On the Approximability of Comparing Genomes with Duplicates
A central problem in comparative genomics consists in computing a (dis-)similarity measure between two genomes, e.g. in order to construct a phylogeny. All the existing measures are defined on genomes without duplicates. However, we know that genes c…
Authors: Sebastien Angibaud (LINA), Guillaume Fertin (LINA), Irena Rusu (LINA)
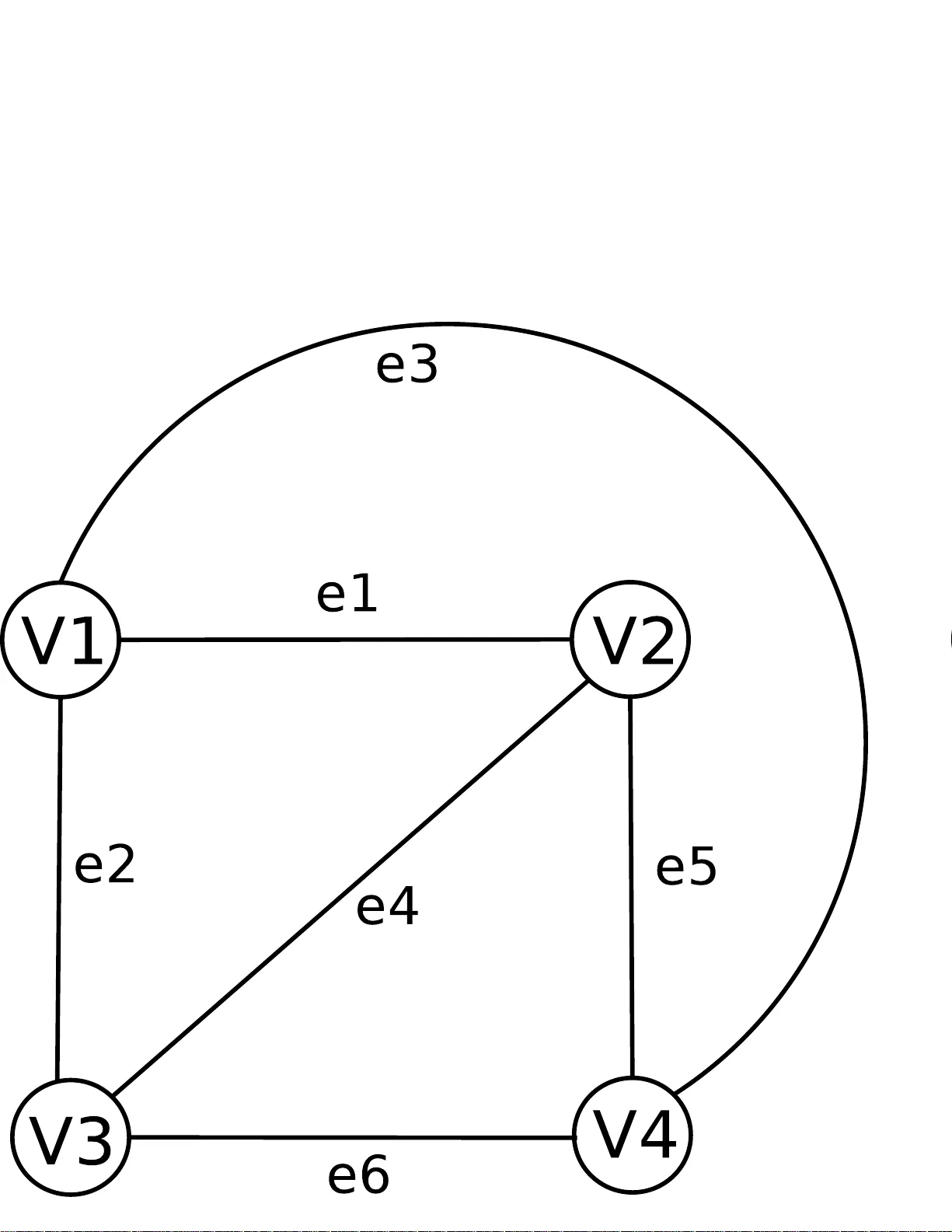
On the Appro ximabilit y of Comparing Genomes with Duplicates S ´ ebastien Angibaud 1 , G uillaume F ertin 1 , Irena Rusu 1 , Ann elyse T h ´ ev en in 2 , and St ´ ephane Vialette 3 1 Lab oratoire d’Informatique de Nantes-A tlan tique (LINA), UMR CNRS 6241, Universit ´ e de Nantes, 2 rue de la Houssini ` ere, 44322 Nantes Cedex 3, F rance { sebastie n.angibaud,g uillaume.fertin,irena.rusu } @univ-nantes.fr 2 Lab oratoire de Recherc he en Informatique (LR I), UMR CNRS 8623, Universit ´ e P aris-Sud, 91405 Orsay , F rance thevenin@l ri.fr 3 IGM-LabInfo, UMR CNRS 8049, U niversi t´ e P aris-Est, 5 Bd Descartes 7745 4 Marne-la-V all ´ ee, F rance vialette@u niv-mlv.fr Abstract. A central problem in comparativ e genomics consists in compu t ing a (dis-)similarit y measure b etw een tw o genomes, e.g. in order to construct a ph ylogenetic tree. A large num b er of such measures has been prop osed in the recent past: numb er of r eversals , numb er of br e akp oi nts , numb er of c ommon or c onserve d intervals , SAD etc. In their initial definitions, all these measures supp ose that genomes conta in n o dup licates. How ever, w e now know that genes can b e d uplicated within the same genome. O ne p ossible approach to ove rcome this difficulty is to establish a one-to-one correspondence (i.e. a matc hing) b etw een genes of b oth genomes, where th e correspondence is chosen in order to optimize the studied measure. Then, after a gene relabeling according to th is matching and a deletion of the u nmatched signed genes, tw o genomes without duplicates are obtained and the measure can b e comp u ted. In this pap er, we are in terested in three measures ( numb er of br e akp oints , numb er of c ommon intervals and numb er of c onserve d intervals ) and three mo dels of matching ( exemplar , interme diate and maximum matching mo dels). W e prove that, for each model and eac h measure M , computing a matching b etw een tw o genomes that optimizes M is APX –hard. W e show that this result remains true even for tw o genomes G 1 and G 2 such that G 1 conta ins n o duplicates and no gene of G 2 app ears more t h an twice. Therefore, our results extend those of [7, 1 0, 13]. Besides, in order to ev aluate the p ossible existence of approximatio n algorithms concerning the num b er of breakp oints, we also stud y the complexity of the follo wing decision problem: is there an exemp larization (resp. an intermediate matc hing, a maximum matc hing) that induces no breakp oint ? I n p articular, w e extend a result of [13] by proving the problem to b e NP –complete in the exemp lar model for a n ew class of instances, w e note th at the problems are eq uiv alent in the intermediate and the ex emplar mo dels and we show that the p roblem is in P in the max imum matching mod el. Finally , we focu s on a fourth measure, closely related to the num b e r of breakp oints: the numb er of adjac encies , for which we giv e sev eral constant ratio approximatio n algorithms in th e maxim um matching model, in the case where genomes contai n the same number of duplications of each gene. Keywords : genome rearrangemen ts, APX –hardness, duplicate genes, breakp oints, adjacencies, com- mon interv als, conserv ed interv als, approximation alg orithms. 1 In tro duction and P reliminaries In comparativ e genomics, computing a measure of (dis-)similarit y b et w een t w o genomes is a cen tral problem: suc h a measure can b e used, for ins tance, to construct phylo genetic trees. Th e measures defined so f ar essent ially fall in to tw o categories: the first one consists in coun ting the minim um n umber of op erations needed to transform a genome into another (e.g. the e dit distanc e [21] or the numb er of r eversals [4]) . The second one contai ns (dis-)similarit y measures based on the genome structure, suc h as the numb er of br e akp oints [7], the c onserve d intervals distanc e [6], th e numb er of c ommon intervals [10], SA D and MAD [24] etc. When genomes co nt ain no du plicates, most m easures can b e computed in p olynomial time. Ho wev er, assum in g that genomes con tain no d u plicates is too limited. Ind eed, it has b een r ecently sho wn that a great num b er of d u plicates exists in some genomes. F or example, in [20], authors estimate that 15% of genes are du plicated in the human genome. A p ossible approac h to o v ercome this d ifficult y is to sp ecify a on e-to-one corresp o ndence (i.e. a matching ) b et w een genes of b oth genomes and to remo v e the unm atc hed genes, thus ob taining t w o genomes with identic al gene con tent and n o du plicates. Usually , the ab ov e mentioned matc hing is c hosen in order to optimize the studied measur e, follo wing the parsimon y principle. Th ree mo dels ac hieving th is corresp ondence ha v e b een prop o sed : the exemplar mo del [23], the interme diate model [3] and the maximum matching mo del [25]. Before definin g precisely the measures and mo dels studied in this pap er, w e need to in tro du ce some n otations. Notations use d in the p ap er. A genome G is represent ed by a sequence of s igned inte gers (called signe d gene s ). F or an y genome G , we denote by F G the set of unsigned in tegers (called genes ) that are present in G . F or an y signed gene g , let − g b e the signed gene ha ving the opp osite s ign and let | g | ∈ F G b e the corresp on d ing (un signed) gene. Giv en a genome G without d u plicates and t w o signed genes a , b suc h that a is lo cated b efore b , let G [ a, b ] b e the set S ⊆ F G of genes lo c ated b et w een genes a and b in G , a and b included. W e also n ote [ a, b ] G the substring (i.e. the sequence of consecutive elements) of G starting at a and finishing at b in G . Let o cc( g , G ) b e the n um b er of o c currences of a giv en gene g in a genome G and let o cc( G ) = max { occ ( g , G ) | g ∈ F G } . A pair of genomes ( G 1 , G 2 ) is said to b e of typ e ( x, y ) if o cc( G 1 ) = x and o cc( G 2 ) = y . A pair of genomes ( G 1 , G 2 ) is said to b e b alanc e d if, for eac h gene g ∈ F G 1 ∪ F G 2 , we ha v e o cc( g , G 1 ) = o cc( g , G 2 ) (otherwise, ( G 1 , G 2 ) will b e said to b e u nb alanc e d ). Note that a pair ( G 1 , G 2 ) of t yp e ( x, x ) is not n ecessary balanced. Denote b y n G the size of genome G , that is the num b er of signed genes it cont ains. Let G [ p ], 1 ≤ p ≤ n G , b e the s igned gene that occurs at p osition p on genome G , and let | G [ p ] | ∈ F G b e the corresp ondin g (unsigned) gene. Let N G [ p ], 1 ≤ p ≤ n G , b e the n umber of o ccurren ces of | G [ p ] | in the first ( p − 1) p ositions of G . W e define a duo in a genome G as a pair of successive signed genes.Giv en a duo d i = ( G [ i ] , G [ i + 1]) in a genome G , w e note − d i the duo equal to ( − G [ i + 1] , − G [ i ]). L et ( d 1 , d 2 ) b e a pair of duos ; ( d 1 , d 2 ) is called a duo match if d 1 is a duo of G 1 , d 2 is a duo of G 2 , and if either d 1 = d 2 or d 1 = − d 2 . F or example, consider th e genome G 1 = +1 + 2 + 3 + 4 + 5 − 1 − 2 + 6 − 2. Then, F G = { 1 , 2 , 3 , 4 , 5 , 6 } , n G 1 = 9, occ (1 , G 1 ) = 2, o cc( G 1 ) = 3, G 1 [7] = − 2, − G 1 [7] = +2, | G 1 [7] | = 2 and N G 1 [7] = 1. Let G 2 b e the genome G 2 = +2 − 1 + 6 + 3 − 5 − 4 + 2 − 1 − 2. Then the pair ( G 1 , G 2 ) is balanced and is of t yp e (3 , 3). Let d 1 = ( G 1 [4] , G 1 [5]) b e the duo (+ 4 , +5) and d 2 b e the duo ( G 2 [5] , G 2 [6]). Th e pair ( d 1 , d 2 ) is a du o matc h . No w, consider th e genome G 3 = +3 − 2 + 6 + 4 − 1 + 5 without d uplicates. W e h a ve G 3 [+6 , − 1] = { 1 , 4 , 6 } and [+6 , − 1] G 3 = (+6 , +4 , − 1). Br e akp oints, adjac encies, c ommon and c onserve d intervals. Let u s no w d efine the f our measures w e will study in this pap er. Let G 1 and G 2 b e tw o genomes withou t duplicates and with the same gene conte nt , that is F G 1 = F G 2 . Br e akp oint and A djac ency. Let ( a, b ) b e a du o in G 1 . W e say that the duo ( a, b ) in d uces a br e akp oint of ( G 1 , G 2 ) if neither ( a, b ) nor ( − b, − a ) is a duo in G 2 . Otherwise, w e say that ( a, b ) induces an adjac ency of ( G 1 , G 2 ). F or example, wh en G 1 = +1 + 2 + 3 + 4 + 5 an d G 2 = +5 − 2 4 − 3 + 2 + 1, the d uo (2 , 3) in G 1 induces a breakp oin t of ( G 1 , G 2 ) while (3 , 4) in G 1 induces an adjacency of ( G 1 , G 2 ). W e note B ( G 1 , G 2 ) (resp. A ( G 1 , G 2 )) th e n umber of breakp oin ts (resp. the n umber of adjacencies) that exist b et w een G 1 and G 2 . Common interval. A c ommon interval of ( G 1 , G 2 ) is a substr ing of G 1 suc h that G 2 con tains a p ermutatio n of this substrin g (not taking signs in to accoun t). F or example, consider G 1 = +1 + 2 + 3 + 4 + 5 and G 2 = +2 − 4 + 3 + 5 + 1. The s ubstring [+3 , +5] G 1 is a common int erv al of ( G 1 , G 2 ). Conserve d interval. Consider tw o signed genes a and b of G 1 suc h th at a precedes b , where the precedence relation is large in the sens e that, p ossibly , a = b . The sub string [ a, b ] G 1 is a c onserve d interval of ( G 1 , G 2 ) if either (i) a precedes b and G 2 [ a, b ] = G 1 [ a, b ], or (ii) − b precedes − a and G 2 [ − b, − a ] = G 1 [ a, b ]. F or example, if G 1 = +1 + 2 + 3 + 4 + 5 and G 2 = − 5 − 4 + 3 − 2 + 1, the substrin g [+2 , +5] G 1 is a conserv ed in terv al of ( G 1 , G 2 ). W e note that the notion of conserv ed in terv al do es not consider the sign of genes. Note also that a conserved int erv al is actually a common in terv al, but with additional restrictions on its extremities. De aling with duplic ates in genomes. When genomes con tain duplicates, w e cannot directly com- pute the measures defin ed in the previous paragraph. A solution consists in fin ding a one-to -one corresp onden ce (i.e. a matc hin g) b etw een dup licated genes of G 1 and G 2 ; w e then use this corre- sp ond ence to rename genes of G 1 and G 2 , and we delete the unmatched signed genes in order to obtain t w o genomes G ′ 1 and G ′ 2 suc h that G ′ 2 is a p ermutation of G ′ 1 ; thus, the measur e compu- tation b ecomes p ossible. In this pap er, we will fo cu s on th ree mo dels of matc hing : the exemplar , interme diate and maximum matching mo dels. – The exemplar mo del [23]: for eac h gene g , we keep in th e matc hing M only one o ccurrence of g in G 1 and in G 2 , and w e remov e all the other o ccurr ences. Hence, w e obtain tw o genomes G E 1 and G E 2 without dup licates. T h e triplet ( G E 1 , G E 2 , M ) is called an exemplariza tion of ( G 1 , G 2 ). Note that in th is mo del, M can b e inferred f rom the exemplarized genomes G E 1 and G E 2 . Thus, in the rest of the pap er, any exemp larization ( G E 1 , G E 2 , M ) of ( G 1 , G 2 ) will b e only describ ed b y the pair ( G E 1 , G E 2 ). – The interme diate mo del [3]: in this mo del, for eac h gene g , w e k eep in the matc hing M an arbitrary n umber k g , 1 ≤ k g ≤ min ( occ ( g , G 1 ) , occ ( g , G 2 )), in order to obtain genomes G I 1 and G I 2 . W e call the trip let ( G I 1 , G I 2 , M ) an interme diate matching of ( G 1 , G 2 ). – The maximum matching mo del [25]: in this case, w e k eep in the matc hing M the maxim u m n umber of signed genes in b oth genomes. More precisely , w e lo ok f or a one-to-one corresp ondence b et w een signed genes of G 1 and G 2 that matc h es, for eac h gene g , exactly min (o cc( g , G 1 ) , o cc( g , G 2 )) occurr en ces. After this op eration, we delete eac h unmatc hed signed gene. The triplet ( G M 1 , G M 2 , M ) obtained b y th is op eration is called a maximum matching of ( G 1 , G 2 ). Pr oblems studie d in this p ap er. Consider t w o genomes G 1 and G 2 with duplicates. Let EComI (resp. IComI , MComI ) b e the problem whic h consists in find ing an exemplarization (resp. inte r- mediate matc hing, maxim um matc hin g) ( G ′ 1 , G ′ 2 , M ) of ( G 1 , G 2 ) suc h that the n um b er of common in terv als of ( G ′ 1 , G ′ 2 ) is maximized. Moreo v er, let EConsI (resp. IConsI , MConsI ) b e the p roblem whic h consists in fi n ding an exemplarization (resp. inte rmediate matc h ing, maxim um matc hing) ( G ′ 1 , G ′ 2 , M ) of ( G 1 , G 2 ) such that the num b er of conserve d in terv als of ( G ′ 1 , G ′ 2 , M ) is maximized. In Section 2, w e pro v e the APX –hardness of EComI and EConsI , ev en for genomes G 1 and G 2 suc h that o cc( G 1 ) = 1 and o cc ( G 2 ) = 2. These results induce th e APX –hardn ess u nder the other mo dels (i.e., IComI , MComI , I ConsI and MCons I are APX –hard). These resu lts extend in p articular those of [7, 10]. 3 Let EBD (resp. IBD , MBD ) b e the problem w h ic h consists in finding an exemplarization (resp. in termediate matc hing, maxim um matc hing) ( G ′ 1 , G ′ 2 , M ) of ( G 1 , G 2 ) that minimizes the n um b er of breakp oints b etw een G ′ 1 and G ′ 2 . In S ection 3, we prov e the APX –hardness of EBD , even for genomes G 1 and G 2 suc h that o cc( G 1 ) = 1 and occ ( G 2 ) = 2. This result implies that IBD and MBD are also APX –hard, and extends th ose of [13]. Let Z EBD (resp. ZIBD , ZMBD ) b e the problem wh ic h consists in d etermining, for t w o genomes G 1 and G 2 , whether th er e exists an exemplarization (resp. in termediate matc h in g, maxim um matc h- ing) w hic h induces zer o br e akp oint . In section 4, w e study the complexity of ZEBD , ZMBD and ZIBD : in particular, we extend a r esult of [13] b y provi ng ZEBD to b e NP –complete for a new class of instances. W e also note that the problems ZEBD and ZIBD are equiv alent , and w e sh o w that Z MBD is in P . Finally , in S ection 5, w e fo cu s on a fourth measure, closely related to the n um b er of breakp oints: the numb er of adjac encies , for w hic h w e giv e sev eral constan t ratio app ro ximation algorithms in the maxim u m matc hing mo del, in the case w h ere genomes are balanced. 2 EComI and EConsI are A PX–hard Consider t wo genomes G 1 and G 2 with d uplicates, and let EComI ( resp. IComI , MComI ) b e the problem wh ic h consists in finding an exemplarization (resp. in termediate matc hing, maximum matc hin g) ( G ′ 1 , G ′ 2 , M ) of ( G 1 , G 2 ) such that the num b er of common interv als of ( G ′ 1 , G ′ 2 ) is maxi- mized. Moreo ver, let EConsI (resp. IConsI , MConsI ) b e the problem whic h consists in finding an exemplarization (r esp. in termediate matc hin g, m axim um matc h ing) ( G ′ 1 , G ′ 2 , M ) of ( G 1 , G 2 ) such that th e n u m b er of conserv ed in terv als of ( G ′ 1 , G ′ 2 , M ) is maximized. EComI and M ComI hav e b een prov ed to b e NP –complete ev en if o cc( G 1 ) = 1 and o cc ( G 2 ) = 2 in [10]. Besides, in [6], Blin and Rizzi h av e stud ied the pr oblem of computing a distance built on the n umber of conserved interv als. This d istance differs from the n umber of conserv ed in terv als w e study in this pap er, mainly in the sen s e th at (i) it can b e applied to t w o sets of genomes (as opp osed to tw o genomes in our case), and (ii) the distance b et w een t w o identica l genomes of length n is equal to 0 (as opp osed to n ( n +1) 2 in our case). Blin and Rizzi [6] prov ed that findin g the minim um distance is NP –complete, u nder b oth the exemplar and maximum matc h ing mo dels. A closer analysis of their p ro of sho ws that it can b e easily ad ap ted to prov e that EConsI and MConsI are NP –complete, ev en in the case o cc( G 1 ) = 1. W e can conclude fr om the ab o v e r esults th at IComI and IConsI are also NP –co mplete , since when one genome con tains n o duplicates, exemplar , interme diate and maximum matching mo dels are equiv alen t. In this section, we improv e the ab o v e results by sho wing that the six problems EComI , IComI , MComI , EConsI , IConsI and MConsI are APX –hard, ev en when genomes G 1 and G 2 are suc h that o cc( G 1 ) = 1 an d o cc( G 2 ) = 2. T he main r esult is T h eorem 1, which will b e completed b y Corollary 1 at the end of the section. Theorem 1. EComI and EConsI ar e APX –ha r d even when genomes G 1 and G 2 ar e such that o cc( G 1 ) = 1 and o cc( G 2 ) = 2 . W e p ro v e Theorem 1 by using an L-r e duction [22] fr om the Min-Ver te x-Co ver problem on cubic graphs, d en oted here Min-Ver tex-Cover-3 . Let G = ( V , E ) b e a cubic graph, i.e. for all v ∈ V , deg r ee ( v ) = 3. A set of vertices V ′ ⊆ V is called a vertex c over of G if for eac h edge e ∈ E , 4 there exists a v ertex v ∈ V ′ suc h that e is inciden t to v . The problem Min-Ver tex-Cover-3 is defined as follo ws: Problem: Min-Ver te x-Co ver-3 Input: A cubic grap h G = ( V , E ). Solution: A v ertex co ver V ′ of G . Measure: Th e cardinalit y of V ′ . Min-Ver tex-Cover-3 w as pro v ed to b e APX –complete in [1]. 2.1 Reduction Let G = ( V , E ) b e an instance of Min-Ver tex-Co ver-3 , wh ere G is a cub ic graph w ith V = { v 1 . . . v n } and E = { e 1 . . . e m } . Consider the tran s formation R wh ic h asso ciates to the graph G t wo genomes G 1 and G 2 in th e follo wing w a y , where eac h gene has a p ositiv e sign. G 1 = b 1 b 2 . . . b m x a 1 C 1 f 1 a 2 C 2 f 2 . . . a n C n f n y b m + n , b m + n − 1 . . . b m +1 (1) G 2 = y a 1 D 1 f 1 b m +1 a 2 D 2 f 2 b m +2 . . . b m + n − 1 a n D n f n b m + n x (2) with : – for eac h i , 1 ≤ i ≤ n, a i = 6 i − 5, f i = 6 i – for eac h i , 1 ≤ i ≤ n, C i = ( a i + 1) , ( a i + 2) , ( a i + 3) , ( a i + 4) – for eac h i , 1 ≤ i ≤ n + m, b i = 6 n + i – x = 7 n + m + 1 and y = 7 n + m + 2 – for eac h i , 1 ≤ i ≤ n, D i = ( a i + 3) , ( b j i ) , ( a i + 1) , ( b k i ) , ( a i + 4) , ( b l i ) , ( a i + 2) where e j i , e k i and e l i are the edges whic h are inciden t to v i in G , with j i < k i < l i . In the follo w in g, genes b i , 1 ≤ i ≤ m , are called markers . There is no du plicated gene in G 1 and the mark ers are th e only duplicated genes in G 2 ; these genes o ccur twice in G 2 . Hence, w e ha v e o cc( G 1 ) = 1 and o cc( G 2 ) = 2. V 2 V 1 V 3 V 4 e 2 e 5 e 1 e 6 e 4 e 3 Fig. 1. The cu bic graph G . T o illustrate the redu ction, consider the cub ic graph G of Figure 1. F rom G , we construct the follo wing genomes G 1 and G 2 : b 1 z}|{ 25 b 2 z}|{ 26 b 3 z}|{ 27 b 4 z}|{ 28 b 5 z}|{ 29 b 6 z}|{ 30 x z}|{ 35 1 C 1 z }| { 2 3 4 5 6 7 C 2 z }| { 8 9 10 11 12 13 C 3 z }| { 14 15 16 17 18 19 C 4 z }| { 20 21 22 23 24 y z}|{ 36 b 10 z}|{ 34 b 9 z}|{ 33 b 8 z}|{ 32 b 7 z}|{ 31 36 |{z} y 1 4 25 2 26 5 27 3 | {z } D 1 6 31 |{z} b 7 7 10 25 8 28 11 29 9 | {z } D 2 12 32 |{z} b 8 13 16 26 14 28 17 30 15 | {z } D 3 18 33 |{z} b 9 19 22 27 20 29 23 30 21 | {z } D 4 24 34 |{z} b 10 35 |{z} x 5 2.2 Preliminary results In order to pr o ve Theorem 1, w e first giv e f our inte rmediate lemmas. In the follo wing, a common in terv al for the EComI problem or a conserv ed int erv al for EConsI is called a r obust interval . Besides, a trivial interval will denote either an inte rv al of length one (i.e. a singleton), or the whole genome. Lemma 1. F or any exe mplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ) , the non trivial r obust intervals of ( G 1 , G E 2 ) ar e ne c essarily c ontaine d in some se quenc e a i C i f i of G 1 ( 1 ≤ i ≤ n ). Pr o of. W e start by p ro ving the lemma for common interv als, and we will then extend it to conserv ed in terv als. First, w e pr o v e that, for an y exemplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ), eac h common int erv al I s u c h that | I | ≥ 2 con tains either b oth of x , y or n one of them. This further implies that I cov ers the whole genome. Supp ose there exists a common in terv al I x (recall that b y definition I x is on G 1 ) suc h that | I x | ≥ 2 and I x con tains x . Let P I x b e the p ermutati on of I x in G E 2 . The interv al I x m ust conta in either b m or a 1 . Let us detail eac h of the t w o cases: (a) If I x con tains b m , then P I x con tains b m to o. Notice that there is some i , 1 ≤ i ≤ n , such that b m b elongs to D i in G E 2 . Then P I x con tains all genes b et ween D i and x in G E 2 . Thus P I x con tains b m + n . Consequen tly , I x con tains b m + n and it also con tains y . (b) If I x con tains a 1 , then P I x con tains a 1 to o. Th en P I x con tains all genes b et ween a 1 and x . Th us P I x con tains b m + n . Hence, I x con tains b m + n and then it also cont ains y . No w, sup p ose that I y is a common interv al suc h that | I y | ≥ 2 and I y con tains y . Let P I y b e the p ermutatio n of I y on G E 2 . The in terv al I y m ust con tain either b m + n or f n . Let us detail eac h of the t wo cases: (a) If I y con tains b m + n , then P I y con tains b m + n to o. Th us P I y con tains all genes b et w een b m + n and y . Hence P I y con tains all the sequen ces D i , 1 ≤ i ≤ n . In p articular, P I y con tains all the mark ers and consequent ly I y m ust con tain x . (b) If I y con tains f n , then P I y con tains f n to o. Then P I y con tains all genes b et ween f n and y . In particular, P I y con tains b m + n − 1 and then I y con tains b m + n − 1 to o. Hence, I y also con tains b m + n , similarly to the previous case. Th us I y con tains x . W e conclude that eac h non singleton common in terv al conta ining either x or y necessarily con tains b oth x and y . Therefore, and by construction of G 2 , there is only one suc h in terv al, that is G 1 itself. Hence, an y non trivial common in terv al is necessarily , in G 1 , either strictly on th e left of x , or b et w een x an d y , or str ictly on the righ t of y . Let u s analyze these differen t cases: – Let I b e a n on trivial common inte rv al situated strictly on the left of x in G 1 . Thus I is a s equence of at least t w o consecutiv e m ark ers. Since in an y exemplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ), ev ery mark er in G E 2 has neigh b orin g genes w hic h are not marke rs, this contradicts the fact that I is a common inte rv al. – Let I b e a non tr ivial common interv al situated strictly on the righ t of y in G 1 . Th en I is a substring of b m + n , . . . , b m +1 con taining at least t wo genes. In an y exemplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ), for eac h pair ( b m + i , b m + i +1 ) of G E 2 , with 1 ≤ i < n , we h a v e a i +1 ∈ G E 2 [ b m + i , b m + i +1 ]. This con tradicts the fact that I is strictly on the righ t of y in G 1 . 6 – Let I b e a non trivial common in terv al lying b etw een x and y in G 1 . F or any exemplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ), a common interv al cannot con tain, in G 1 , b oth f i and a i +1 for some i , 1 ≤ i ≤ n − 1 (since b m + i is situated b et w een f i and a i +1 in G E 2 and on the r igh t of x in G 1 ). Hence, a non trivial common in terv al of ( G 1 , G E 2 ) is included in some s equence a i C i f i in G 1 , 1 ≤ i ≤ n . This pro v es the lemma for common in terv als. By definition, an y conserv ed in terv al is necessarily a common inte rv al. So, a n on tr ivial conserv ed int erv al of ( G 1 , G E 2 ) is includ ed in some s equ ence a i C i f i in G 1 , 1 ≤ i ≤ n . The lemma is pro ved. ⊓ ⊔ Lemma 2. L et ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and i ∈ [1 . . . n ] . L et ∆ i b e a substring of [ a i + 3 , a i + 2] G E 2 that do es not c ontain any marker. If | ∆ i | ∈ { 2 , 3 } , then ther e is no r obust interval I of ( G 1 , G E 2 ) such that ∆ i is a p ermutation of I . Pr o of. First, we prov e that there is no p ermutatio n I of ∆ i suc h that I is a common in terv al of ( G 1 , G E 2 ). Next, w e show that th ere is no p ermutati on I of ∆ i suc h that I is a conserved int erv al. By Lemma 1, w e kn o w that a non trivial common interv al of ( G 1 , G E 2 ) is a su bstring of some sequence a i C i f i , 1 ≤ i ≤ n . This substring con tains only consecutiv e in tegers. Therefore, if there exists a p ermutatio n I of ∆ i suc h that I is a common interv al of ( G 1 , G E 2 ), then ∆ i m ust b e a p ermuta tion of consecutiv e in tegers. If | ∆ i | = 2, we ha v e ∆ i = ( p, q ) w here p and q are not consecutive inte gers and if | ∆ i | = 3, then we h av e ∆ i = ( a i + 3 , a i + 1 , a i + 4) or ∆ i = ( a i + 1 , a i + 4 , a i + 2). In these three cases, ∆ i is not a p ermutati on of consecutiv e in tegers. Hence, there is n o p ermutatio n I of ∆ i suc h that I is a common interv al of ( G 1 , G E 2 ). Moreo v er, any conserv ed int erv al is also a common in terv al. T h us, there is no p ermutati on I of ∆ i suc h that I is a conserv ed in terv al of ( G 1 , G E 2 ). ⊓ ⊔ F or more clarit y , let us n o w introd uce some notations. Giv en a graph G = ( V , E ), let V C = { v i 1 , v i 2 . . . v i k } b e a vertex co ver of G . Let R ( G ) = ( G 1 , G 2 ) b e the p air of genomes d efined b y the construction describ ed in (1) and (2). No w, let F b e the function whic h asso ciates to V C , G 1 and G 2 an exemplarizatio n F ( V C ) of ( G 1 , G 2 ) as follo ws . In G 2 , all the mark ers are remov ed from the sequences D i for all i 6 = i 1 , i 2 . . . i k . Next, for eac h mark er wh ich is still present t wice, one of its o ccur rences is arbitrarily remov ed. S ince in G 2 only mark ers are du p licated, we conclude that F ( V C ) is an exemplarizatio n of ( G 1 , G 2 ). Giv en a cubic graph G and genomes G 1 and G 2 obtained b y the transformation R ( G ), let us define the function S wh ic h asso ciates to an exemplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ) the v ertex cov er V C of G defined as follo ws : V C = { v i | 1 ≤ i ≤ n ∧ ∃ j ∈ { 1 . . . m } , b j ∈ G E 2 [ a i , f i ] } . In other words, w e k eep in V C the vertice s v i of G for w hic h there exists some gene b j suc h that b j is in G E 2 [ a i , f i ]. W e no w prov e that V C is a v ertex co v er. Consid er an edge e p of G . By construction of G 1 and G 2 , there exists some i , 1 ≤ i ≤ n , su c h that gene b p is lo cated b et w een a i and f i in G E 2 . Th e pr esence of gene b p b et w een a i and f i implies that v ertex v i b elongs to V C . W e conclude th at eac h edge is inciden t to at least on e v ertex of V C . Let W b e the fun ction defined on { EConsI , EComI } by W (pb) = 1 if p b = ECons I and W (pb) = 4 if pb = EComI . Let opt P ( A ) b e the optim um resu lt of an instance A for an optimization problem pb, pb ∈ { EcomI , EConsI , Min-Ver tex -Co ver-3 } . W e now d efine the function T w hose argumen ts are a problem pb ∈ { EConsI , EComI } and a cubic graph G . Let R ( G ) = ( G 1 , G E 2 ) as usual. Th en T (pb , G ) is d efined as the num b er of robust trivial interv als of ( G 1 , G E 2 ) with resp ect to pb. Let n and m b e r esp ectiv ely th e n um b er of v ertices 7 and the num b er of edges of G . W e ha v e T ( EConsI , G ) = 7 n + m + 2 and T ( EComI , G ) = 7 n + m + 3. Indeed, for EComI , there are 7 n + m + 2 singletons and w e also need to consider the whole genome. Lemma 3. L et pb ∈ { EcomI , EConsI } . L e t G b e a cubic gr aph and R ( G ) = ( G 1 , G 2 ) . L et ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and let i , 1 ≤ i ≤ n . Then only two c ases c an o c- cur with r esp e ct to D i . 1. Either in G E 2 , al l the markers fr om D i wer e r emove d, and in this c ase, ther e ar e exactly W ( pb ) non trivial r obust intervals involving D i . 2. Or in G E 2 , at le ast one marker was kept in D i , and i n this c ase, ther e i s no non trivial r obust interval involving D i . Pr o of. W e fir st prov e th e lemma for the EComI problem and then we extend it to EConsI . Lemma 1 im p lies that eac h non trivial common in terv al I of ( G 1 , G E 2 ) is cont ained in some su bstring of a i C i f i , 1 ≤ i ≤ n . So, th e p ermuta tion of I on G E 2 is conta ined in a su bstring of a i D i f i , 1 ≤ i ≤ n . Consider i , 1 ≤ i ≤ n , and supp ose that all the mark ers from D i are remov ed on G E 2 . Thus, a i C i f i , C i , a i C i and C i f i are common in terv als of ( G 1 , G E 2 ). L et us n o w sh ow that there is no other non trivial common interv al inv olving D i . Let ∆ i b e a substring of [ a i + 3 , a i + 2] G E 2 suc h that | ∆ i | ∈ { 2 , 3 } . By Lemma 2, we kno w that ∆ i is not a common interv al. The remaining interv als are ( a i , a i + 3), ( a i , a i + 3 , a i + 1), ( a i , a i + 3 , a i + 1 , a i + 4), ( a i + 1 , a i + 4 , a i + 2 , f i ), ( a i + 4 , a i + 2 , f i ) and ( a i + 2 , f i ). By construction, none of them can b e a common interv al, b ecause none of them is a p ermutation of consecutiv e inte gers. Hence, th ere are only four non trivial common inte rv als in v olving D i in G E 2 . Among these four common inte rv als, only a i C i f i is a conserv ed interv al to o. In the end, if all the mark ers are remo v ed from D i , there are exactly four non trivial common in terv als and one non trivial conserv ed inte rv al inv olving D i . So, giv en a p r oblem pb ∈ { EcomI , EconsI } , there are exactly W (pb) n on trivial robust in terv als inv olving D i . No w, supp ose that at least one mark er of D i is kept in G E 2 . Lemma 1 sho ws that eac h n on trivial common in terv al I of ( G 1 , G E 2 ) is con tained in some subs tring of a i C i f i , 1 ≤ i ≤ n . Since no mark er is p r esen t in a s equ ence a i C i f i , w e ded u ce that there do es n ot exist an y tr ivial common in terv al contai ning a mark er. So, a non trivial common in terv al inv olving D i only must con tain a substring ∆ i of [ a i + 3 , a i + 2] G E 2 suc h that ∆ i con tains no marker. Sin ce n o mark er is an extremit y of [ a i + 3 , a i + 2] G E 2 , w e ha v e | ∆ i | ≤ 3. By Lemma 2, w e kno w that ∆ i is not a common in terv al. The remaining interv als to b e consid ered are the interv als a i ∆ i and ∆ i f i . By constru ction of a i C i f i , these interv als are not common interv als (the abs en ce of gene a i + 2 for a i ∆ i and of gene a i + 3 for ∆ i f i implies that these in terv als are not a p ermutatio n of consecutiv e int egers). Hence, these in terv als cannot b e conserved interv als either. ⊓ ⊔ Lemma 4. L et pb ∈ { EcomI , EConsI } . L et G = ( V , E ) b e a cubic gr aph with V = { v 1 . . . v n } and E = { e 1 . . . e m } and let G 1 , G 2 b e the two genomes obtaine d by R ( G ) . 1. L et V C b e a vertex c over of G and denote k = | V C | . Then the exemplarization F ( V C ) of ( G 1 , G 2 ) has at le ast N = n W ( p b ) + T ( pb , G ) − W ( pb ) · k r obust intervals. 2. L et ( G 1 , G E 2 ) b e an e xe mplarization of ( G 1 , G 2 ) and let V C ′ b e the v ertex c over of G obtaine d by S ( G 1 , G E 2 ) . Then | V C ′ | = W ( pb ) · n + T ( pb ,G ) − N W ( pb ) , wher e N is the numb er of r obust intervals of ( G 1 , G E 2 ) . Pr o of. 1. Let pb ∈ { EcomI , EConsI } . Let G b e a cubic graph and let G 1 and G 2 b e the tw o genomes obtained by R ( G ). Let V C b e a vertex co v er of G and denote k = | V C | . Let ( G 1 , G E 2 ) b e the 8 exemplarization of ( G 1 , G 2 ) obtained by F ( V C ). By constr u ction, w e ha v e at least ( n − k ) su bstrings D i in G E 2 for which all the mark ers are remov ed. By L emm a 3, we kn o w that eac h of these substrings implies the existence of W (pb) n on trivial robu st in terv als. So, we hav e at least W (pb)( n − k ) non trivial robust in terv als. Moreo v er , by d efinition of T (pb , G ), the num b er of trivial robust in terv als of ( G 1 , G E 2 ) is exactly T (pb , G ) . Th us, w e ha v e at least N = W (pb) · n + T (pb , G ) − W (pb) · k robust in terv als of ( G 1 , G E 2 ). 2. Let ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and let n − j b e the n um b er of sequences D i , 1 ≤ i ≤ n , for whic h all mark er s ha v e b een d eleted in G E 2 . Then, by Lemmas 1 and 3, the n umber of robust interv als of ( G 1 , G E 2 ) is equal to N = W (pb) · n + T (p b , G ) − W (pb) · j . Let V C ′ b e the v ertex co ver obtained b y S ( G 1 , G E 2 ). Eac h mark er has one occurr ence in G E 2 and th ese o ccurrences lie in j s equ ences D i . So, by definition of S , we conclude th at | V C ′ | = j = W (pb) · n + T (pb ,G ) − N W (pb) . ⊓ ⊔ 2.3 Main result Let us firs t defin e the n otion of L-r e duction [22]: let A and B b e tw o optimizat ion problems and c A , c B b e resp ectiv ely their cost functions. An L-r e duction from problem A to p roblem B is a pair of p olynomial-time computable functions R an d S w ith the follo wing prop erties: ( a ) If x is an instance of A , then R ( x ) is an instance of B ; ( b ) If x is an instance of A and y is a solution of R ( x ), then S ( y ) is a solution of A ; ( c ) If x is an instance of A and R ( x ) is its corresp onding instance of B , then there is some p ositiv e constan t α s uc h th at opt B ( R ( x )) ≤ α. opt A ( x ) ; ( d ) If s is a solution of R ( x ), then ther e is some p ositive constan t β such that | opt A ( x ) − c A ( S ( s )) | ≤ β | opt B ( R ( x )) − c B ( s ) | . W e prov e Theorem 1 by showing th at the pair ( R, S ) defined p reviously is an L-r e duction from Min-Ver tex-Cover-3 to EConsI and fr om Min-Ver tex-Cover-3 to EComI . First n ote that prop erties ( a ) and ( b ) are ob viously satisfied b y R and S . Consider pb ∈ { EcomI , EC onsI } . Let G = ( V , E ) b e a cubic graph with n v ertices and m edges. W e n o w pro v e pr op erties ( c ) and ( d ). Con s ider the genomes G 1 and G 2 obtained b y R ( G ). F or sak e of clarit y , we abbreviate here and in the follo wing opt Min-Ver tex-Cover-3 to opt Min-VC . First, w e need to pro v e that there exists α ≥ 0 suc h that opt pb ( G 1 , G 2 ) ≤ α. opt Min-Ver tex-Cover-3 ( G ). Since G is cubic, we ha v e the follo wing prop erties: n ≥ 4 (3) m = 1 2 n X i =1 deg r ee ( v i ) = 3 n 2 (4) opt Min-VC ( G ) ≥ m 3 = n 2 (5) T o explain prop erty (5), remark that, in a cubic graph G with n v ertices and m edges, eac h v ertex co v er s thr ee edges. Th us, a set of k v ertices co ve rs at most 3 k edges. Hence, an y v ertex co ver of G m ust con tain at least m 3 v ertices. By Lemma 3, we kno w that sequences of the f orm a i C i f i , 1 ≤ i ≤ n , conta in either zero or W (pb) non trivial robust in terv als. By Lemm a 1, there are no other n on trivial robust inte rv als. So, w e ha v e the f ollo wing inequalit y : 9 opt pb ( G 1 , G 2 ) ≤ T (pb , G ) | {z } triv ial robus t inter v als + W (pb) · n If p b = EComI , w e ha ve: opt EComI ( G 1 , G 2 ) ≤ 7 n + m + 3 + 4 n opt EComI ( G 1 , G 2 ) ≤ 27 n 2 by (3) an d (4) (6) And if pb = EConsI , we ha v e : opt EConsI ( G 1 , G 2 ) ≤ 7 n + m + 2 + n opt EConsI ( G 1 , G 2 ) ≤ 21 n 2 by (3) and (4) (7) Altoget her, b y (5), (6) and (7), w e pr o v e p rop erty ( c ) with α = 27. No w, let us p ro v e p rop erty ( d ). Let V C = { v i 1 , v i 2 . . . v i P } b e a min im um v ertex co v er of G . Then P = opt Min-VC ( G ). Let G 1 and G 2 b e th e genomes obtained by R ( G ). Let ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and let k ′ b e the num b er of r obust in terv als of ( G 1 , G E 2 ). Finally , let V C ′ b e the v ertex cov er of G suc h that V C ′ = S ( G 1 , G E 2 ). W e need to find a p ositiv e constant β suc h th at | P − | V C ′ || ≤ β | opt pb ( G 1 , G 2 ) − k ′ | . F or pb ∈ { EcomI , EC onsI } , let N pb b e the num b er of robust in terv als b etw een the t w o genomes obtained b y F ( V C ). By the fir st pr op erty of Lemma 4, we hav e opt pb ( G 1 , G 2 ) ≥ N pb ≥ W (pb) · n + T (pb , G ) − W (pb) · P So, it is sufficien t to prov e that there exists some β ≥ 0 suc h that | P − | V C ′ || ≤ β | W (pb) · n + T (pb , G ) − W (pb) · P − k ′ | . By th e second prop ert y of Lemm a 4, w e h av e | V C ′ | = W (pb) · n + T (pb ,G ) − k ′ W (pb) . Since P ≤ | V C ′ | , w e ha v e | P − | V C ′ || = | V C ′ | − P = W (pb) · n + T (pb ,G ) − k ′ W (pb) − P = 1 W (pb) ( W (pb) · n + T (pb , G ) − W (pb) · P − k ′ ). So β = 1 is su fficien t in b oth cases, since W ( EComI ) = 4 and W ( EConsI ) = 1, whic h implies 1 W (pb) ≤ 1. Altoget her, w e th en h a ve | opt Min-VC ( G ) − | V C ′ || ≤ 1 · | opt pb ( G 1 , G 2 ) − k ′ | . W e prov ed that the reduction ( R, S ) is an L-r e duction . This implies that for t w o genomes G 1 and G 2 , b oth p roblems EConsI and EComI are APX –hard even if o cc( G 1 ) = 1 and o cc( G 2 ) = 2. Theorem 1 is pro ved. ⊓ ⊔ W e extend in Corollary 1 our results for the interme diate and maximum matching mo dels. Corollary 1. IComI , MComI , IConsI and M ConsI ar e APX –ha r d even when genomes G 1 and G 2 ar e such that o cc( G 1 ) = 1 and o cc( G 2 ) = 2 . Pr o of. The interme diate and maximum matc h ing mo dels are iden tical to the exemplar mo del when one of the t w o genomes con tains no du plicates. Hence, the APX –hardness result for EComI (resp. EConsI ) also holds for IComI and MComI (resp. IConsI and MCons I ). ⊓ ⊔ 10 3 EBD is APX–hard Consider t wo genomes G 1 and G 2 with duplicates, and let EBD (resp . IBD , MBD ) b e the problem whic h consists in fi n ding an exemplarization (resp. inte rmediate matc h ing, maxim um matc hing) ( G ′ 1 , G ′ 2 , M ) of ( G 1 , G 2 )that min imizes th e n u m b er of breakp oints b etw een G ′ 1 and G ′ 2 . EBD h as b een prov ed to b e NP –complete ev en if o cc( G 1 ) = 1 and o cc( G 2 ) = 2 [7]. Some inapproxi mabilit y results also exist: in particular, it has b een pro ved in [13] th at, in the general case, EBD cannot b e appr o x im ated within a factor c log n , where c > 0 is a constan t, and cannot b e appro ximated within a factor 1 . 36 w hen o cc ( G 1 ) = o cc( G 2 ) = 2. Moreo v er , for t w o balanced genomes G 1 and G 2 suc h th at k = o cc( G 1 ) = occ( G 2 ), sev eral approxima tion algorithms for MBD are giv en. These approximat ion algo rithms admit resp ectiv ely a ratio of 1 . 103 7 when k = 2 [17], 4 wh en k = 3 [17] and 4 k in th e general case [19]. W e can conclude fr om the ab o v e results that IBD and MBD problems are also NP –complete, s in ce when one genome con tains no du p licates, exemplar , i nterme diate and maximum matching mo dels are equiv alent . In this section, w e impro v e the ab ov e results b y showing that the three pr oblems EBD , IBD and MBD are APX –hard, ev en when genomes G 1 and G 2 are s uc h that o cc( G 1 ) = 1 and o cc( G 2 ) = 2. The main resu lt is Theorem 2 b elo w, which will b e completed by Corollary 2 at the end of the section. Theorem 2. EBD is APX –h ar d even when genomes G 1 and G 2 ar e such that o cc( G 1 ) = 1 and o cc( G 2 ) = 2 . T o pr ov e Th eorem 2, we u se an L-R e duction from Min-Ver tex-Cover-3 to EBD . Let G = ( V , E ) b e a cubic graph with V = { v 1 . . . v n } and E = { e 1 . . . e m } . F or eac h i , 1 ≤ i ≤ n , let e f i , e g i and e h i b e the three edges which are in ciden t to v i in G with f i < g i < h i . Let R ′ b e the p olynomial transformation w hic h asso ciates to G the follo w ing genomes G 1 and G 2 , where eac h gene has a p ositiv e sign: G 1 = a 0 a 1 b 1 a 2 b 2 . . . a n b n c 1 d 1 c 2 d 2 . . . c m d m c m +1 G 2 = a 0 a n d f n d g n d h n b n . . . a 2 d f 2 d g 2 d h 2 b 2 a 1 d f 1 d g 1 d h 1 b 1 c 1 c 2 . . . c m c m +1 with : – a 0 = 0, and for eac h i , 1 ≤ i ≤ n , a i = i and b i = n + i – c m +1 = 2 n + m + 1, and for eac h i , 1 ≤ i ≤ m , c i = 2 n + i and d i = 2 n + m + 1 + i W e remark that there is no d uplication in G 1 , so o cc( G 1 ) = 1. In G 2 , only the genes d i , 1 ≤ i ≤ m , are duplicated and occur t wice. Th us o cc( G 2 ) = 2. Let G b e a cubic graph and V C b e a v ertex co v er of G . Let G 1 and G 2 b e the genomes obtained b y R ′ ( G ). W e define F ′ to b e the p olynomial transf orm ation wh ic h asso ciates to V C , G 1 and G 2 the exemplarization F ′ ( V C ) = ( G 1 , G E 2 ) of ( G 1 , G 2 ) as follo ws. F or eac h i such that v i / ∈ V C , w e remo v e fr om G 2 the genes d f i , d g i and d h i . Then, for eac h j , 1 ≤ j ≤ m suc h that d j still h as t w o o ccurrences in G 2 , we arb itrarily r emo v e one of these o ccurr ences in ord er to obtain the genome G E 2 . Hence, ( G 1 , G E 2 ) is an exemplarizat ion of ( G 1 , G 2 ). Giv en a cubic graph G , w e construct G 1 and G 2 b y the transformation R ′ ( G ). Giv en an ex- emplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ), let S ′ b e the p olynomial transformation which asso ciates to ( G 1 , G E 2 ) the set V C = { v i | 1 ≤ i ≤ n , a i and b i are not consecutiv e in G E 2 } . W e claim that V C is a v ertex co ver of G . Indeed, let e p , 1 ≤ p ≤ m , b e an edge of G . Genome G E 2 con tains one o ccurr ence of gene d p since G E 2 is an exemplarization of G 2 . By constru ction, there exists i , 1 ≤ i ≤ n , su c h 11 that d p is in G E 2 [ a i , b i ] and suc h that e p is inciden t to v i . The pr esence of d p in G E 2 [ a i , b i ] implies that v ertex v i b elongs to V C . W e can conclude th at eac h edge of G is in ciden t to at least one vertex of V C . Lemmas 5 and 6 b elo w are us ed to p ro v e th at ( R ′ , S ′ ) is an L- R e duction from th e Min-Ver tex- Co ver-3 problem to the EBD problem. Let G = ( V , E ) b e a cubic graph with V = { v 1 , v 2 . . . v n } and E = { e 1 , e 2 . . . e m } an d let us construct ( G 1 , G 2 ) by the transformation R ′ ( G ). Lemma 5. L et V C b e a vertex c over of G and ( G 1 , G E 2 ) the exemplarizat ion given by F ′ ( V C ) . Then | V C | = k ⇒ B ( G 1 , G E 2 ) ≤ n + 2 m + k + 1 , wher e B ( G 1 , G E 2 ) is the numb er of br e akp oints b etwe en G 1 and G E 2 . Pr o of. Supp ose | V C | = k . Let us list the breakp oin ts b et ween genomes G 1 and G E 2 obtained by F ′ ( V C ). T he pairs ( b i , a i +1 ), 1 ≤ i ≤ n − 1, and ( b n , c 1 ) ind uce one br eakp oin t eac h . F or all i , 1 ≤ i ≤ m , eac h pair of the form ( c i , d i ) (resp. ( d i , c i +1 )) indu ces one breakp oint. F or all i , 1 ≤ i ≤ n , suc h that v i ∈ V C , ( a i , b i ) induces at most one breakp oint . Fin ally , the pair ( a 0 , a 1 ) induces one breakp oin t. Thus there are at most n + 2 m + k + 1 breakp oints of ( G 1 , G E 2 ). ⊓ ⊔ Lemma 6. L et ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and V C ′ b e the vertex c over of G obtaine d by S ′ ( G 1 , G E 2 ) . We have B ( G 1 , G E 2 ) = k ′ ⇒ | V C ′ | = k ′ − n − 2 m − 1 . Pr o of. Let ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and V C ′ b e the v ertex cov er obtained b y S ′ ( G 1 , G E 2 ). Su p p ose B ( G 1 , G E 2 ) = k ′ . F or an y exemplarization ( G 1 , G E 2 ) of ( G 1 , G 2 ), th e follo win g breakp oints alw ays o ccur: the pair ( a 0 , a 1 ) ; for eac h i , 1 ≤ i ≤ m , eac h pair ( c i , d i ) and ( d i , c i +1 ) ; for eac h i , 1 ≤ i ≤ n − 1, the pair ( b i , a i +1 ) ; the pair ( b n , c 1 ). Thus, w e h a ve at least n + 2 m + 1 breakp oints. The other p ossible br eakp oin ts are in duced b y pairs of the form of ( a i , b i ). Since we ha v e B ( G 1 , G E 2 ) = k ′ , there are exactly k ′ − n − 2 m − 1 suc h breakp oints. By construction of V C ′ , the cardinalit y of V C ′ is equal to the n um b er of breakp oints indu ced b y pairs of the f orm ( a i , b i ). So, w e ha v e: | V C ′ | = k ′ − n − 2 m − 1. ⊓ ⊔ T o p ro v e that ( R ′ , S ′ ) is an L-r e duction , w e first notice that p rop erties (a) an d (b) of an L- r e duction are trivially ve rified. The next lemma pro v es prop ert y (c ) . Lemma 7. The ine quality opt EBD ( G 1 , G 2 ) ≤ 12 · opt Min-VC ( G ) holds. Pr o of. F or a cubic graph G with n v ertices and m edges, we hav e 2 m = 3 n (see (4)) and opt Min-VC ( G ) ≥ n 2 (see (5)). By construction of th e genomes G 1 and G 2 , any exemplarizatio n of ( G 1 , G 2 ) con tains 2 n + 2 m + 2 genes in eac h genome. Th us, we ha v e opt EBD ( G 1 , G 2 ) ≤ 2 n + 2 m + 2 ≤ 6 n ( n ≥ 4 in a cubic graph). Hence, w e conclude that opt EBD ( G 1 , G 2 ) ≤ 12 · opt Min-VC ( G ). ⊓ ⊔ No w, we prov e prop ert y ( d ) of our L-r e duction . Lemma 8. L et ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and let V C ′ b e the vertex c over of G obtaine d by S ′ ( G 1 , G E 2 ) . Then, we have | opt Min-VC ( G ) − | V C ′ || ≤ | opt EBD ( G 1 , G 2 ) − B ( G 1 , G E 2 ) | Pr o of. Let ( G 1 , G E 2 ) b e an exemplarization of ( G 1 , G 2 ) and V C ′ b e the vertex co ver of G obtained b y S ′ ( G 1 , G E 2 ). L et V C b e a ve rtex co ve r of G su ch that | V C | = opt Min-VC ( G ). W e kn ow that opt Min-VC ( G ) ≤ | V C ′ | and opt EBD ( G 1 , G 2 ) ≤ B ( G 1 , G E 2 ). So, it is sufficient to pro v e | V C ′ | − opt Min-VC ( G ) ≤ B ( G 1 , G E 2 ) − opt EBD ( G 1 , G 2 ). 12 By Lemma 5, we ha v e B ( F ′ ( V C )) ≤ n + 2 m + 1 + opt Min-VC , wh ic h implies opt EBD ( G 1 , G 2 ) ≤ B ( F ′ ( V C )) ≤ n + 2 m + 1 + opt Min-VC . Then B ( G 1 , G E 2 ) − opt E B D ( G 1 , G 2 ) ≥ B ( G 1 , G E 2 ) − n − 2 m − 1 − opt Min-VC ( G ) (8) By Lemma 6, w e hav e: | V C ′ | = B ( G 1 , G E 2 ) − n − 2 m − 1 whic h imp lies | V C ′ | − opt Min-VC ( G ) = B ( G 1 , G E 2 ) − n − 2 m − 1 − opt Min-VC ( G ) (9) Finally , b y (8) and (9), we get | V C ′ | − opt Min-VC ≤ B ( G 1 , G E 2 ) − opt EBD ( G 1 , G 2 ). ⊓ ⊔ Lemmas 7 and 8 pro v e that the p air ( R ′ , S ′ ) is an L-r e duction from Min-Ver tex-Cover-3 to EBD . Hence, EBD is APX –hard ev en if o cc( G 1 ) = 1 and o cc( G 2 ) = 2, and T heorem 2 is p ro v ed. W e extend in Corollary 2 our results for the interme diate and maximum matching mo dels. Corollary 2. The IBD and MBD pr oblems ar e APX –har d even when genomes G 1 and G 2 ar e such that o cc( G 1 ) = 1 and o cc( G 2 ) = 2 . Pr o of. The in termediate and maxim um matc hing mo d els are iden tical to the exemplar mo d el when one of the t w o genomes con tains n o duplicates. Hence, the APX –hardn ess result for EBD also holds for IBD and MBD . ⊓ ⊔ 4 Zero breakp oin t distance This section is dev oted to zero breakp oint d istance recognition issues. Indeed, in [13], the authors sho w ed that deciding whether the exemplar breakp oin t distance b et w een any tw o genomes is zero or not is NP –complete ev en wh en no gene o ccurs m ore than th ree times in b oth genomes, i. e. , instances of t yp e (3 , 3). This imp ortan t resu lt implies that the exemplar b reakp oint distance problem do es not admit an y appr o ximation in p olynomial-time, unless P = NP . F ollo wing this line of r esearc h , w e first complemen t the result of [13] by proving that deciding whether the exemplar br eakp oin t distance b et w een an y t w o genomes is zero or not is NP –complete, ev en wh en no gene is du plicated more than twice in one of th e genomes (the m axim um num b er of dup lications is ho wev er unboun ded in the other genome). Th is result is n ext extended to the inte rmediate matc hing mo d el and we giv e a practical - but exp onenti al - algorithm for deciding whether the exemplar breakp oint distance b et w een any tw o genomes is zero or not in case no gene o ccurs more than t wice in b oth genomes (a problem whose complexit y , P versus NP –co mplete, r emains op en). Finally , we show that deciding whether the maxim u m matc hing b reakp oint distance b et w een an y t wo genomes is zero or not is p olynomial-time solv able and hence that such negativ e app ro ximation r esults (the ones w e obtained for the exemplar and intermediate mo d els) do no propagate to the maximum m atc hing mo del. The f ollo wing easy observ ation will pro ve extremely useful in the sequel of the presen t sectio n. Observ ation 3 L et G 1 and G 2 b e two genomes. If the exemplar br e akp oint distanc e b etwe en G 1 and G 2 is zer o, then ther e exists an exemplarizatio n ( G E 1 , G E 2 ) of ( G 1 , G 2 ) such that (1) G E 1 = G E 2 , or (2) − ( G E 1 ) r = G E 2 , wher e − ( G E 1 ) r is the signe d r eversal of genome G 1 . The same observation c an b e made for the interme diate and maximum matching mo dels. 13 4.1 Zero exemplar brea kp oin t distance The zero exemplar breakp oint distance ( ZEBD ) pr ob lem is form ally defined as follo ws. Problem: ZEBD Input: Tw o genomes G 1 and G 2 . Question: I s the exemplar breakp oin t d istance b et w een G 1 and G 2 equal to zero? Aiming at p recisely defining the in appro ximabilit y landscap e of computing th e exemplar b r eak- p oint distance b et w een t w o genomes, we complemen t the result of [13], who show ed ZEBD to b e NP –complete ev en for instances of t yp e (3 , 3), by the follo wing theorem. Theorem 4. ZEBD i s NP –c omplete even if no gene o c c u rs mor e than twic e in G 1 . Pr o of. Mem b ership of ZEBD to NP is immediate. The r eduction w e u s e to pro v e hardn ess is from Min-Ver tex-Cover [16]. Let an arbitrary instance of Min-Ver tex-Cover b e giv en b y a graph G = ( V , E ) and a p ositiv e int eger k . W r ite V = { v 1 , v 2 . . . v n } and E = { e 1 , e 2 . . . e m } . In th e rest of the pro of, elements of V (resp. E ) will b e seen either as v ertices (resp. edges) or genes, dep endin g on the con text. The corresp onding instance ( G 1 , G 2 ) of ZEBD is defined as follo ws: G 1 = v 1 X 1 v 2 X 2 . . . v n X n G 2 = Y [1] Y [2] . . . Y [ k ] Y V . F or eac h i = 1 , 2 , . . . , n , X i is defined to b e X i = e i 1 e i 2 . . . e i j , where e i 1 , e i 2 , . . . , e i j , i 1 < i 2 < . . . < i j , are the edges inciden t to v ertex v i . Th e strings Y [ i ], 1 ≤ i ≤ k , are all equ al and are defined by Y [ i ] = Y V Y E where Y V = v 1 v 2 . . . v n and Y E = e 1 e 2 . . . e m . Notice that no gene o ccur s more than t wice in G 1 (actually genes v i o ccur once and genes e i o ccur twice) . Ho w ev er, the num b er of o ccurrences of eac h gene in G 2 is up p er b ounded by k + 1. F urthermore, all genes ha ve p ositiv e sign, and h ence according to Observ ation 3 we only need to consider exemplarizations ( G E 1 , G E 2 ) of ( G 1 , G 2 ) suc h that G E 1 = G E 2 . It is imm ed iate to chec k th at our constru ction can b e carried out in p olynomial-time. W e no w claim that there exists a v ertex co ver of size k in G iff the exemplar br eakp oin t distance b et w een G 1 and G 2 is zero. Supp ose fi r st that there exists a vertex co ver V ′ ⊆ V of size k in G . W rite V ′ = { v i 1 , v i 2 , . . . , v i k } , i 1 < i 2 < . . . < i k . F or conv enience, we also d efine i 0 to b e 0. F r om V ′ w e construct an exemplar- ization ( G E 1 , G E 2 ) as follo ws. W e obtain G E 1 from G 1 b y a tw o step pro cedure. First we delete in G 1 all strings X i suc h that v i / ∈ V ′ . Second, for eac h 1 ≤ j ≤ m , if gene e j still o ccurs t wice, w e delete its second o ccurr ence (this second step is concerned with edges connecting t wo v ertices in V ′ ). W e no w turn to G E 2 . F or 1 ≤ j ≤ k , we consider the strin g Y [ j ] = Y V Y E that w e pro cess as follo ws: (1) we d elete in Y V all genes b ut v i j and th ose genes v ℓ / ∈ V ′ suc h that i j − 1 < ℓ < i j , and (2) w e delete in Y E all genes but those e ℓ that are n ot incident to v i j or incident to v i j and some smaller v ertex in V ′ ( i.e. , e ℓ = { v i j ′ , v i j } f or some j ′ < j ). Finally , w e d elete in the trailing string Y V = v 1 v 2 . . . v n all genes but those v ℓ ( / ∈ V ′ ) suc h that i k < ℓ . Since V ′ is a v ertex co v er in G , then it follo ws th at eac h gene o ccurs once in the obtained genomes, i.e. , ( G E 1 , G E 2 ) is ind eed an exemplarizatio n of ( G 1 , G 2 ). It is n o w easily seen th at G E 1 = G E 2 , and hence that the exemplar breakp oint distance b et w een G 1 and G 2 is zero. 14 Con v ersely , sup p ose that the exemplar breakp oin t distance b etw een G 1 and G 2 is zero. Since all genes ha v e a p ositiv e sign, then it follo ws that there exists an exemplarization ( G E 1 , G E 2 ) of ( G 1 , G 2 ) suc h th at G E 1 = G E 2 . Exemplarizatio n G E 2 can b e wr itten as G E 2 = Y V [1] Y E [1] Y V [2] Y E [2] . . . Y V [ k ] Y E [ k ] Y V [ k + 1] where, Y V [ i ], 1 ≤ i ≤ k + 1, is a string on V and Y E [ i ], 1 ≤ i ≤ k , is a str ing on E , V and E b eing viewe d as alphab ets. No w, defi ne V ′ ⊆ V as f ollo ws: v i ∈ V ′ iff gene v i o ccurs in some Y V [ j ], 1 ≤ j ≤ k , as the last gene. By construction, | V ′ | ≤ k (w e m a y ind eed hav e | V ′ | < k if some Y V [ j ], 1 ≤ j ≤ k , denotes the empt y str in g). W e no w observe that, sin ce no gene v i is du plicated in G 1 , all genes e ℓ that o ccur b et w een some gene v i ∈ V ′ and some gene v j ∈ V in G E 2 should matc h genes in str ing X i in G 1 . Then it follo ws that V ′ is a v ertex co ver of size at most k in G . ⊓ ⊔ The complexit y of ZEBD remains op en in case n o gene o ccur s more than twic e in G 1 and more than a constan t times in G 2 , i.e. , instances of t yp e (2 , c ) for some c = O (1) ; recall here th at ZEBD is NP –complete if no gene occur s more than th r ee times in G 1 or in G 2 (instances of t yp e (3 , 3), [13] ). In particular, the complexit y of ZEBD for instances of type (2 , 2) is op en. Ho wev er, w e p r op ose here a p ractical - but exp onen tial - algorithm f or ZEBD for instances of type (2 , 2), whic h is w ell-suited in case the n umber of genes that o ccur twice b oth in G 1 and in G 2 is relativ ely small. Prop osition 1. ZEBD for i nstanc es of typ e (2 , 2) (no gene o c curs mor e than twic e in G 1 and in G 2 ) is solvable in O ∗ (1 . 618 2 2 k ) time, wher e k is upp er-b ounde d by the numb er of genes that o c cu r exactly twic e in G 1 and in G 2 . Pr o of. According to Observ ation 3, for any instance ( G 1 , G 2 ), we only need to fo cu s on exemplar- izations ( G E 1 , G E 2 ) such that G E 1 = G E 2 or − ( G E 1 ) r = G E 2 , where − ( G E 1 ) r is the signed reve rsal of G E 1 . Let us first consider the case G E 1 = G E 2 (the case − ( G E 1 ) r = G E 2 is id en tical up to a signed rev ersal and will thereb y b e briefly d iscussed at the end of the p ro of ). Let ( G 1 , G 2 ) b e an instance of type (2 , 2) of ZEBD . Our algorithm is by transforming instance ( G 1 , G 2 ) in to a CNF b o olean formula φ with only few large clauses su c h that φ is satisfiable iff the exemplar breakp oint distance b et w een G 1 and G 2 is zero. By hyp othesis, eac h signed gene o ccurs at most t w ice in G 1 and in G 2 . Therefore, for an y signed gene g , we hav e one out of four p ossible distinct configurations depicted in Figure 2, where p 1 , p 2 , q 1 and q 2 are p ositions of o ccurrence of g in G 1 and G 2 . F urthermore, since w e are lo oking for an exemplarization ( G E 2 , G E 2 ) of ( G 1 , G 2 ) suc h that G E 1 = G E 2 , w e may assu me, in case g o ccurs only once in G 1 or in G 2 , that all o ccurren ces of G ha v e the same sign (otherwise a trivial self-reduction wo uld indeed apply). In other wo rds, referring at Figure 2, w e assume G 1 [ p 1 ] = G 2 [ q 1 ] = G 2 [ q 2 ] in case (2), G 1 [ p 1 ] = G 1 [ p 2 ] = G 2 [ q 1 ] in case (3), and G 1 [ p 1 ] = G 2 [ q 1 ] in case (4). Finally , as for case (1), w e ma y assume that either all o ccurrences ha v e the same sign, or G 1 [ p 1 ] = − G 1 [ p 2 ] and G 2 [ q 1 ] = − G 2 [ q 2 ] (otherwise a trivial self-reduction w ould again apply). W e n o w describ e the construction of the CNF b o olean formula φ . Firs t, th e set of b o olean v ariables X is defin ed as follo ws: for eac h gene g o ccur ring at p osition p in G 1 and at p osition q in G 2 ( i.e. , | G 1 [ p ] | = | G 2 [ q ]) | ) we add to X the b o olean v ariable x p q . W e n o w turn to d efining the clauses of φ . Let g b e an y gene, and let the o ccurrence p ositions of g in G 1 and in G 2 b e noted as in Figure 2. – if o cc( g , G 1 ) = occ( g , G 2 ) = 2 (case(1)), 15 (1) (2) (3) (4) G 1 G 2 q 2 q 1 p 1 p 2 q 2 p 1 q 1 q 1 p 2 p 1 q 1 p 1 Fig. 2. The 4 gene-configurations for instances of type (2 , 2): p 1 and p 2 are the o ccurrence p ositions of gene g in G 1 , and q 1 and q 2 are the occurrence positions of gene g in G 2 . – if G 1 [ p 1 ] = G 1 [ p 2 ] = G 2 [ q 1 ] = G 2 [ q 2 ], w e add to φ the clauses ( x p 1 q 1 ∨ x p 1 q 2 ∨ x p 2 q 1 ∨ x p 2 q 2 ), ( x p 1 q 1 ∨ x p 1 q 2 ), ( x p 1 q 1 ∨ x p 2 q 1 ), ( x p 1 q 1 ∨ x p 2 q 2 ), ( x p 1 q 2 ∨ x p 2 q 1 ), ( x p 1 q 2 ∨ x p 2 q 2 ) and ( x p 2 q 1 ∨ x p 2 q 2 ), – otherwise, w e ha ve G 1 [ p 1 ] = − G 1 [ p 2 ] and G 2 [ q 1 ] = − G 2 [ q 2 ] (see ab ov e discussion), – if G 1 [ p 1 ] = G 2 [ q 1 ] and G 1 [ p 2 ] = G 2 [ q 2 ])), we add to φ the clauses ( x p 1 q 1 ∨ x p 2 q 2 ) and ( x p 1 q 1 ∨ x p 2 q 2 ), – if G 1 [ p 1 ] = G 2 [ q 2 ] and G 1 [ p 2 ] = G 2 [ q 1 ])), we add to φ the clauses ( x p 1 q 2 ∨ x p 2 q 1 ) and ( x p 1 q 2 ∨ x p 2 q 1 ), – if o cc( g , G 1 ) = 1 and occ( g , G 2 ) = 2 (case (2)), we add to φ the clauses ( x p 1 q 1 ∨ x p 1 q 2 ) and ( x p 1 q 1 ∨ x p 1 q 2 ), – if o cc( g , G 1 ) = 2 and occ( g , G 2 ) = 1 (case (3)), we add to φ the clauses ( x p 1 q 1 ∨ x p 2 q 1 ) and ( x p 1 q 1 ∨ x p 2 q 1 ), and – if o cc( g , G 1 ) = occ( g , G 2 ) = 1 (case (4)), we add to φ the clause ( x p 1 q 1 ). The rationale of this construction is that if form ula φ ev aluates to true for some assignm ent f and f ( x p q ) is tru e for some gene g o ccurrin g at p osition p in G 1 and q in G 2 , then all o ccurrences of g but the one at p osition p should b e deleted in G 1 and all o ccurrences of g bu t the one at p osition q should b e deleted in G 2 , in order to obtain the exemplar solution. What is left is to enforce that φ ev aluates to true iff the exemplar breakp oint distance b et w een G 1 and G 2 is zero. T o this aim, w e add to φ the follo wing clauses. F or eac h pair of v ariables ( x i 1 j 1 , x i 2 j 2 ) suc h that | G 1 [ i 1 ] | 6 = | G 1 [ i 2 ] | , i 1 < i 2 and j 1 > j 2 , we add to φ the clause ( x i 1 j 1 ∨ x i 2 j 2 ). T he constru ction of φ is now complete. Clearly , φ ev aluates to tru e iff the exemplar b reakp oint distance b et w een G 1 and G 2 is zero. Let k b e the num b er of genes g that occur twice in G 1 and in G 2 with the same sign, i.e. , G 1 [ p 1 ] = G 1 [ p 2 ] = G 2 [ q 1 ] = G 2 [ q 2 ]. W e no w m ak e the imp ortant observ ation th at all clauses in φ ha v e size less than or equal to 2 except those k clauses of size 4 in tro duced in case gene g o ccurs t wice in G 1 and in G 2 with th e same sign. By introd ucing a new b o olean v ariable, we can easily replace in φ eac h clause of size 4 by t wo clauses of size 3, and hence we m a y no w assume that φ is a 3-CNF form ula ( i.e. , eac h clause has size at most 3) with exactly 2 k clauses of size 3. As for the case − ( G E 1 ) r = G E 2 , w e replace G 1 b y − ( G 1 ) r and construct another 3-CNF formula φ ′ as describ ed ab o ve. The t wo 3-CNF formulas need, ho wev er, to b e examined separately . F ernau prop osed in [15] an algorithm for solving 3-CNF b o olean form ulas that ru ns in O ∗ (1 . 618 2 ℓ ) time, where ℓ is the n umber of clauses of size 3. Therefore, ZEBD for instances of type (2 , 2) is solv able in O ∗ (1 . 618 2 2 k ) time, where k is the num b er of genes g that o ccur twice in G 1 and in G 2 . ⊓ ⊔ 16 4.2 Zero in termediate matching breakp oin t dista nce W e no w turn to the zero in termediate br eakp oin t distance ( ZIBD ) problem. It is defi n ed as follo ws. Problem: ZI BD Input: Tw o genomes G 1 and G 2 . Question: I s the in termediate breakp oin t distance b et w een G 1 and G 2 equal to zero ? W e sho w h ere that ZEBD and ZIBD are equiv alent p roblems. W e need the f ollo wing lemma. Lemma 9 ([2]). L e t G 1 and G 2 b e two genomes without duplic ates and with the same gene c on- tent, and G ′ 1 and G ′ 2 b e the two ge nomes obtaine d fr om G 1 and G 2 by deleting any ge ne g . Then B ( G ′ 1 , G ′ 2 ) ≤ B ( G 1 , G 2 ) . Theorem 5. ZEBD and ZIBD ar e e qui v alent pr oblems. Pr o of. One direction is trivial (an y exemplarizatio n is ind eed an inte rmediate matc hing). The other direction follo ws from Lemma 9. ⊓ ⊔ It follo ws f rom Theorem 5 that the problem IBD is not appr o xim ab le ev en for instances of t yp e (3 , 3) (see [13]) and if no gene o ccurs more than t wice in G 1 (see Th eorem 4). 4.3 Zero maxim um matc hing breakp oin t distance W e sho w here that, opp ositely to the exemplar and the in termediate matc hing mo dels, deciding whether the maxim um matc h ing breakp oin t distance b et w een t w o genomes is equal to zero is p olynomial-time solv able, and hence we cannot rule out the existence of accurate app ro ximation algorithms f or the maxim um matc hing m o del. W e refer to this problem as ZMBD . Problem: ZM BD Input: Tw o genomes G 1 and G 2 . Question: Is the maxim um matc hing breakp oint distance b et ween G 1 and G 2 equal to zero ? The main idea of our approac h is to transf orm an y instance of ZMBD into a matching diagr am and next use an efficient algorithm for finding a large set of non-intersecti ng line segmen ts. Note that th is latter problem is equiv alent to findin g a large increasing subsequence in p erm utations. A matc hing d iagram [18] consists of, sa y n , p oin ts on eac h of t wo parallel lines, and n str aight line segmen ts matching distinct pairs of p oint s. The in tersection graph of the line segment s is called a p ermutation gr aph (the reason f or the name is that if the p oin ts on the top line are num b ered 1 , 2 , . . . , n , then the p oin ts on the other line are num b ered by a p ermutatio n on 1 , 2 , . . . , n ). W e describ e h o w to turn the pair of genomes ( G 1 , G 2 ) into a matc hing diagram D ( G 1 , G 2 ). F or sake of present ation we introdu ce the f ollo wing n otations. F or eac h gene family g , we write o cc pos ( G, g ) (resp. o cc neg ( G, g )) for the num b er of p ositiv e (resp. n egativ e) o ccurrences of gene g in genome G . According to Observ ation 3, it is enough to consider t wo cases: G M 1 = G M 2 or − ( G M 1 ) r = G M 2 , wh ere ( G M 1 , G M 2 , M ) is a maximum matc hing of ( G 1 , G 2 ). Let us first fo cus on testing G M 1 = G M 2 (the case − ( G M 1 ) r = G M 2 is iden tical up to a signed rev ersal). W e describ e the construction of th e top lab eled p oin ts. Reading genome G 1 from left to righ t, we rep lace gene g by the sequence of lab eled p oin ts + g 1 ( i, o cc pos ( G 2 , g )) + g 1 ( i, o cc pos ( G 2 , g ) − 1) . . . + g 1 ( i, 1) 17 if g is the i -th p ositiv e o ccurrence of gene g in genome G 1 or by the sequence of lab eled p oints − g 1 ( i, o cc neg ( G 2 , g )) − g 1 ( i, o cc neg ( G 2 , g ) − 1) . . . − g 1 ( i, 1) if g is the i -th negativ e o ccurrence of gene g in genome G 1 . A symm etric constru ction is p erformed for the lab eled p oint s of the b ottom line, i.e. , reading genome G 2 from left to righ t, we replace gene g b y the sequence of lab eled p oin ts + g 2 ( i, o cc pos ( G 1 , g )) + g 2 ( i, o cc pos ( G 1 , g ) − 1) . . . + g 2 ( i, 1) if g is the i -th p ositiv e o ccurrence of gene g in genome G 2 or by the sequence of lab eled p oints − g 2 ( i, o cc neg ( G 1 , g )) − g 2 ( i, o cc neg ( G 1 , g ) − 1) . . . − g 2 ( i, 1) if g is the i -th negativ e o ccurrence of gene g in genome G 2 . W e no w obtain the matc hing diagram D ( G 1 , G 2 ) as follo ws: eac h lab eled p oint + g 1 ( i, j ) (resp . − g 1 ( i, j )) of the top lin e is connecte d to the lab eled p oint + g 2 ( j, i ) (resp. − g 2 ( j, i )) of the b ottom line b y a line segmen t. Clearly , eac h lab eled p oint is in ciden t to exactly one line segmen t, and hen ce D ( G 1 , G 2 ) is indeed a matc hing diagram. Of particular imp ortance, observ e that b y construction, for an y x ∈ { 1 , 2 } and any t wo lab eled p oints + g x ( i, j ) and + g x ( i, k ), j 6 = k , th e t w o line segments in ciden t to these tw o p oin ts are in tersecting ; the s ame conclusion can b e d ra wn for any tw o lab eled p oin ts − g x ( i, j ) and − g x ( i, k ), j 6 = k . The follo wing lemma states this prop ert y in a suitable wa y . Lemma 10. If [+ g 1 ( i, j ) , + g 2 ( j, i )] and [+ g 1 ( k , ℓ ) , + g 2 ( ℓ, k )] (r esp. [ − g 1 ( i, j ) , − g 2 ( j, i )] and [ − g 1 ( k , ℓ ) , − g 2 ( ℓ, k )] ) ar e two non-i nterse cting line se gments in the matching diagr am D ( G 1 , G 2 ) , then i 6 = k and j 6 = ℓ . Theorem 6. ZMBD is p olynomial-time solvable. Pr o of. Let G 1 and G 2 b e tw o genomes, and m the size of a maxim um matc hin g b et w een G 1 and G 2 . According to Lemma 10, th ere exists a maxim um matc hin g ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) su c h th at G M 1 = G M 2 if there exists m non-intersecting line segmen ts in D ( G 1 , G 2 ). The maxim um num b er of non-in tersecting line segmen ts in a matc h ing diagram with n p oin ts on eac h line can b e f ou n d in O ( n log log n ) time [8]. As for the case − ( G M 1 ) r = G M 2 , we replace G 1 b y − ( G 1 ) r and r un the same algorithm on the obtained matc hing diagram. ⊓ ⊔ 5 Appro ximating the n um b er of adjacencies in the maxim um matc hing mo del F or t w o balanced genomes G 1 and G 2 , sev eral approximat ion algorithms for computing th e n u m b er of br e akp oints b et ween G 1 and G 2 are given for th e maxim um m atc hin g mo d el [17, 19]. W e prop ose in this section three appro ximation algorithms to maximize the n u m b er of adjac encies (as opp osed to minimizing the num b er of breakp oin ts). The appro ximation r atios we obtain are 1 . 1442 when o cc( G 1 ) = 2, 3 wh en o cc( G 1 ) = 3 and 4 in the general case. Ob serv e th at in the latter case, opp ositely to [17, 19], our approxima tion ratio is indep endent of the maximum num b er of dup licates. Note also that in [12], inapp ro ximation results are given for t wo unb alanc e d genomes G 1 and G 2 ev en wh en o cc( G 1 ) = 1 and o cc( G 1 ) = 2. W e first defin e the problem M ax- k -Adj w e are inte rested in ( k ≥ 1 is a fixed in teger). 18 Problem: Ma x- k -Adj Input: Two balanced genomes G 1 and G 2 with o cc( G 1 ) = k (and consequen tly o cc( G 2 ) = k ). Solution: A maxim um matc hing ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ). Measure: Th e num b er of adjacencies b et w een G M 1 and G M 2 . W e define Max-Ad j to b e the pr oblem MAX k -Adj , in wh ic h k is unb ounded . 5.1 A 1 . 1442-appro ximat ion for Max-2-Adj W e f o cus here on balanced genomes G 1 and G 2 suc h th at o cc( G 1 ) = 2, and w e giv e an appro ximation algorithm for Max -2-Adj based on the Max-2-CSP problem (defined b elo w ), for w hic h a 1 . 1442- appro ximation algorithm is giv en in [9]. T he main idea is to constru ct a b o olean formula ϕ for eac h p ossible adjacency , and next to maximize the n umber of b o olean formulas φ that can b e sim ultaneously satisfied in a truth assignment ; the num b er of simultaneously satisfied f ormulas will b e exactly the n umber of adjacencies, an d hence any approxi mation ratio for Max-2-CSP is an app ro ximation ratio for Max-2-Adj . Problem: Ma x- k -CSP Input: A p air ( χ, Φ ), w here χ is a set of b o olean v ariables and Φ is a set of b o olean form ulas su c h that eac h form ula conta ins at most k literals of χ . Solution: An assignment of χ . Measure: Th e num b er of formulas that are satisfied b y the assignmen t. W e define the follo win g tr ansformation M ak eCSP that asso ciates to any instance of Max-2-Adj an instance of M ax-2-CSP . Give n an instance ( G 1 , G 2 ) of Max-2-Adj , w e create a v ariable X g for eac h gene g and define χ as the set of v ariables X g . Th en, we construct the set Φ of formula s. F or eac h duo d i = ( G 1 [ i ] , G 1 [ i + 1]), 1 ≤ i ≤ n G 1 − 1, suc h that d i or − d i app ears in G 2 , w e distinguish three cases in order to create a formula ϕ i of Φ : 1. There exists a unique duo d j = ( G 2 [ j ] , G 2 [ j + 1]) in G 2 suc h that d j = d i or d j = − d i . F or sake of readabilit y , w e define the literal Y q p , 1 ≤ p ≤ n G 1 , 1 ≤ q ≤ n G 2 , where | G 1 [ p ] | = | G 2 [ q ] | , as follo ws: Y q p = X | G 1 [ p ] | if N G 1 [ p ] = N G 2 [ q ] and Y q p = X | G 1 [ p ] | otherwise. W e now consider t wo cases: – (a) d i = d j : in that case, ϕ i = ( Y j i ∧ Y j +1 i +1 ). – (b) d i = − d j : in that case, ϕ i = ( Y j +1 i ∧ Y j i +1 ). 2. The duo d i app ears twice in G 2 . W e consider t wo cases: – (c) N G 1 [ i ] = N G 1 [ i + 1]: in that case, ϕ i = ( X | G 1 [ i ] | ⊕ X | G 1 [ i +1] | ) wh ere ⊕ is the b o olean function X O R . – (d) N G 1 [ i ] 6 = N G 1 [ i + 1]: in th at case, ϕ i = ( X | G 1 [ i ] | ⊕ X | G 1 [ i +1] | ). Remark that eac h formula ϕ i con tains t w o literals. Hence, ( χ, Φ ) is an instance of Max-2-CSP . Lemma 11. L et G 1 and G 2 b e two b alanc e d genomes such that o cc( G 1 ) = 2 . L et ( χ, Φ ) b e the instanc e of Max-2-CSP obtaine d by Make CSP ( G 1 , G 2 ) . F or any inte ger k , i f ther e exists a max- imum matching ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) which induc es at le ast k adjac e nc i es, then ther e exists an assignment of the v ariables of χ such that at le ast k formulas of Φ ar e satisfie d. 19 Pr o of. Let G 1 and G 2 b e t w o balanced genomes su c h that o cc( G 1 ) = 2 and let ( χ, Φ ) b e the instance of Max-2-CSP obtained by MakeCSP ( G 1 , G 2 ). Let k b e an int eger. Supp ose there exists a m axim um matc hin g ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) whic h ind uces at least k adjacencies. W e construct the follo w ing assignment of v ariables of χ . F or eac h gene g , we define X g = 1 if g is not dup licated, else we defin e X g = 1 iff the o ccurr ences of g are matc hed in the reading order (see Figure 3). W e now show that for eac h duo wh ich induces an adjacency b et w een G M 1 and G M 2 , there exists a distinct satisfied form ula of Φ . Let d i = ( G M 1 [ i ] , G M 1 [ i + 1]), 1 ≤ i ≤ n G 1 − 1, b e a duo whic h in duces an adjacency , and let d j = ( G M 2 [ j ] , G M 2 [ j + 1]) b e the related duo on G M 2 . By constru ction of Φ , there exists a f orm ula ϕ i ∈ Φ whic h has b een previously defined in one of the cases (a), (b), (c) or (d) of the definition of Make CSP . W e claim that, for eac h of these cases, ϕ i is satisfied: – (a) ϕ i = ( Y j i ∧ Y j +1 i +1 ) and d i = d j . W e first pro v e that literal Y j i is true. Three cases are p ossible. (i) The gene | G 1 [ i ] | is not du plicated ; then we ha v e defin ed in our assignmen t X | G 1 [ i ] | = 1. Moreo ver, w e ha v e Y j i = X | G 1 [ i ] | (since N G 1 [ i ] = N G 2 [ j ] = 0), hence Y j i is true. (ii) T he gene | G 1 [ i ] | is duplicated and N G 1 [ i ] = N G 2 [ j ] ; then, by definition of our assignment and since G 1 [ i ] and G 2 [ j ] are matc hed together in the maxim um matc hing ( G M 1 , G M 2 , M ), w e hav e X | G 1 [ i ] | = 1 (w e matc h signed genes in the reading order). Moreo ver, w e ha v e Y j i = X | G 1 [ i ] | whic h induces that Y j i is true. (iii) The gene | G 1 [ i ] | is d uplicated and N G 1 [ i ] 6 = N G 2 [ j ] ; then, by d efi nition of our assignment and since G 1 [ i ] and G 2 [ j ] are matc hed toge ther in the maximum matc h ing ( G M 1 , G M 2 , M ), we ha v e X | G 1 [ i ] | = 0 (w e do not matc h signed genes in the reading order). Moreo ver, w e h a ve in this case Y j i = X | G 1 [ i ] | whic h in duces that Y j i is true. In eac h case, w e ha v e p ro v ed that Y j i is true. W e can also p ro v e that Y j +1 i +1 is true, u sing the same argumen ts. Hence, w e conclude that ϕ i is tru e. – (b) ϕ i = Y j +1 i ∧ Y j i +1 and d i = − d j . By similar arguments as in case (a), we can prov e that Y j +1 i and Y j i +1 are true. – (c) W e hav e N G 1 [ i ] = N G 1 [ i + 1] and the d uo d i app ears twice in G 2 (noted d j and d j ′ ). Since d i induces an adj acency , the d uo d i matc hes either d j or d j ′ . In these t w o cases, w e h a v e X | G 1 [ i ] | = X | G 1 [ i +1] | (otherwise G 1 [ i ] and G 1 [ i + 1] would not matc h su ccessiv e signed genes). Moreo ver, ϕ i = ( X | G 1 [ i ] | ⊕ X | G 1 [ i +1] | ) and thus, ϕ i is tru e. – (d) W e ha v e N G 1 [ i ] 6 = N G 1 [ i + 1] and the d uo d i app ears t wice in G 2 (noted d j and d j ′ ). Since d i induces an adj acency , the d uo d i matc hes either d j or d j ′ . In these t w o cases, w e h a v e X | G 1 [ i ] | 6 = X | G 1 [ i +1] | (otherwise G 1 [ i ] and G 1 [ i + 1] would not matc h su ccessiv e signed genes). Moreo ver, ϕ i = ( X | G 1 [ i ] | ⊕ X | G 1 [ i +1] | ) and thus, ϕ i is tru e. W e ha v e constru cted a v ariable assignmen t of χ su c h that, for eac h du o d i in G M 1 whic h im p lies an adj acency , there exists a distinct satisfied formula ϕ i ∈ Φ . Thus, if there exists a m axim um matc hin g of ( G 1 , G 2 ) wh ic h induces at least k adjacencies, then the corresp onding assignment implies at least k satisfied form ulas. ⊓ ⊔ Lemma 12. L et G 1 and G 2 b e two b alanc e d genomes such that o cc( G 1 ) = 2 . L et ( χ, Φ ) b e the instanc e of Max-2-CSP obtaine d by Mak eCSP ( G 1 , G 2 ) . F or any inte ger k , if ther e exists an as- signment of χ su ch that at le ast k formulas of Φ ar e satisfie d, then ther e exists a maximum matching ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) which induc e s at le ast k adjac e nc i es. 20 Fig. 3. All possibilities of assignmen t: X A = 1 (gene A o ccurs tw ice and signed genes are m atched in the reading order), X B = 1 or X B = 0 (gene B occurs once) and X C = 0 (gene C o ccurs t wice and signed genes are not matched in the reading order). Note that this construction is indepen dent of the sign of the genes. Pr o of. Let G 1 and G 2 b e t w o balanced genomes su c h that o cc( G 1 ) = 2 and let ( χ, Φ ) b e the instance of Max-2-CSP obtained by MakeCSP ( G 1 , G 2 ). Let k b e an int eger. Supp ose there exists an assignmen t of χ suc h th at at least k form ulas ϕ i ∈ Φ are satisfied. W e create th e follo wing maxim um matc hing ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ). F or eac h v ariable X g suc h th at the gene g is d uplicated, w e matc h th e o ccurrences of g in the reading order if X g = 1 (suc h as gene A in Figure 3). I f w e hav e X g = 0, we matc h the first o ccurren ce of g on G 1 with the second one on G 2 and the second o ccurrence of g on G 1 with the first one on G 2 (suc h as gene C in Figure 3). Then, w e matc h signed genes w hic h are not duplicated. Now, we pro v e that eac h satisfied form u la ϕ i ∈ Φ in d uces a distinct adjacency for ( G M 1 , G M 2 , M ). Let ϕ i ∈ Φ b e a satisfied formula whic h is defined in one of the cases (a), (b), (c) or (d) of th e definition of Mak eCSP : – (a) W e h av e ϕ i = ( Y j i ∧ Y j +1 i +1 ) and the duos d i = ( G 1 [ i ] , G 1 [ i + 1]) and d j = ( G 2 [ j ] , G 2 [ j + 1]) are iden tical. Here, we must pro v e that d i and d j are matc h ed together in ( G M 1 , G M 2 , M ) and thus induce an ad- jacency . First, we sho w that signed genes G 1 [ i ] and G 2 [ j ] are matc hed together in ( G M 1 , G M 2 , M ). Since ϕ i is satisfied, we ha v e Y j i = 1. W e must disso ciate thr ee cases: (i) the gene | G 1 [ i ] | is not duplicated: in that case, the signed gene G 1 [ i ] can b e matc hed only with G 2 [ j ]. (ii) T h e gene | G 1 [ i ] | is du plicated and w e ha v e N G 1 [ i ] = N G 2 [ j ]. In that case, we ha v e d efined Y j i = X | G 1 [ i ] | whic h implies X | G 1 [ i ] | = 1. Thus, since N G 1 [ i ] = N G 2 [ j ], the signed genes G 1 [ i ] and G 2 [ j ] are matc hed together. ( iii) The gene | G 1 [ i ] | is d uplicated and we h a v e N G 1 [ i ] 6 = N G 2 [ j ]. In th at case, w e hav e defined Y j i = X | G 1 [ i ] | whic h im p lies X | G 1 [ i ] | = 0. Thus, since N G 1 [ i ] 6 = N G 2 [ j ], the signed genes G 1 [ i ] and G 2 [ j ] are matc h ed toge ther. F or eac h case, the signed genes G 1 [ i ] and G 2 [ j ] are matc h ed together. W e can conclude in the same w a y that G 1 [ i + 1] and G 2 [ j + 1] are also matc hed together, whic h implies that d i induces an adjacency . – (b) W e hav e ϕ i = ( Y j +1 i ∧ Y j i +1 ) = 1 and the duos d i = ( G 1 [ i ] , G 1 [ i + 1]) and d j = ( G 2 [ j ] , G 2 [ j + 1]) are rev ersed. W e can use the same reasoning used in case (a) to pro v e th at d i induces an adjacency . – (c) The d uo d i app ears twice in G 2 (noted d j and d j ′ ). W e ha v e ϕ i = ( X | G 1 [ i ] | ⊕ X | G 1 [ i +1] | ) and N G 1 [ i ] = N G 1 [ i + 1]. Since ϕ i is true, we h a ve X | G 1 [ i ] | = X | G 1 [ i +1] | whic h implies by construction of the m axim um matc hin g that d i matc hes d j or d j ′ . – (d) The duo d i app ears twice in G 2 (noted d j and d j ′ ). W e h a v e ϕ i = ( X | G 1 [ i ] | ⊕ X | G 1 [ i +1] | ) and N G 1 [ i ] 6 = N G 1 [ i + 1 ]. S in ce ϕ i is tru e, w e h a v e X | G 1 [ i ] | 6 = X | G 1 [ i +1] | whic h im p lies by construction of the maxim um matc hing that d i matc hes d j or d j ′ . 21 Consequent ly , for eac h satisfied formula, there exists a distinct adjacency b et w een G M 1 and G M 2 . Th us, if there exists an assignmen t of χ whic h implies at least k satisfied form u las of Φ , then there exists a maximum matc h ing of ( G 1 , G 2 ) wh ich implies at least k ad j acencies. ⊓ ⊔ Lemmas 11 and 12 pro v e that an y α -appr o ximation for Max-2-CSP implies an α -appro ximation for M ax-2-Adj . In [9], an appro ximation alg orithm is giv en for Ma x-2-CSP , whose appro ximation ratio is equal to 1 0 . 874 ≤ 1 . 144 2. Thus, w e ha ve the follo win g th eorem. Theorem 7. Max-2-Adj is 1 . 1442 -appr oximable. 5.2 A 3-appro ximation for Max-3-Adj No w, we present a 3-approxi mation for Max-3-Adj by using the Maximum I ndepend ent Set problem defined as follo ws: Problem: Ma x-Indepen dent-Set Input: A graph G = ( V , E ). Solution: An in dep end en t set of G (i.e. a subset V ′ of V suc h that no t wo v ertices in V ′ are joined by an edge in E ). Measure: Th e cardinalit y of V ′ . In [17] , Goldstein et al . used Max-Inde penden t-Set to appr o x im ate the Minim um Comm on String P artition pr oblem b y creating a c onflict gr aph . W e construct in the same wa y an in s tance of Ma x-Independ ent-Set wh ere a v ertex represents a p ossible adj acency and where an edge represent s a conflict b etw een t w o adjacencies. W e d efine M ak eMIS to b e the follo wing transformation whic h asso ciates to t w o balanced genomes G 1 and G 2 an instance of Max-In depende nt-Set . W e construct a v ertex for eac h duo matc h, and then we create an edge b et w een tw o vertic es when they are in confl ict, i.e. when t wo matc hes are incompatible. Figure 4 illustrates the graph obtained b y Mak eMIS ( G 1 , G 2 ) where G 1 = +3 + 1 + 2 + 3 + 4 + 2 + 5 and G 2 = +3 + 4 + 2 + 3 + 1 + 2 + 5. Fig. 4. The conflict graph obtained by MakeMIS ( G 1 , G 2 ) where G 1 = +3 + 1 + 2 + 3 + 4 + 2 + 5 and G 2 = +3 + 4 + 2 + 3 + 1 + 2 + 5 (for sake of readabilit y , p ositiv e signs are not display ed). In order to prov e that ther e exists a 3-appro xim ation for Max-3-Adj , we give the follo wing in termediate lemmas. 22 Lemma 13. L et G 1 and G 2 b e two b alanc e d genomes and let G b e the gr aph obtaine d by Mak eMIS ( G 1 , G 2 ) . F or any inte ger k , ther e exists an indep endent set V ′ of G such that | V ′ | ≥ k iff ther e exists a maximum matching ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) which induc e s at le ast k adjac e nc i es. Pr o of. Let G 1 and G 2 b e tw o b alanced genomes an d let G b e the graph obtained by Mak eMIS ( G 1 , G 2 ). L et k b e an in teger. ( ⇒ ) Su pp ose there exists an ind ep endent set V ′ of G su c h that | V ′ | ≥ k . W e construct a matc hin g ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) as follo ws: first, for eac h v ertex of V ′ , w e matc h together the t wo corresp ond ing d uos, th us indu cing one adjacency (calle d a definite adjacency). By construction of G , this op eration is p ossible. I ndeed, tw o v ertices wh ic h are not connected in G imply t w o compatible adj acencies. Then, w e matc h arbitrarily the un m atc hed genes. This op eration cannot break an y definite adjacency . Finally , w e obtain a maxim um matc hing ( G M 1 , G M 2 , M ) which ind uces at least | V ′ | adjacencies, and consequen tly at least k adjacencies. ( ⇐ ) Supp ose there exists a maxim u m matc hin g ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) whic h induces at least k adjacencies. W e construct a set V ′ b y taking eac h v ertex whic h r epresen ts a duo matc h b et w een G M 1 and G M 2 . By construction of G , V ′ is an indep endent set (no pair of adjacencies can create a conflict), and then w e ha ve | V ′ | ≥ k . ⊓ ⊔ Lemma 14. L et G 1 and G 2 b e two b alanc e d g enomes such that occ ( G 1 ) = k . The maximum de g r e e ∆ of the gr aph G obtaine d by Mak eMIS ( G 1 , G 2 ) satisfies ∆ ≤ 6( k − 1) . Pr o of. Let G 1 and G 2 b e t wo balanced genomes suc h that occ( G 1 ) = k and let G b e the graph obtained b y MakeMIS ( G 1 , G 2 ). Consider a d u o matc h m = ( d 1 , d 2 ) with d 1 = ( G 1 [ i ] , G 1 [ i + 1]) and d 2 = ( G 2 [ j ] , G 2 [ j + 1]) where 1 ≤ i ≤ n G 1 − 1 and 1 ≤ j ≤ n G 2 − 1. W e claim that the vertex v m of G , whic h represents the d uo matc h m , is connected to at most 6( k − 1) v ertices. F or this, we list the p ossible d u o matc hes m ′ = ( d ′ 1 , d ′ 2 ) suc h that the v ertex v m ′ of G wh ic h represen ts m ′ is connected to v m . Remark that if v m ′ is connected to v m (i.e. m and m ′ are in conflict), then at least one of the duos d ′ 1 and d ′ 2 o verlaps, resp ectiv ely , either d 1 or d 2 . Let d ′ 1 b e a du o in G 1 whic h o v erlaps d 1 . First, w e list the p ossible d u os d ′ 2 suc h that the du o matc h es m = ( d 1 , d 2 ) and m ′ = ( d ′ 1 , d ′ 2 ) are in conflict. Remark that d ′ 1 (or − d ′ 1 ) app ears at most k times on G 2 since a gene can o ccur at most k times. W e th en distinguish three cases: – (a) d ′ 1 = ( G 1 [ i − 1] , G 1 [ i ]): if d ′ 1 (or − d ′ 1 ) app ears k times in G 2 , one of these o ccurr ences is necessary d ′ 2 = ( G 2 [ j − 1] , G 2 [ j ]) if d 1 = d 2 , or d ′ 2 = ( G 2 [ j + 1] , G 2 [ j + 2]) if d 1 = − d 2 . F or these t wo cases, the duo matc hes m and ( d ′ 1 , d ′ 2 ) are not in conflict. – (b) d ′ 1 = d 1 : if d ′ 1 (or − d ′ 1 ) app ears k times on G 2 , one of these occurr ences is necessary d 2 , whic h in duces in this case n o conflict with m . – (c) d ′ 1 = ( G 1 [ i + 1] , G 1 [ i + 2]): if d ′ 1 (or − d ′ 1 ) app ears k times on G 2 , one of these o ccurrences is necessary d ′ 2 = ( G 2 [ j + 1] , G 2 [ j + 2]) if d 1 = d 2 , or d ′ 2 = ( G 2 [ j − 1] , G 2 [ j ]) if d 1 = − d 2 . F or these t wo cases, the duo matc hes m and m ′ are not in conflict. F or eac h case, one of the k p ossible duos d ′ 2 do es not imply a conflict b et w een m and m ′ . Thus, for an y duo d ′ 1 whic h o verlaps d 1 , there exists at most k − 1 duos d ′ 2 on G 2 suc h that m and m ′ are in conflict. Using the same arguments, we can easily p ro v e that for an y duo d ′ 2 whic h o verlaps d 2 , there exists at most k − 1 duos d ′ 1 on G 1 suc h that m and m ′ are in conflict. Hence, eac h of th e six duos which o v erlaps d 1 or d 2 implies at most k − 1 confl icts. Thus, we obtain at m ost 6( k − 1) v ertices whic h are connected to the v ertex v m in th e conflict graph. ⊓ ⊔ 23 According to Lemma 13, any α -app ro ximation for Ma x-Indepen dent-Set is thus also an α -appro ximation f or Max- k -Adj . It is prov ed in [5] that Max-Inde penden t-Set that is ap- pro ximable within ratio ∆ +3 5 , where ∆ is the maximum degree of the graph. Com bining this with Lemma 14, we obtain the follo w ing result. Theorem 8. Max- k -Adj is 6 k − 3 5 -appr oximable. Note that in the case wh ere k = 2, we obtain a ratio of 1 . 8, w hic h is not b etter than the one obtained in Theorem 7. Moreo v er, we in tro duce in the next sectio n a 4-appr oximati on in the general case. Hence, th e only interesting case of Th eorem 8 ab o ve is w hen k = 3, ind ucing a 3-approxi mation for Ma x-3-Adj . 5.3 A 4-appro ximation for Max-Adj In [14], a 4-approxima tion algorithm for the M ax-Weighted 2-inter v al P a ttern pr oblem ( Max-W2IP ) is giv en. In th e follo wing, we first defin e Max-W 2IP , and next w e p r esen t how w e can relate an y instance of Max-Adj to an instance of Max-W2IP . The Maximum W e ighte d 2 -Interval Pattern pr oblem. A 2- interval is the u nion of t w o disj oin t in terv als defined o v er a sin gle line. F or a 2-in terv al D = ( I , J ), we alw ays assume that th e in terv al I < J , i.e. , I is completely on the left of J do es not ov erlap J . W e sa y that t w o 2-inte rv als D 1 = ( I 1 , J 1 ) and D 2 = ( I 2 , J 2 ) are disjoint if D 1 and D 2 ha v e no common p oin t (i.e. ( I 1 ∪ J 1 ) ∩ ( I 2 ∪ J 2 ) = ∅ ). Three p ossible relations exist b et ween t wo disjoin t 2-in terv als: we wr ite (1) D 1 ≺ D 2 , if I 1 < J 1 < I 2 < J 2 , (2) D 1 ⊏ D 2 , if I 2 < I 1 < J 1 < J 2 and (3) D 1 ≬ D 2 , if I 1 < I 2 < J 1 < J 2 . W e sa y that a pair of 2-in terv als D 1 and D 2 is R - compar able for some R ∈ {≺ , ⊏ , ≬ } , if either ( D 1 , D 2 ) ∈ R or ( D 2 , D 1 ) ∈ R . A set of 2-in terv als D is R -comparable for some R ⊆ {≺ , ⊏ , ≬ } , R 6 = ∅ , if an y pair of distinct 2-in terv als in D is R -comparable for some R ∈ R . The non-empty set R is called a mo del . Th e Max -Weighted 2-i nter v al P a ttern ( Max-W2IP ) pr oblem is formally defined as follo ws. Problem: Ma x-Weighted 2-inter v al P a ttern ( Max-W2IP ) Input: A set D of 2-in terv als, a mo d el R ⊆ {≺ , ⊏ , ≬ } with R 6 = ∅ , and a weig ht function ω : D → N + . Solution: An R -comparable subset D ′ of D . Measure: Th e weig h t of D ′ . T r ansformation. W e fir st d escrib e h o w to trans form an y in s tance ( G 1 , G 2 ) of Max-Adj into an instance, referred hereafter as M ak e2I ( G 1 , G 2 ) = ( D , R , ω ), of M ax-W2IP . W e need a new defin i- tion. Let G 1 and G 2 b e t w o b alanced genomes. An interv al I 1 of G 1 and an interv al I 2 of G 2 , b oth of size at least 2, are said to b e identic al if they corresp ond to the same string up to a complete rev ersal, where a reve rsal also c hanges all the s igns in the string. C learly , t wo iden tical in terv als ha v e the same length. The weigh ted 2-inte rv al set D is obtained as follo ws . W e first concatenate G 1 and G 2 , an d for any p air ( I 1 , I 2 ) of identica l in terv als ( I 1 is an interv al of G 1 and I 2 is an interv al of G 2 ), w e construct the 2-inte rv al D = ( I 1 , I 2 ) of w eigh t ω ( D ) = | I 1 | − 1 (= | I 2 | − 1) and add it to D . Notice that, since iden tical in terv als ha ve length at least 2, eac h 2-in terv al of D has w eigh t at least 1. Figure 5 giv es an example of suc h a construction. Observ e that, b y constru ction, n o t w o 2-in terv als 24 of D are {≺} -comparable. The construction of the instance of Max-W2IP is complete b y setting R = {≺ , ⊏ , ≬ } , i. e. , w e are lo oking for disjoint 2-inte rv als, no matter what the relation b etw een an y t w o disjoint 2-in terv al is. Therefore, for sak e of abb reviation, we sh all d enote the corresp ond in g instance simply as Mak e2I ( G 1 , G 2 ) = ( D , ω ) and forget ab out the mo d el. Fig. 5. 2-interv als induced by genomes G 1 = +1 + 2 − 3 + 2 + 1 and G 2 = +2 + 1 + 3 − 2 − 1. F or readability , singleton interv als are not dra wn. The dotted 2-interv al is of weigh t 2, while all other 2-interv als are of w eigh t 1. W e no w describ e h o w to transf orm an y solution of Max-W2IP int o a solution of Ma x-Adj . Let G 1 and G 2 b e t wo balanced genomes and Make2 I ( G 1 , G 2 ) = ( D , ω ). F u rthermore, let S ⊆ D b e a set of disj oint 2-in terv als, i.e. a solution for Max -W2IP for mo del the {≺ , ⊏ , ≬ } for the instance ( D , ω ). W e w rite Max-W2IP to Adj ( S ) for the transform ation of S into a maxim um matc hing ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) defined as follo ws. First, for eac h 2-interv al D = ( I 1 , I 2 ) of S , w e match the signed genes of I 1 and I 2 in the natural wa y ; th en , in order to ac hiev e a maximum matc hin g (since eac h signed gene is not necessarily co v ered by a 2-in terv al in S ), w e apply the follo wing greedy algorithm: iterativ ely , w e matc h, arbitrarily , t w o un matc h ed signed genes g 1 and g 2 suc h that | g 1 | = | g 2 | and g i is a gene of G i ( i = 1 , 2), until no suc h p air of signed genes exists. After a r elab eling of signed genes according to this matc hin g (denoted M ), w e obtain a maxim um m atching ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ). The r ationale of this construction stems from t wo follo wing lemmas. Lemma 15. L et G 1 and G 2 b e two b alanc e d genomes, Ma k e2I ( G 1 , G 2 ) = ( D , ω ) and S b e any set of disjoint 2-intervals of D . If we denote by W S the total weight of S , then the maximum matching ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) obtaine d by Max-W2IP to Adj ( S ) induc es at le ast W S adjac encies. Pr o of. F or eac h 2-in terv al D = ( I 1 , I 2 ) of S , w e ha v e matc hed the signed genes of I 1 and I 2 in the natur al wa y . Therefore, for eac h 2-int erv al D = ( I 1 , I 2 ) of S , w e obtain | I 1 | − 1 adja- cencies in ( G M 1 , G M 2 , M ) since I 1 and I 2 are iden tical int erv als. S ince the fin al greedy p art of Max-W2IP to Adj ( S ) d o es not d elete an y adjacency , we h a v e at least W S adjacencies in ( G M 1 , G M 2 , M ). ⊓ ⊔ Lemma 16. L et G 1 and G 2 b e two b alanc e d genomes, ( G M 1 , G M 2 , M ) b e a maximum matching of ( G 1 , G 2 ) , Mak e2I ( G 1 , G 2 ) = ( D , ω ) and W b e the numb er of adjac encies induc e d by ( G M 1 , G M 2 , M ) . Then ther e exists a sub set S ⊆ D of disjoint 2-i ntervals of total weig ht W . Pr o of. Denote b y n the size of G M 1 . Consider any factorization G M 1 = s 1 s 2 . . . s p suc h that, for eac h 1 ≤ i < p , s i and s i +1 are separated by one breakp oint and no b reakp oint app ears in s i , 1 ≤ i ≤ p . Therefore, there exists p − 1 breakp oin ts b et w een G M 1 and G M 2 , and hence n − p adjacencies b et w een G M 1 and G M 2 . T o eac h substring s i of the factorization of G M 1 corresp onds a su b string t i in G M 2 suc h that s i and t i are id en tical. Moreo ver, eac h substring s i of size l i , 1 ≤ i ≤ p , con tains l i − 1 adjacencies. W e construct the 2-in terv al set S as th e un ion of D i = ( ˆ s i , ˆ t i ), 1 ≤ i ≤ p , w here ˆ s i (resp. 25 ˆ t i ) is the interv al obtained f r om s i (resp. t i ). The f actorizat ion of G M 1 implies that the constructed 2-in terv als are disjoint, and hence the total weig ht of S is P p i =1 ( l i − 1) = P p i =1 l i − P p i =1 1 = n − p = W . ⊓ ⊔ W e no w d escrib e Algorithm Appro xAdj and then pr o v e it to b e a 4-approxi mation algorithm for Max-Adj . Algorithm 1 App ro xA dj Require: Two balanced genomes G 1 and G 2 . Ensure: A maximum matc hing ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ). – Let Make2I ( G 1 , G 2 ) = ( D , ω ). – Invok e the 4-approximation algorithm of Crochemore et al . [14] to obtain a set of disjoint 2-interv als S ⊆ D . – Construct t h e maximal matching ( G M 1 , G M 2 , M ) = Max-W2IP to Adj ( S ). Theorem 9. Algorithm Appro xAdj is a 4 -appr oximation algorithm for M ax-Adj . Pr o of. According to Lemmas 15 and 16, there exists a maximum matc hin g ( G M 1 , G M 2 , M ) of ( G 1 , G 2 ) that in duces W adjacencies iff there exists a subset of disjoin t 2-interv als S ⊆ D w ith total w eigh t W . Therefore, an y appro ximation ratio for Max-W2IP implies the same appr o x im a- tion ratio for Max-Adj . In [14], a 4-appr o x im ation algorithm is p rop osed for Max -W2IP . Hence, Algorithm A pp ro x Adj is a 4-appro ximation algo rithm for Max-Adj . ⊓ ⊔ 6 Conclusions and future work In this p ap er, we h av e fir st giv en new approxima tion complexit y results for seve ral optimization problems in genomic rearr an gement. W e fo cus ed on conserv ed interv als, common interv als and breakp oints, and w e to ok into accoun t the pr esence of du plicates. W e restricted our pro ofs to cases where one genome con tains no duplicates and the other con tains n o more than t wo o ccurrences of eac h gene. With this assump tion, we pro v ed that the p roblems consisting in computing an ex- emplarization (resp. an in termediate matc h ing, a maxim um matc hin g) optimizing any of the three ab o v e mentio ned measures is APX –hard, thus extending the results of [7, 10, 13 ]. In a second p art of the p ap er, w e hav e fo cused on the ZEBD (resp. ZIBD , ZMBD ) problems, where the q u estion is whether there exists an exemplarization (resp. int ermediate matc hing, maxim um matc hing) th at in- duces zero breakp oin t. W e h a v e extended a r esult fr om [13] by sho wing that Z EBD is NP –complete ev en for instances of t y p e (2 , k ), where k is u n b oun ded. W e also ha v e noted that ZEBD and ZIBD are equiv alen t p roblems, and sh o wn that ZMBD is in P . Finally , w e ga ve sev eral appr o ximation algorithms for computing the maximum num b er of adjacencies of tw o balanced genomes un der the maximum matching mo del. Th e appr o xim ation ratios w e get are 1.1442 for instances of t yp e (2 , 2), 3 for instances of t y p e (3 , 3) and 4 in the general case. Concerning the latter result, we note that the appro ximation ratio we obtain is constan t, eve n when the num b er of o ccurrences in genomes is unboun ded. Ho wev er, several problems r emain unsolve d. In particular, concerning appro ximation algorithms, virtually nothing is kno wn (i) in the case of unbalanced genomes and (ii) in the exemplar and in termediate mo d els. In deed, all the existing results (see for instance [17, 19] for the num b er of breakp oints), in clud ing ou r s, fo cus on the maximum matc hing problem for balanced genomes, 26 whic h imp lies th at no gene is d eleted fr om genomes G 1 and G 2 . Now, if we allo w genes to b e deleted, th e prob lem seems muc h more difficult to tac kle. Finally , we w ould lik e to recall th e follo win g op en p r oblem from [11]: what is the complexity of ZEBD f or instances of t yp e (2 , 2) ? References 1. P . Alimon ti and V . Kann . Some APX-completeness results for cubic graphs. The or etic al Computer Scienc e , 237(1-2):123 –134, 2000. 2. S . Angibaud, G. F ertin, I. Ru su , A . Th ´ evenin, and S. Vialette. Efficient to ols for computing the num b er of breakp oints and th e num b er of adjacencies b etw een tw o genomes with duplicate genes. Journal of Computational Biolo gy , 2008. S ubmitted. 3. S . Angibaud, G. F ertin, I. Rusu, and S. Vialette. A general framew ork for comput in g rearrangement distances b etw een genomes with du plicates. Journal of Computational Biolo gy , 14(4):379 –393, 2007. 4. V . Bafna and P . Pe vzner. Sorting by reversa ls: genome rearrangemen ts in plant organelles and evolutionary history of X chromo some. Mole cular Biolo gy and Evolution , pages 239– 246, 1995. 5. P . Berman and T. F ujito. Approximating indep endent sets in d egree 3 graphs. I n Pr o c. 4th Workshop on Algor ithms and Data Structur es , vo lume 955 of LNCS , pages 449–460. Sp ringer, 1995. 6. G. Blin and R. Rizzi. Conserv ed interv al distance computation b etw een non-trivial genomes. In Pr o c. COCOON 2005 , vo lume 3595 of LNCS , p ages 22–31. S pringer, 2005. 7. D . Bryan t. The complexit y of calculating exemplar distances. In Comp ar ative Genomics: Empiric al and Analytic al Appr o aches to Gen e Or der Dynamics, Map A lignement, and the Evolution of Gene F amilies , pages 207–212. Kluw er Academic Publisher, 2000. 8. M.-S . Chang and F.-H. W ang. Efficien t algorithms for the maximum w eigh t clique and maxim um weigh t inde- p endent set problems on p ermutatio n graphs. Information Pr o c essing L etter s , 43(6):293–2 95, 1992. 9. M. Charik ar, K. Mak arychev, and Y. Mak arychev. Near-optimal algorithms for maxim um constraint satisfaction problems. In SODA ’07: Pr o c e e dings of the eighte enth annual ACM-SIAM symp osium on Discr ete algorithms , pages 62–68, Philadelphia, P A, USA, 2007. So ciety for Industrial and Applied Mathematics. 10. C. Chauve, G. F ertin, R. Rizzi, and S. Vialette. Genomes con taining duplicates are hard t o compare. In Pr o c. IWBRA 2006 , volume 3992 of LNCS , pages 783–790. Springer, 2006. 11. Z. Chen, R. F owler , B. F u, and B. Zhu. Low er b ounds on the approximation of t he exemplar conserved in terv al distance problem of genomes. In Pr o c COCOON 2006 , v olume 4112 of LNCS , pages 245–254. S pringer, 2006. 12. Z. Chen, B. F u, J. Xu, B. Y ang, Z. Zh ao, and B. Zh u. Non-breaking similarit y of genomes with gene rep etitions. In Pr o c. CPM 2007 , volume 4580 of LNCS , pages 119–130 , 2007. 13. Z. Chen, B. F u, and B. Zhu. The approximabili ty of the exemplar b reakp oint distance problem. In Pr o c. AAIM 2006 , vo lume 4041 of LNCS , p ages 291–302. Springer, 2006. 14. M. Crochemore, D . Hermelin, G. M. Landau, and S. Vialette. Ap p ro ximating the 2-interv al pattern problem. In Pr o c. ESA 2005 , volume 366 9 of LNCS , pages 426–437 . Springer, 2005. 15. H . F ernau. Pa rameterized algorithms: A graph-theoretic approac h. Habilitationsschrif t, 2005. 16. M.R. Garey and D.S. Johnson. Com puters and Intr actability: a guide to the the ory of N P-c ompleteness . W .H. F ree- man, San F ranciso, 1979. 17. A . Goldstein, P . Kolman, and Z. Zheng. Minimum common string partition problem: Hardn ess and appro xima- tions. In Pr o c. ISAAC 2004 , v olume 3341 of LNCS , pages 473–484. S pringer, 2004. 18. M.C. Golumbic. Algor ithmic Gr aph The ory and Perfe ct Gr aphs . Academic Press, New Y ork, 1980. 19. P . Kolman and T. W ale ´ n. Reversal distance for strings with duplicates: Linear time appro ximation u sing hittin g set. I n Pr o c. W AOA 2006 , volume 436 8 of LNCS , pages 279– 289. Springer, 2006 . 20. W. Li, Z. Gu , H. W ang, and A. N ek rutenko. Ev olutionary analysis of th e h uman genome. Natur e , 409:847–849, 2001. 21. M. Marron, K. M. Swenson, and B. M. E. Moret. Genomic distances under d eletions and in sertions. The or etic al Computer Scienc e , 325(3):347–3 60, 2004. 22. C. Pa padimitriou and M. Y annak ak is. Op timization, approximation, and complexity classes. Journal of Computer and System Scienc es , 43(3):425– 440, 1991. 23. D. Sankoff. Genome rearrangemen t with gene families. Bioi nformatics , 15(11):90 9–917, 1999. 24. D. Sankoff and L. Haque. P ow er b o osts for cluster tests. In Pr o c. RECOMB-CG 2005 , v olume 3678 of LNCS , pages 121–13 0. Springer, 2005. 27 25. J. T ang and B. M. E. Moret. Phyloge netic reconstruction from gene-rearrangemen t data with un equal gene conten t. In Pr o c. W ADS 2003 , v olume 2748 of LNCS , pages 37–46. Sp rin ger, 2003. 28
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment