FindZebra: A search engine for rare diseases
Background: The web has become a primary information resource about illnesses and treatments for both medical and non-medical users. Standard web search is by far the most common interface for such information. It is therefore of interest to find out how well web search engines work for diagnostic queries and what factors contribute to successes and failures. Among diseases, rare (or orphan) diseases represent an especially challenging and thus interesting class to diagnose as each is rare, diverse in symptoms and usually has scattered resources associated with it. Methods: We use an evaluation approach for web search engines for rare disease diagnosis which includes 56 real life diagnostic cases, state-of-the-art evaluation measures, and curated information resources. In addition, we introduce FindZebra, a specialized (vertical) rare disease search engine. FindZebra is powered by open source search technology and uses curated freely available online medical information. Results: FindZebra outperforms Google Search in both default setup and customised to the resources used by FindZebra. We extend FindZebra with specialized functionalities exploiting medical ontological information and UMLS medical concepts to demonstrate different ways of displaying the retrieved results to medical experts. Conclusions: Our results indicate that a specialized search engine can improve the diagnostic quality without compromising the ease of use of the currently widely popular web search engines. The proposed evaluation approach can be valuable for future development and benchmarking. The FindZebra search engine is available at http://www.findzebra.com/.
💡 Research Summary
The paper presents FindZebra, a vertical search engine specifically designed to aid the diagnosis of rare (or orphan) diseases, and evaluates its performance against general-purpose web search engines (Google Search and Google Custom Search) as well as PubMed. The authors begin by highlighting the growing reliance of both laypeople and clinicians on web-based information for medical queries, noting that standard search engines are optimized for short, keyword‑centric queries and heavily rely on link‑based ranking signals such as PageRank. These characteristics make them ill‑suited for the long, symptom‑list queries typical of rare‑disease diagnostics, where terminology is often highly specialized, multi‑word, and sparsely represented on the web.
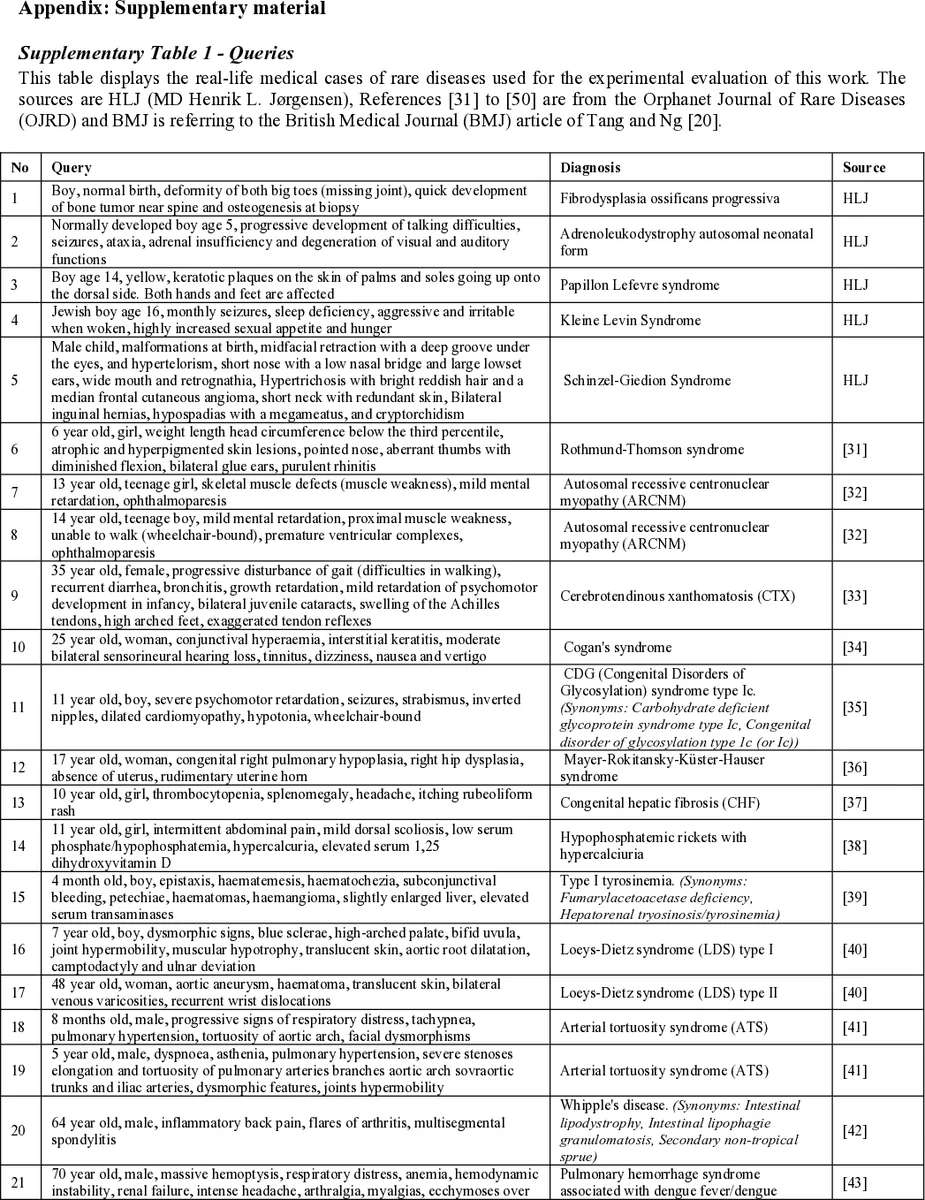
To address this gap, the authors construct an evaluation framework using 56 real‑world diagnostic cases. Each case is expressed as a list of patient symptoms, forming a complex query. Standard information‑retrieval metrics—precision at k (P@k), mean average precision (MAP), normalized discounted cumulative gain (nDCG), and recall—are employed to quantify retrieval quality. The evaluation is designed to isolate two factors: the quality of the underlying index (the document collection) and the effectiveness of the retrieval algorithm.
FindZebra’s architecture consists of three main components. First, a curated corpus is assembled by crawling authoritative, freely available resources such as Orphanet, OMIM, and POSSUMweb. Duplicate removal, metadata normalization, and language preprocessing are applied to produce a clean dataset. Second, the indexing layer uses Apache Lucene to build an inverted index, but crucially augments term matching with medical ontologies: the Unified Medical Language System (UMLS) provides synonym and hierarchical relationships, while the Human Phenotype Ontology (HPO) supplies phenotype‑disease mappings. This enables query expansion that respects medical semantics, allowing “sleep deficiency” and “insomnia” to be treated as related concepts. Third, the search interface presents results with enriched context—ICD‑10 codes, links to relevant literature, and brief snippets—so clinicians can quickly assess relevance.
In the experimental phase, the same curated dataset is fed into Google Custom Search Engine (CSE) to evaluate whether the Google ranking algorithm itself is a limiting factor when the index is controlled. Results show that while Google CSE improves over default Google Search by focusing on the same documents, it still underperforms FindZebra because the underlying ranking model is tuned for general web relevance rather than medical specificity. PubMed, which indexes biomedical literature, performs poorly on symptom‑list queries because it defaults to Boolean “AND” matching, discarding many potentially relevant articles that lack every term.
FindZebra consistently outperforms the baselines. In the top‑10 results, it achieves an average precision exceeding 70 %, substantially higher than Google Search (≈30 %) and Google CSE (≈45 %). The advantage is most pronounced for cases with rare or highly technical symptoms, where the specialized index and ontology‑driven expansion mitigate term‑independence assumptions and low‑frequency term pruning that cripple generic engines. The authors also demonstrate a prototype UI that visualizes UMLS concepts and HPO hierarchies alongside search hits, further supporting clinicians’ hypothesis‑driven reasoning.
The discussion acknowledges limitations: the current corpus excludes proprietary clinical databases and may lag behind the latest research; query expansion can occasionally introduce noise if overly aggressive; and the evaluation, while realistic, is limited to 56 cases. Future work includes integrating machine‑learning‑based relevance feedback, expanding the corpus with newer datasets, and conducting user studies in real clinical settings.
In conclusion, the study provides strong evidence that a domain‑specific search engine, built on a curated medical corpus and enriched with biomedical ontologies, can substantially improve diagnostic search performance for rare diseases without sacrificing the simplicity and speed that make general web search attractive to clinicians. The proposed evaluation methodology also offers a reproducible benchmark for future medical IR systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment