ). The prior annealing sc heme can b e implemen ted similarly . F or simplicity and demonstration purpose, we only allo w gmm_move_smc to b e configured with differen t annealing schemes, and hard co de the Zhou, Y an 35 prop osal scales. An industry strength design ma y mak e this class a template with annealing sc heme and prop osal scales as p olicy template parameters. class gmm_move_smc { public : typedef cxx11::function &)> alpha_setter_type; gmm_move_smc (const alpha_setter_t ype &alp ha_setter) : alpha_setter_(alpha_set ter) {} std::size_t operator() (std::size_ t iter, Particle &particle) { alpha_setter_(iter, particle); double alpha = particle.value().alpha(); alpha = alpha < 0.02 ? 0.02 : alpha; particle.value().mu_sd( 0.15 / alpha); particle.value().lambda _sd((1 + std::sqrt(1 / alpha)) * 0.15); particle.value().weight _sd((1 + std::sqrt(1 / alpha)) * 0.2); incw_.resize(particle.s ize()); weight_.resize(particle .size()); particle.read_weight(we ight_.begin()); double coeff = particle.value().alpha_inc(); for (v smc::Particle:: size_type i = 0; i != particle.size(); ++i) { incw_[i] = coeff * particle.value().state(i, 0).log_likelihood(); } particle.value().nc().a dd_log_weight(&incw_[0], particle.weight_set()); particle.weight_set().a dd_log_weight(&incw_[0]); return 0; } private : alpha_setter_type alpha_setter_; std::vector incw_; std::vector weight_; }; Note that, cxx11::function is an alias to either std::function or boost::function , de- 36 vSMC: Parallel Sequential Monte Carlo in C++ p ending on the v alue of the configuration macro VSMC_HAS_CXX11LIB_FUNCTIONAL . Ob jects of this class t yp e can b e added to a sampler as a mo v e. The operator() satisfies the in terface requiremen t of the core mo dule. First it uses alpha_setter_ to set the dist ribution parameter α ( t/T r ). Second, it sets the prop osal scales for the three Metrop olis random walks according to the current v alue of α . Then it computes the unnormalize d incremen tal weigh ts. The NormalizingConstnat class has member function add_log_weight , whic h is not unlik e the one with the same name in WeightSet . It accepts the logarithm of the incremen tal weigh ts and a WeightSet ob ject. The standard normalizing constant estimates will be computed using these v alues. The last, w e also modify the WeightSet t yp e ob ject itself b y adding the logarithm of the incremen tal weigh ts. A t each iteration, ramdom walks are also p erfo emd. The implemen tations of the random w alks are straigh tforward. Belo w is the implementation of the random w alk on the mean parameters. The random walks on the other parameters are similar. class gmm_move_mu : public BASE_MOVE { public : std::size_t move_state (std::size_ t iter, SingleParticle sp) { const gmm_state &state = sp.particle().value (); gmm_param ¶m = sp.state(0); cxx11::normal_distribut ion<> rmu (0, state .mu_sd()); cxx11::uniform_real_dis tribution<> runif(0, 1); double p = param.log_prior() + state.alpha() * param.lo g_likelihood(); param.save_old(); for (s td::size_t i = 0; i != param.comp_num(); ++i) param.mu(i) += rmu(sp.rng()); p = state.update_log_prior(param) + state.alpha() * state.update_log_l ikelihood(param) - p; double u = std::l og(runif(sp.rng())); return param.mh_reject_mu(p, u); } }; First we sav e the logarithm of the v alue of target density computed using the old v alues in p , the acceptance probability . And then we call gmm_param::save_old to sav e the old v alues. Next we update each parameter with a prop osed Normal random v ariates and compute the new log-prior and the log-likelihoo d as w ell as the new v alue of the target densit y . Then w e reject it according to the Metrop olis algorithm, as implemen ted in gmm_param , whic h manages b oth the current states as well as the backup. Last we need to monitor certain quantities for interferen ce purp ose. Recall that, in the main function we used sampler.path_sampling(gmm_path()) to set the monitoring of path Zhou, Y an 37 sampling integrands. The path_sampling mem b er function requires a callable ob jects with the following signature, double path_eval (std::size_t iter, const Particle &, double *res); The input parameter iter is the iteration num b er, the v alue of t in Equation 18 . The return v alue shall b e the v alue of α t . The output parameter res shall store the array of v alues of d log q α ( X ( i ) t ) d α α = α t . Our implementation of gmm_path is a sub-class of an SMP mo dule base class, which pro vides an operator() that satisfies the ab ov e interfac e requirement. Its usage is similar to the MoveSEQ template introduced in Section 4.2 . The path sampling in tegrands under this geometry annealing scheme are simply the log- lik eliho o d. Therefore the implementation of gmm_path class is rather simple, class gmm_path : public BASE_PATH { public : double path_state (std::size_t, ConstSingleParticle sp) {return sp.state(0).log_likelihood();} double path_grid (std::size_t, const Particl e &particle) {return particle.value().alpha();} }; R esults After compiling and running the algorithm, the results were consisten t with those rep orted in Del Moral et al. ( 2006b ). F or a more in depth analyze of the metho dologies, extensions and the results see Zhou et al. ( 2013 ). Extending the implementation using MPI vSMC ’s MPI mo dule assumes that iden tical samplers are constructed on each node, with p ossible different num b er of particles to accommo date the difference in capacities among no des. T o extend the ab ov e SMP implemen tation for use with MPI , first at the b eginning of the main function, we add the follo wing, MPIEnvironment env(argc, argv); to initialize the MPI en vironment. When the ob ject env is destro y ed at the exit of the main function, the MPI environmen t is finalized. Second, we need to replace base v alue collection class template with StateMPI . So no w gmm_state is declared as the follo wing, class gmm_state : public StateMPI > >; 38 vSMC: Parallel Sequential Monte Carlo in C++ The implement ation is exactly the same as b efore. Third, the gmm_param class now needs to b e transferable using MPI . Unlik e the SMP situations, a simple copy constructor is not enough. vSMC uses Bo ost MPI library , and t h us one only needs to write a serialize member function for gmm_param such that the data can be serialized in to bytes. See do cument s of Boost MPI and serialization libraries for details. In summary , the following member function accept an Archive ob ject as input, and it can p erform a store or a load op eration based on the Archive t yp e. In a load op eration, the Archive ob ject is like an input stream and in a store operation, it is like an output stream. template void serialize (Archive &ar, const unsigned) { int nu m = comp_num_; ar & num; comp_num(num); ar & log_prior_; ar & log_likelihood_; for (s td::size_t i = 0; i != comp_num_; ++i) { ar & mu_[i]; ar & lambda_[i]; ar & weight_[i]; ar & log_lambda_[i]; } } F ourth, after user input of the sampler parameters, w e need to sync them with all no des. F or example, for the ParticleNum parameter, boost::mpi::communicato r World; boost::mpi::broadcast(W orld, Par ticleNum, 0); Last, any imp ortance sampling estimates that are computed on each no de, need to be com- bined into final results. F or example, the path sampling results are now obtained through adding the results from each no de together, double ps_sum = 0; boost::mpi::reduce(Worl d, ps, ps_sum, std: :plus(), 0); ps = ps_sum; F or the standard normalizing constan t ratio estimator, w e will replace NormalizingConstant with NormalizingConstantMPI , which will p erform such and other tasks. After these few lines of c hange, the sampler is now p arallelized using MPI and can b e deplo yed to clusters and other distributed memory architecture. On eac h no de, the selected SMP parallelization is used to perform multi- threading parallelization locally . vSMC ’s MPI module will take care of normalizing w eights and other tasks. Zhou, Y an 39 Par al lelization p erformanc e One of the main motiv ation b ehind the creation of vSMC is to ease the parallelization with differen t programming mo dels. The same implementation can b e used to built differen t sam- plers based on what kind of parallel programming mo del is supp orted on the users’ platforms. In this section w e compare the performance of v arious SMP parallel programmin g mo dels and Op enCL parallelization. W e consider five differen t implemen tations supp orted by In tel C++ Complier 2013: sequen tial, In tel TBB , Cilk Plus , Op enMP and C++11 . The samplers are compiled with CXX=icpc -std=c++11 -gcc-name=gcc- 4.7 -gxx -name=g++-4.7 CXXFLAGS=-O3 -xHost -fp-model precise \ -DVSMC_HAS_CXX11LIB_FUN CTIONAL=1 \ -DVSMC_HAS_CXX11LIB_RAN DOM=1 on a Ubun tu 12.10 workstation with an Xeon W3550 (3.06GHz, 4 cores, 8 hardware threads through hyper-threading) CPU. A four components mo del and 100 iterations with a prior annealing scheme is used for all implementations. A range of num b ers of particles are tested, from 2 3 to 2 17 . F or differen t n umber of particles, the wall clo ck time and speedup are shown in Figure 2 . F or 10 4 or more particles, the differences are minimal among all the programming mo dels. They all ha ve roughly 550% sp eedup. With smaller n umber of particles, vSMC ’s C++11 parallelization is less efficient than other industry strength programming mo dels. Ho wev er, with 1000 or more particles, which is less than t ypical applications, the difference is not very significan t. Op enCL implementations are also compared on the same workstation, which also has an NVIDIA Quadro 2000 graphic card. OpenCL programs can b e compiled to run on b oth CPUs and GPUs. F or CPU implementation, there are Intel Op enCL ( Intel Co op eration 2013b ) and AMD APP Op enCL ( Adv anced Micro Devices, Inc. 2012 ) platforms. W e use the Intel TBB implemen tation as a baseline for comparison. The same Op enCL implemen tation are used for all the CPU and GPU runtimes. Therefore they are not particularly optimized for an y of them. F or the GPU implemen tation, in addition to double precision, we also tested a single precision configuration. Unlike modern CPUs, whic h ha ve the same p erformance for double and single precision floating p oint op erations (unless SIMD instructions are used, whic h can ha ve at most a sp eedup b y a factor of 2), GPUs penalize double precision performance hea vily . F or different n umber of particles, the wall clo c k time and sp eed up are plotted in Figure 3 . With smaller n um b er of particles, the Op enCL implementati ons hav e a high ov erhead when compared to the Intel TBB implementation. With a large num b er of particles, AMD APP Op enCL has a similar p erformance as the In tel TBB implementation. Intel Op enCL is about 40% faster than the In tel TBB implementation. This is due to more efficien t v ectorization and compiler optimizations. The double precision p erformance of the NVIDIA GPU has a 220% sp eedup and the single precision p erformance has near 1600% sp eedup. As a rough reference for the exp ected p erformance gain, the CPU has a theoretical p eak performance of 24.48 GFLOPS. The GPU has a theoretical p eak p erformance of 60 GFLOPS in double precision and 480 GFLOPS in single precision. This represen ts 245% and 1960% speedup compared to the CPU, resp ectiv ely . 40 vSMC: Parallel Sequential Monte Carlo in C++ ● ● ● ● ● ● ● ● ● ● ● ● 10 2 10 3 10 4 10 5 10 2 10 3 10 4 10 5 Number of par ticles W all clock time (second) Implementation ● Sequential Intel TBB Intel Cilk Plus OpenMP vSMC STDTBB ● ● ● ● ● ● ● ● ● ● ● ● 2 0 2 1 2 2 10 2 10 3 10 4 10 5 Number of par ticles Speedup Implementation ● Sequential Intel TBB Intel Cilk Plus OpenMP vSMC STDTBB Figure 2: Performance of C++ implemen tations of Bay esian mo deling for Gaussian mixture mo del (Linux; Xeon W3550, 3.06GHz, 4 cores, 8 threads). Zhou, Y an 41 ● ● ● ● ● ● ● ● ● ● ● ● 10 −1 10 0 10 1 10 2 10 2 10 3 10 4 10 5 Number of par ticles W all clock time (second) Implementation ● Intel TBB AMD CPU Intel CPU NVIDIA GPU (single) NVIDIA GPU (double) ● ● ● ● ● ● ● ● ● ● ● ● 2 −2 2 −1 2 0 2 1 2 2 2 3 2 4 10 2 10 3 10 4 10 5 Number of par ticles Speedup Implementation ● Intel TBB AMD CPU Intel CPU NVIDIA GPU (single) NVIDIA GPU (double) Figure 3: P erformance of Op enCL implemen tations of Bay esian mo deling for Gaussian mixture mo del (Linux; Xeon W3550 GPU, 3.06GHz, 4 cores, 8 threads; NVIDIA Quadro 2000). 42 vSMC: Parallel Sequential Monte Carlo in C++ It is widely believed that Op enCL programming is tedious and hard. Ho w ever, vSMC provides facilities to manage Op enCL platforms and devices as well as common op erations. Limited b y the scope of this pap er, the OpenCL implemen tation (distributed with the vSMC source) is not do cumented in this pap er. Ov erall the Op enCL implementat ion has ab out 800 lines including b oth host and device co de. It is not an enormous increase in effort when compared to the 500 lines SMP implementation. Less than doubling the co de base but gaining more than 15 times performance speedup, w e consider the programming effort is relativ ely small. 6. Discussion This pap er in tro duced a C++ template library in tended for implementing general SMC al- gorithms and constructing parallel samplers with differen t programming models. While it is p ossible to implemen t man y realistic application with the present ed framew ork, some tech- nical proficiency is still required to implement some problem specific part of the algorithms. Some basic knowledge of C++ in general and how to use a template library are also required. It is shown that with the presen ted framew ork it is possible to implement parallelized, scalable SMC samplers in an efficient and reusable wa y . The p erformance of some common parallel programming mo dels are compared using an example. Some future w ork ma y w orth the effort to ease the implemen tation of SMC algorithms further. Ho wev er, there is a balance b etw een p erformance, flexibilit y and the ease of use. vSMC aims to b e developer-friendly and to provide users as muc h control as possible for all p erformance related asp ects. F or a BUGS -like interface, users may b e interest ed in other softw are such as BiiPS ( Caron, T o deschini, Legrand, and Moral 2012 ). In addition LibBi ( Murra y 2013 ) pro vides a user friendly and high p erformance alternative with a fo cus on state-space mo dels. Compared with these recent dev elopments, vSMC is less accessible to those with little or no kno wledge of C++ . How ev er, for researchers with exp ertise in C++ and template generic programming in particular, vSMC provides a framework within whic h p oten tial sup erior p erformance can b e obtained and greater flexibilit y and extensibilit y are p ossible. References Adv anced Micro Devices, Inc (2012). AMD A c c eler ate d Par al lel Pr o c essing Op enCL Pr o- gr amming Guide . V ersion 2.8, URL http://developer.amd.com/tools- and- sdks/ heterogeneous- computing/amd- accelerated- parallel- processing- app- sdk . Capp ´ e O, Go dsill SJ, Moulines E (2007). “An Overview of Existing Metho ds and Recent Adv ances in Sequen tial Mon te Carlo.” Pr o c e e dings of the IEEE , 95 (5), 899–924. Caron F, T o deschin i A, Legrand P , Moral PD (2012). BiiPS : Bayesian Infer enc e with Inter- acting Particle Systems . V ersion 0.7.2, URL https://alea.bordeaux.inria.fr/ biips/ doku.php . Da wes B, et al. (2013). Bo ost – C++ Libr aries . V ersion 1.53, URL http://www.boost.org/ . Del Moral P , Doucet A, Jasra A (2006a). “Sequential Monte Carlo Metho ds for Bay esian Computation.” In Bayesian Statistics 8 . Oxford Universit y Press. Zhou, Y an 43 Del Moral P , Doucet A, Jasra A (2006b). “Sequential Monte Carlo Samplers.” Journal of R oyal Statistic al So ciety B , 68 (3), 411–436. Douc R, Capp ´ e O, Moulines E (2005). “Comparison of Resampling Sc hemes for P article Filter- ing.” In Pr o c e e dings of the 4th International Symp osium on Imange and Signal Pr o c essing and A nalysis , pp. 1–6. Doucet A, Johansen AM (2011). “A T utorial on Particle Filtering and Smo othing: Fifteen Y ears Later.” In The Oxfor d Handb o ok of Non-line ar Filtering . Oxford Universit y Press. Gelman A, Meng XL (1998). “Sim ulating Normalizi ng Constan ts: F rom Importance Sampling to Bridge Sampling to P ath Sampling.” Statistic al Scienc e , 13 (2), 163–185. GNU Pro ject (2013). GCC – GNU Compiler Col le ction . V ersion 4.8.1, URL http://gcc. gnu.org . In tel Co op eration (2011). Intel Cilk Plus L anguage Sp e cific ation . V ersion 1.1, URL http: //cilkplus.org/ . In tel Co op eration (2013a). Intel C++ Comp oser XE . V ersion 13.1, URL http://software. intel.com/en- us/intel- compilers . In tel Co op eration (2013b). Intel SDK for Op enCL Applic ations 2013 . URL http://software. intel.com/en- us/vcsource/tools/opencl- sdk . In tel Co op eration (2013c). Intel Thr e ading Building Blo cks . V ersion 4.1, URL http: //threadingbuildingbloc ks.org/ . Johansen AM (2009). “SMCTC: Sequential Monte Carlo in C++ .” Journal of Statistic al Softwar e , 30 (6), 1–41. Kronos Op enCL W orking Group (2012). The Op enCL Sp e cific ation . V ersion 1.2, URL http: //www.khronos.org/openc l/ . Lee A, Y au C, Giles MB, Doucet A, Holmes CC (2010). “On the Utilit y of Graphics Cards to P erform Massiv ely P arallel Simulation of Adv anced Monte Carlo Methods.” Journal of Computational and Gr aphic al Statistics , 19 (4), 769–789. Liu JS, Chen R (1998). “Sequen tial Monte Carlo Metho ds for Dynamic Systems.” Journal of the A meric an Statistic al Asso ciation , 93 (443), 1032–1044. Martin K, Hoffman B (2010). Mastering CMake . V ersion 2.8, URL http://www.cmake.org . Message Passing Inter face F orum (2012). MPI : A Message-Passing Interfac e Starndar d . V er- sion 3.0, URL http://www.mpi- forum.org/ . Microsoft Co op eration (2012). Micr osoft Visual C++ 2012 . URL http://msdn.microsoft. com/en- us/vstudio/hh386302 . Murra y LM (2013). “Bay esain State-Space Mo delling on High-P erformance Hardware Using LibBi .” arXiv , pp. 1–28. URL http://arxiv.org/abs/1306. 3277 . Neal RM (2001). “Annealed Imp ortance Sampling.” Statistics and Computing , 11 (2), 125–139. 44 vSMC: Parallel Sequential Monte Carlo in C++ Op enMP Architecture Review Board (2011). Op enMP Applic ation Pr o gr am Interfac e . V er- sion 3.1, URL http://www.openmp. org . P eters GW (2005). T opics in Se quential Monte Carlo Samplers . Master’s thesis, Department of Engineering, Universit y of Cam bridge. R Core T eam (2013). R : A L anguage and Envir onment for Statistic al Computing . R F oundation for Statistical Computing, Vienna, Austria. V ersion 2.15.2, URL http: //www.r- project.org . Ric hardson S, Green PJ (1997). “On Bay esian An alysis of Mixtures with an Un kno wn Number of Comp onents.” Journal of R oyal Statistic al So ciety B , 59 (4), 731–792. Rob ert CP (2007). The Bayesian Choic e: F r om De cision-the or etic F oundations to Computa- tional Implementation . 2nd edition. Springer-V erlag, New Y ork. Salmon JK, Moraes MA, Dror RO , Shaw DE (2011). Par al lel R andom Numb ers: As Easy as 1, 2, 3 . Spiegelhalter D, Thomas A, Best N (1996). BUGS – Bayesian Infer enc e Using Gibbs Sampling . V ersion 0.5, URL http://www.mrc- bsu.cam.ac.uk/bugs/ . The LL VM Developer Group (2013a). clang : A C L anguage F amily F r ontend for LL VM . V ersion 3.2, URL http://clang.llvm.org/ . The LL VM Developer Group (2013b). “ lib c++ ” C++ Standar d Libr ary . V ersion 1.1, URL http://libcxx.llvm.org . T orv alds L, et al. (2013). Git – Distribute d Sour c e Co de Management . V ersion 1.8.3.1, URL http://git- scm . v an Heesch D (2013). Doxygen – Gener ating Do cumentation fr om Sour c e Co de . V ersion 1.8.4, URL http://www.stack.nl/~dimitri/doxyg en/index.html . Zhou Y, Johansen AM, Aston J AD (2013). “T o wards Automatic Model Comparison: Adaptive Sequen tial Monte Carlo Approac h.” arXiv , pp. 1–33. URL 3123 . Affiliation: Y an Zhou Departmen t of Statistics Univ ersity of W arwic k Co ven try , CV4 7AL, United Kingdom E-mail: Yan.Zhou@warwick. ac.uk URL: http://zhouyan.github.com
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment