Clustering and Feature Selection using Sparse Principal Component Analysis
In this paper, we study the application of sparse principal component analysis (PCA) to clustering and feature selection problems. Sparse PCA seeks sparse factors, or linear combinations of the data variables, explaining a maximum amount of variance …
Authors: Ronny Luss, Alex, re dAspremont
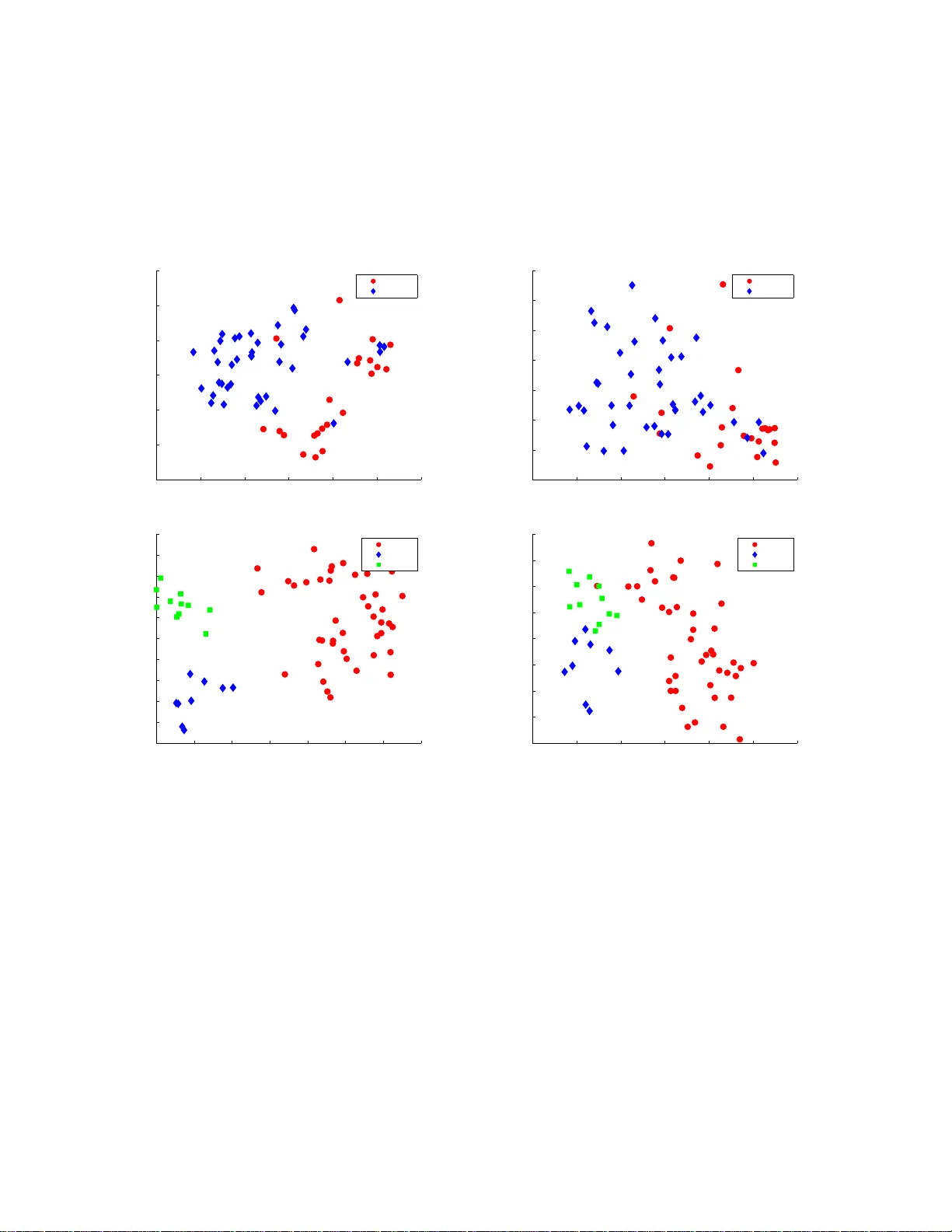
Clustering and Feature Selection using Sparse Principal Component Analysis Ronny Luss ∗ Ale xandre d’Aspremon t † No vember 26, 2024 Abstract In this paper , we study the applic ation of sp arse principal compone nt analys is (PCA ) to cluste ring and feature selecti on problems. Sparse PC A seeks sparse factors, or linear com- binati ons of the data v ariable s, explainin g a maximum amount of varian ce in the data while ha ving on ly a limited number of nonz ero coef fi cients . PCA i s ofte n used as a simple clusteri ng techni que and sparse factors allo w us here to interp ret the clusters in terms of a reduced set of varia bles. W e begin w ith a bri ef introd uction and moti v ation on sparse PCA and detail our implementa tion of the algorithm in d’Aspremont et al. (200 5). W e then apply these resul ts to some classic cluster ing and featu re selection pro blems arising in biology . Keyw ords: Sparse principal com ponent analys is, semi definite programmi ng, clustering, fea- ture selection. 1 1 Introduction This paper focuses o n application s of sp arse principal component analy sis to clustering and fea- ture selection problem s, with a particular focus on gene expression data analysi s. Sparse met hods hav e had a significant im pact in many areas of statistics, in parti cular regression and classification (see [CT05], [DT05] and [V ap95] among others). As in these areas, our moti vation for de veloping sparse mult iv ariate visualizatio n tool s is th e potential of these meth ods for yieldi ng statistical re- sults that are bot h more interpretable and more robust than classical analyses, while giving up little statistical ef ficiency . Principal com ponent analysi s (PCA) is a classic tool for analyzin g large scale multivariate data. It seeks lin ear combin ations of the data variables (often called factors or principal com ponents) that capture a maximum amount of variance . Numerically , PCA only amounts to computi ng a few ∗ ORFE Departmen t, Princeton Uni versity , Princeton, NJ 08544. rluss@princeton. edu † ORFE Departmen t, Princeton Uni versity , Princeton, NJ 08544. aspremon@princet on.edu 1 Mathematical Subject Classification: 90C90, 62H25, 65K05 . 1 leading eigen vectors of the data’ s covariance m atrix, so it can be applied to very large scale data sets. One of the key shortcom ings o f PCA howe ver is that these factors are linear com binations of al l va riables; that is, all factor coefficients (or loadi ngs) are non-zero. This m eans th at whi le PCA facilitates mod el interpretation and visualization b y concentrating the informati on in a few key factors, the f actors themselves are still constructed using all observed v ariables. In many applications of PCA, the coordinate axes hav e a di rect physi cal interpretation; in finance or biology for e xample, each axis might correspond to a sp ecific fina ncial asset o r gene. In such cases, having only a few nonzero coefficients i n the principal components would greatly im prove the relev ance and i nterpretability of the factors. In sparse PCA, we seek a trade-of f between the two go als of e xpr essive power (explaining m ost of th e variance or inform ation i n the data) and interpr etability (making sure that t he factors in volve only a few coordinate axes or variables). When PCA is used as a clusterin g t ool, sparse factors wi ll allow us to identify the clusters with the action of only a fe w variables. Earlier methods to produ ce sparse factors incl ude Cadima and Joll iffe [CJ95] where the load- ings w ith s mallest absolu te value are t hresholded to zero and n oncon ve x algorith ms called SCoT - LASS by [JTU03], SLRA [ZZS02, ZZS04] and SPCA b y [ZHT0 6]. This last method works by writing PCA as a regression-type optim ization probl em and applies LASSO [Tib96 ], a penaliza- tion technique based o n the l 1 norm. V ery recently , [MW A06b] and [MW A06a] also proposed a greedy app roach which seeks global ly opti mal solu tions on small problems and uses a greedy method to approximate the solut ion of larger ones. In what follo ws, we giv e a brief introductio n t o the relaxation of this problem in [dEGJL07] and describe ho w this sm ooth optimization algorithm was imp lemented. The m ost expensi ve numerical s tep in th is algo rithm is the computati on of the gradient as a matrix e xponential and our key numerical contrib ution here is to sho w t hat using only a partial eigen v alue decompos ition of the current it erate can produce a sufficiently precise gradi- ent approxim ation while drastically i mproving computational effic iency . W e then show on classic gene expression data sets that using sparse PCA as a sim ple clus tering tool isol ates very relev ant genes compared to other techniques such as recursiv e feature e limin ation or ranking. The paper is or gani zed as follows. In Section 2, we begin with a brief introduction and motiv a- tion on sparse PCA and detail our implement ation of the algorithm in a numerical toolbox called DSPCA, which is av ai lable for d ownload on the authors’ websi tes. In Section 3, we describe the application of sparse PCA to clustering and feature selection on gene expression data. 2 Algorithm and im plementa tion In this s ection, we begin by int roducing the sparse PCA problem and the corresponding semid ef- inite relaxati on derived in [dEGJL0 7]. W e then di scuss h ow to use a parti al eigen value decompo- sition of the current it erate to produ ce a sufficiently precise g radient approximation and im prove computational ef ficiency . 2 2.1 Sparse PCA & Semidefinite Relaxation Giv en a cov ariance matrix A ∈ S n , where S n is the s et of sym metric matrices of dim ension n , the problem o f findi ng a sparse factor which explains a maximum amo unt of variance in the d ata can be written: maximize x T Ax subject to k x k 2 = 1 Card ( x ) ≤ k , (1) in t he variable x ∈ R n where Card ( x ) denotes the cardinality of x and k > 0 is a parameter controlling this cardinality . Computing sparse factors with maxim um variance is a combin atorial problem and is num erically hard and [dEGJL0 7] use semi definite relaxation t echniques to compute approximate solutions ef ficiently by solving the following con ve x program: maximize T r ( AX ) subject to T r ( X ) = 1 1 T | X | 1 ≤ k X 0 , (2) which is a semidefinite program in the var iable X ∈ S n , where 1 T | X | 1 = P ij | X ij | can be seen as a con ve x lower bound on t he function Card ( X ) . When th e so lution of the above problem has rank one, we have X = xx T where x is an approx imate soluti on to (1). When the sol ution of th is relaxation is not rank one, we use the leading eigen vector of X as a principal component. While small inst ances of problem (2) can be solved effi ciently using interior point semidefinite programming solvers su ch as SEDUMI by [Stu99], the O ( n 2 ) li near constraint s make these solvers ineffi cient for reasonably l ar ge ins tances. Furthermore, interior poi nt m ethods are geared towards solving small problems with high precisi on requirement s, while here we need to sol ve very large instances with relatively low precisio n. In [dEGJL07] it was shown that a smo othing t echnique due to [Nes05] could be applied to probl em (2) to get a complexity esti mate o f O ( n 4 √ log n/ǫ ) and a much l ower memory cost per iteratio n. The ke y numerical s tep in this algorithm is the computation of a sm ooth approximati on of problem (2) and t he gradient of the objective, wh ich amounts to computing a matrix exponential. 2.2 Implementation Again, giv en a cova riance matrix A ∈ S n , the DSPCA code solves a penalized formulati on o f problem (2), written as: maximize T r ( AX ) − ρ 1 T | X | 1 subject to T r ( X ) = 1 X 0 , (3) in the va riable X ∈ S n . The dual of this program can be written as: minimize f ( U ) = λ max ( A + U ) subject to | U ij | ≤ ρ. (4) 3 in the variable U ∈ S n . Th e algorithm in [dEGJL07] regularizes the objective f ( U ) in (4), replac- ing it by the smooth (i.e. with Lipschit z continuous gradient) uniform approximation: f µ ( U ) = µ log ( T r exp(( A + U ) /µ )) − µ log n, whose gradient can be computed explicitly as: ∇ f µ ( U ) := exp (( A + U ) /µ ) / ( T r exp (( A + U ) /µ )) . Follo wing [Nes83], solving the smooth problem: min { U ∈ S n , | U ij |≤ ρ } f µ ( U ) with µ = ǫ/ 2 log( n ) th en produces an ǫ -a pproximate sol ution to (3) and requires O ρ n √ log n ǫ (5) iterations. The main s tep at each iteratio n is computing the matrix exponential exp(( A + U ) / µ ) . This is a classic problem in linear algebra (see [MVL03] for a comprehensive survey) and in what follows, we detail three dif ferent methods implemented in DSPC A and their relativ e performance. Full eigen value decomposition An exact compu tation of the matrix exponential can be done through a full eigen value d ecomposition of the matrix. Giv en the spectral decompo sition V D V T of a m atrix A where the columns of V are t he eigen vectors and D is a diagonal matri x compris ed of the eigen v alues ( d i ) n i =1 , the matrix exponential can be computed as: exp( A ) = V H V T , where H is the diagon al matrix with ( h i = e d i ) n i =1 on its diago nal. While thi s i s a sim ple procedure, it is also relativ ely inef ficient. Pad ´ e approximation The next metho d implem ented in DSPCA is called Pad ´ e approxim ation and approximates the exponential by a rational function. The (p,q) P ad ´ e approximation for exp( A ) is defined by (see [MVL03]): R pq ( A ) = [ D pq ] − 1 N pq ( A ) , (6) where N pq ( A ) = p X j =0 ( p + q − j )! p ! ( p + q )! j !( p − j )! A j and D pq ( A ) = q X j =0 ( p + q − j )! q ! ( p + q )! j !( q − j )! ( − A ) j . Here p and q control the degree and precisi on of the approximation and we set p = q = 6 (we set p = q i n practice due to computati onal issues; see [MVL03]). The approximation is only valid in a sm all neighborhood o f zero, whi ch means t hat we need to scale down the matrix before 4 approximating its exponential using (6), and then scale it back to its origin al size. This sca ling and squaring can be done efficiently usi ng th e p roperty that e A = ( e A m ) m . W e first s cale th e m atrix A so that 1 m k A k ≤ 10 − 6 and find the smallest integer s such that this is true for m = 2 s . W e then use the Pad ´ e approxim ation to compute e A , and simply square the result s t imes to scale it back. Pad ´ e approximation only requires computin g one matrix in version and se veral matrix products which can be done very efficiently . Ho wev er , if n or s get somewhat lar ge, scaling and squaring can be cost ly , in which case a full eigen v alue decomp osition has better performance. While Pad ´ e approximations appear to be the current m ethod of choice for comput ing matrix exponentials (see [MVL03]), it does not per form as well as e xpected on our problems compared to partial eigen va lue decompositio n discuss ed below , because o f the particul ar structure of our optim ization p roblem. Numerical results illustrating this issue are detailed in the last section. Partial eigen value decomposition The first two classi c methods we described for comput ing the exponential of a m atrix are both geared towards produci ng a s olution up to (or clos e t o) machine precision. In most of the sparse P CA problems we solve here, the target precision for the a lgorithm is of the order 10 − 1 . Compu ting the gradi ent up to machine precisio n in this context is u nneces- sarily costly . In fact, [d’A05] shows that the op timal con vergence of the alg orithm in [Nes0 5] can be achieved using approximate gradient va lues ˜ ∇ f µ ( U ) , provided the error satisfies t he following condition: |h ˜ ∇ f µ ( U ) − ∇ f µ ( U ) , Y i| ≤ δ, | U ij | , | Y ij | ≤ ρ, i, j = 1 , . . . , n, (7) where δ is a parameter con trolling the approxim ation error . In practice, thi s m eans that we on ly need to compute a few leading eigen values of t he m atrix ( A + U ) /µ t o get a sufficient gradient approximation. More precisely , if we denote by λ ∈ R n the eigen values of ( A + U ) /µ , condition (7) can be used to show th at, to get con ver gence, we need only comput e the j l ar gest eigen values with j such that: ( n − j ) e λ j q P j i =1 e 2 λ i ( P j i =1 e λ i ) 2 + √ n − j e λ j P j i =1 e λ i ≤ δ ρn . (8) The terms o n the left si de decrease as j increases meanin g the cond ition is satisfied by increasing the number of eigenv alues used. Computing the j largest eigen values of a matrix can be done very ef ficiently usin g packages such as ARP A CK if j << n . When j becomes lar ge, the algorithm switches t o full eigen v alue decomp osition. Finally , t he leading ei gen values tend to coal esce close to the optim um, potentially increasing t he nu mber o f eigen values required at each iteration (see [Pat98] for example) but th is phenom enon does no t seem to appear at the low precision tar gets considered here. W e detail empirical results on the performance of thi s technique in the following sections illustratin g ho w this technique clearly do minates the two others for lar ge scale problems. 2.3 Comparison with interior point solvers W e giv e an example illus trating the necessity of a lar g e-scale solver using a smooth approxim ation to problem (2). The Sedumi imp lementation to problem (2) runs out of memory for problem sizes lar ger than 60 , so we comp are the example on very sm all dim ensions against our implementati on 5 with partial eigen va lue decomposition (DSPCA). The covariance matrix is formed using colon cancer gene expression dat a detailed in the following section. T able 1 shows runni ng tim es for DSPCA and S edumi on for various (small) problem dimensio ns. DSPCA clearly beats the interior point so lver in computational tim e whi le achieving comparable precision (m easured as the per- centage of v ariance explained by the sparse fa ctor). For refere nce, we sho w ho w much variation is explained by the leading principal component. The decrease i n variance using Sedumi and DS PCA represents the cost of sparsity here. Dim. NonZeros DSPCA time DSPCA V ar . Sedumi time Sedumi V ar . PCA V ar . 10 3 0.08 42.88 % 0.39 43.00 % 49.36 % 20 8 0.27 45.67 % 5.29 47.50 % 52.58 % 30 11 0.99 40.74 % 41.9 9 42.44 % 51.13 % 40 14 4.55 40.26 % 229 .44 41.89 % 52.40 % 50 15 7.52 36.76 % 903 .36 38.53 % 50.24 % T able 1: CPU time (in seconds) and explained varian ce for Sedumi and smooth optimiza- tion with par tial ei gen va lue decomposi tions (DSPC A). V ar measures th e p ercenta ge of to tal v ariance ex plained by the sparse factors . DS PCA significantly outperfor ms in terms o f CPU time while reachi ng comparable varia nce targe ts. 3 Clusterin g and F eature Select i o n In this section, we use our code for sparse P CA (DSPCA), to analyze lar ge sets of gene expression data and we discus s applications of this technique to clustering and feature selectio n. PCA is very often used as a simple tool for data visualization and clustering (see [SSR06] for a recent a nalysis), here sparse factors allo w us to interpret the low dimensional projection of the data in terms of only a fe w variables. 3.1 Gene expr ession data W e first test the performance of DSPCA on cov ariance matrices generated from a gene e xpression data set from [ABN + 99] o n colon cancer . The data set is com prised of 62 ti ssue sampl es, 22 from normal colon tissues and 40 f rom cancerous colon tissues with 2000 genes measured in e ach sample. W e p reprocess the data by taking the log o f all data intensit ies and then normalize the log data of each sam ple to be mean zero and standard deviation one, which is a classic procedure t o minimize experimental ef fects (see [HK05]). Since the semidefinit e programs prod uced by these data sets are relativ ely large (with n = 1000 after preprocessing ), we first test the n umerical performance of the algorit hm and run the cod e on increasingly l ar ge p roblems us ing each of the th ree method s described in section 2. 2 (ful l eigen- value d ecomposition , Pad ´ e approximati on and partial eigen v alue decompositio n). Theoretically , 6 the p erformance increase of us ing partial, rather than full, eigen value decompos itions shoul d be substantial when only a fe w eigen v alues are required. In practice there is overhead due to t he necessity of testing cond ition (8) iterativ ely . Figu re 1 depicts the results of these tests on a 3.0 GHz CPU in a loglog plot of run time (in seconds) versus probl em d imension (on t he left). W e plot the av erage number of eigen va lues required by condition (8) versus problem d imension (on the right ), with d ashed lin es at plus and min us one s tandard deviation. W e cannot include i nterior point algorit hms in this comp arison because memory problem s occur for d imensions g reater than 50. 10 2 10 3 10 0 10 1 10 2 10 3 Exact Pade Approximation Partial Eig Approximation CPU time (secon ds) Dimension 100 200 300 400 500 600 700 800 900 1000 0 1 2 3 4 5 6 7 8 9 10 % eigs. require d Dimension Figur e 1: Performance of smooth optimization using exact gradient, Pad ´ e approximat ions and partial eig en valu e decompositi on. CP U time v ersus pro blem dimension (left). A verage percen tage of eigen v alues required versus proble m dimensio n (right) for partial eigen valu e decompo sition, w ith dash ed lines at plus and minus one standard devi ation. In th ese test s, partial eigen value is mo re than three times faster than the other methods for lar ge problem sizes, meaning the approxi mate gradient is the dominatin g alternative compared t o the exact gradient. The plot of the a verage percentage of eigen va lues for each test s hows t hat on a verage on ly 2-3% o f the eigen values are necessary . Give n the gain performance, t he main lim- itation becomes m emory; th e parti al eigen value i mplementatio n has memory allocation problems at di mensions 1900 and larger on a com puter with 3 .62 Gb RAM. Notice also that in ou r exper- iments, computing th e matrix exponential using Pad ´ e approxim ation is slower than performin g a full eigen v alue decomposition, which reflects the high numerical cost of m atrix multi plications necessary for scaling and squaring in Pad ´ e approxim ations. Next, we test the impact of the sparsity targe t on compu tational complexity . W e observe in (5) that the num ber of iteration s required b y the algorithm scales l inearly with ρ . Furthermore, as ρ increases, conditi on (8) means that more eigen va lues are required to satis fy to prove c on ver gence, which increases the computational ef fort. Figure 2 shows how computing time (left) and d uality gap (right ) vary with the number of eigen values required at each iteration (sho wn as the percentage of total eigen v alues). The four 7 tests are all of dimens ion 1000 and v ary in degrees of sparsity; for a fixed data set at a fixed point i n t he algo rithm, m ore sparsi ty (ie. higher ρ and less genes) requires more eigen values as noted above. The in crease in required eigen v alues is much more prono unced when viewed as t he algorithm proceeds, measured by the increasing computin g time (left) and decreasing duality gap (right). More eigen values are also required as the current i terate gets c loser to the op timal soluti on. Note that, to reduce overhead, th e code does not allow the number of required eigen va lues to decrease from one iteration to the ne xt. Figure 2 shows as was expected that computational cost of an iteration increases with both tar get precision and target sparsity . 0 50 100 150 200 250 300 350 0 2 4 6 8 10 12 14 14 Genes, Rho = 32 16 Genes, Rho = 30 21 Genes, Rho = 28 28 Genes, Rho = 24 % eigs. require d CPU time (secon ds) 0 0.5 1 1.5 2 2.5 x 10 4 0 2 4 6 8 10 12 14 14 Genes, Rho = 32 16 Genes, Rho = 30 21 Genes, Rho = 28 28 Genes, Rho = 24 % eigs. require d Duality gap Figur e 2: Percentage of eigen va lues requi red versus CPU time (left), and percenta ge of eigen value s require d versus duality gap (right). T he computatio nal cost of an iteration increa ses w ith both tar get precision and targ et sparsity . 3.2 Clustering In this section, w e com pare t he clu stering (class discovery) performance of sparse PCA to t hat of standard PCA. W e analyze the col on cancer data set of [ABN + 99] as well as a lymph oma data set from [AED + 00]. The lymphoma data set con sists of 3 classes of lymphoma denot ed DLCL (diffuse lar ge B-cell lympho ma), FL (foll icular lymphom a) and CL (chronic lymp hocytic leukemia). The top two plo ts of Figure 3 displ ay the clust ering effec ts of using two factors on the col on cancer data while the bott om two pl ots of Figure 3 disp lay t he resul ts on the lymp homa data. On both data sets, clust ers are represented usi ng PCA factors on the left plot s and sparse factors from DSPCA on the rig ht plots. For the colon cancer data, t he second factor has greater power in predicting the class of eac h sample, while for the lym phoma data, the first factor classifies DLCL and the sec ond factor differentiates between CL and FL. In these examples, we observe that DSPCA maintains good cluster separation while requiring far fewer genes: a total of 13 instead of 1000 for the colon cancer data, and 108 genes instead of 1000 for the lymphoma data set. While t he clustering remains visuall y clear , we now analyze and quantify the quality of the 8 −15 −10 −5 0 5 10 15 −15 −10 −5 0 5 10 15 Normal Cancer Factor thre e (500 genes) Factor t wo (500 genes) −6 −4 −2 0 2 4 6 −3 −2 −1 0 1 2 3 4 Normal Cancer Factor t hree (1 gene) Factor two (12 genes) −30 −20 −10 0 10 20 30 40 −35 −30 −25 −20 −15 −10 −5 0 5 10 15 DLCL FL CL Factor t wo (500 genes) Factor o ne (500 genes) −20 −10 0 10 20 30 40 −25 −20 −15 −10 −5 0 5 10 15 DLCL FL CL Factor two (53 genes) Factor one (55 gen es) Figur e 3: Clustering: The top two graphs display the results on the colon cancer data set using PCA (left) and DSPCA (righ t). Normal patients a re red circles and cance r patie nts are blue diamonds. The bottom two graphs display the results on the lymphoma data set using PCA (left) and DSPCA (right ). For lymphoma, we denote dif fuse l ar ge B-cell ly mphoma as DLCL (red circles), follicular lymphoma as FL (blue diamonds), and chron ic lymphoc ytic leukae mia as C L (green square s). 9 clusters deri ved from PCA and DSPCA num erically us ing the Rand index. W e first clu ster th e data (after reducing to two dimensi ons) usi ng K -means clu stering, and then use the Rand index to compare the partitions obtained from PCA and DSPCA to the true partiti ons. The Rand index measures the similarity between two partitions X and Y and is computed as the ratio R ( X , Y ) = p + q n 2 where p is the number of pairs of elements that are in the same cluster in both p artitions X and Y (correct pairs), q is the number of pairs o f element s in diffe rent clust ers in both X and Y (error pairs), and n 2 is the total nu mber of element pairs. The Rand index for va rying levels of sparsity is pl otted in Figure 4. The Rand ind ex of standard PCA is .6 54 for colon cancer (.804 for lymphoma) as m arked in Figure 4. The Rand in dex for the DSPCA factors of colon cancer usin g 13 genes i s .6 69 and i s t he leftm ost po int above the PCA line. This sh ows that clusters derive d using sparse factors can achieve equ iv alent performance to clusters derived from dense factors. Howe ver , DSPCA on the lymphom a data does not achieve a high Rand index wit h very sparse factors and it takes about 50 genes per factor to get good clusters. 0 10 20 30 40 50 0.45 0.5 0.55 0.6 0.65 0.7 0.75 PCA DSPCA Rand inde x Number of nonze ro coefficien ts 0 100 200 300 400 500 600 0.45 0.5 0.55 0.6 0.65 0.7 0.75 0.8 0.85 0.9 PCA DSPCA Rand inde x Number of nonze ro coefficien ts Figur e 4: Rand inde x versus sparsi ty: colon cancer (left) & lymphoma (right). For lymphom a, we can also look at another m easure of cluster validity . W e measure the impact of sparsity on t he separation betw een the true clusters, d efined as th e distance between th e cluster centers. Figure 5 s hows how this separation varies with the sparsity of the factors. The lymp homa clusters wi th 108 genes have a separation of 63, after whi ch separation drops sharply . Noti ce that the s eparation of CL and FL is very small to begin with and the sharp decrease in separation is mostly due to CL and FL getting closer to DLCL. 3.3 F eatur e s election Using the s parse factor analysis derive d above, our next obj ectiv e is to track the action of the selected genes. See [GWBV02] for an example il lustrating the poss ible relation of selected genes 10 0 100 200 300 400 500 600 0 20 40 60 80 100 120 CL−FL CL−DLCL DLCL−FL Sum Separatio n # of nonzero s Figur e 5: Separation vers us sparsity for the lymphoma data set. in this data set to colon cancer using Recursive Feature Elimination with Support V ector Machines (RFE-SVM) for feature selection. Another study in [HK05] comp ares RFE-SVM results on the colon cancer data set to those genes selected by the Rankgene software in [SMP + 03] on the colon cancer and lymphom a data sets. W e implemented RFE-SVM and determined the regularization p arameter using cross -va lidation. On the colon cancer data, the clust ering results show th at factor two contains t he most predictive power . T abl e 2 s hows 7 genes that appeared in the top 10 m ost important genes (ranked b y mag- nitude) in t he results of DSPCA and RFE-SVM. For comparison, we include ranks comp uted using another sparse PCA p ackage: SPCA from [ZHT 06] with a sparsity level equal to the result s of DSPCA (we use t he functi on spca for colon cancer and arrayspca for lym phoma, bot h from [ZH05]). DSPCA id entifies 2 genes from the top 10 genes o f RFE-SVM that are not identified by SPCA. After removing the true first comp onent which explains 58.1% of the variation, DSPCA, SPCA, and PCA explain 1.4%, 1.1%, and 7.2% of the remaining v ariatio n respecti vely . Since the first factor of the lymphoma DSPCA re sults classifies DLCL, we combine FL and CL into a single group for RFE-SVM and compare the results to f actor one of DSPCA. T able 3 shows 5 genes that appeared in the top 15 mos t impo rtant genes of all th ree feature selection methods. An additio nal 2 g enes were di scovere d by DSPCA and SPCA but not by RFE-SVM. DSPCA, SPCA, and PCA explain 13.5%, 9 .7%, and 28 .5% of th e variation respecti vely , providin g another example where DSPCA m aintains more of the clusterin g effects t han SPCA given equal sparsity . Both RFE-SVM and SPCA are faster than DS PCA, but RFE-SVM does not produce f actors in the sense of PCA and SPCA is noncon vex and has no con ver gence guarantees. 4 Conclusio n W e showed that efficient approximations of the gradient allowed large scale instances of the sp arse PCA relaxation algorithm in [dEGJ L07] to be solved at a m oderate comput ational cost. This 11 DSPCA RFE-SVM SPCA GAN Description 2 1 11 J 02854 Myosin re gulator light chain 2, smooth muscle isoform (huma n) 4 3 9 X86693 H.sapiens mRN A f or he vin like protein 5 6 1 T92451 T ropom yosin, fib roblast and epithelial muscle-typ e ( human) 7 4 5 H43887 Complement factor D precursor (H. sapiens) 8 9 N A H06524 Gelsolin precursor, plasm a (human) 9 2 N A M63391 Hum an desmin gene, complete cds 10 7 6 T47377 S-100P Protein (Human) . T able 2: Feature selection on colon cancer data set: C olumns 1-3 are respecti ve ranks. 7 genes (out of 2000) were in the top 10 list of genes for both DS PCA (fac tor two) and RFE-SVM (with C=.005). 5 of the genes were top genes in SPCA. DSPCA RFE-SVM SPCA GAN Description 1 1 1 GENE1636X *Fibron ectin 1; Clone=139 009 2 N A 6 GENE1637X Cyclin D2/KIAK000 2=overlaps with middle of KIAK0002 cDN A; Clone=359 412 3 14 2 GENE1641X *Fibron ectin 1; Clone=139 009 4 N A 9 GENE1638X MMP-2=Matrix metallopr oteinase 2=72 kD type IV collagenase precurso r=72 kD gelatinase=gelatinase A=TBE-1;Clone=3 2365 6 6 2 3 GENE1610X *Mig=Hum ig=chemo kine targeting T cells; Clone=8 10 3 4 GENE1648X *cathepsin B; Clone=297 219 15 8 8 GENE1647X *cathepsin B; Clone=261 517 T able 3: Feature selection on lymphoma data set: Columns 1-3 are respecti ve ranks. 5 genes (out of 4026) were in the top 15 list of genes for both D SPCA (factor one), RFE- SVM (classifyin g D LCL or not DL CL with C=.001), and SPC A (fac tor one). DSP CA and SPCA ha ve an addit ional 2 top genes not found in the top 15 genes of RF E-SVM. 12 allowed us to apply sparse PCA t o clust ering and feature selection o n two classic gene expression data s ets. In both cases, sparse PCA efficiently i solate relev ant genes whi le m aintaining m ost of the original clustering powe r of PCA. Acknowledge m ents W e are very g rateful to the organizers of th e BIRS workshop on Optimization and Engineering Applications where this work was presented. The autho rs would also like to ackno wledge support from grants NSF DMS-0625352, Eurocontrol C20083E/BM/05 and a gift from Google, Inc. References [ABN + 99] A. Alon, N. Barkai, D. A. Notterman, K. Gish, S . Ybarra, D . Mack, and A. J. Levine. Broad pattern s of gene expres sion rev ealed by cluste ring analysis of tumor and normal colon tissues probed by oligon ucleotid e arrays. Cell Biology , 96:67 45–67 50, 1999. [AED + 00] A. Alizadeh, M. Eisen, R. Davis , C. Ma , I . Lossos, and A. Rosenwald . Distinct typ es of dif fuse lar ge b-cell lymphoma identified by gene expre ssion profiling. Natur e , 403:503–51 1, 2000. [CJ95] J. Cadima and I. T . Jollif fe. Loadings and correlati ons in the interpreta tion of princi pal com- ponen ts. J ourna l of Applied Statist ics , 22:203–214 , 1995. [CT05] E. J. Cand ` es and T . T ao. Decoding by linear programming. Information Theory , IEEE T ra ns- action s on , 51(12):4203 –4215, 2005. [d’A05] A. d ’Aspremont. Smooth optimizati on with approx imate gradient . ArXiv: math.OC/05123 44 , 2005. [dEGJL07] A. d’Aspremont, L. El G haoui, M. I. Jordan, and G. R. G. Lanckriet . A direct formulation for sparse PCA using semidefinit e programming. SIAM Revie w , 49(3):43 4–448, 2007. [DT05] D. L. Don oho a nd J. T anner . Sparse n onne gati ve s olution s of underdete rmined linear equa tions by linear programming . Pr oc. of the National Academy of Sciences , 102(27): 9446–9 451, 2005. [GWBV02] I. Guyon, J. W eston, S. Barnhill, and V . V apnik. Gene selection for cancer classification using suppo rt vector machines. Machine Learning , 46:389– 422, 2002. [HK05] T . M. Huang and V . Kec man. Gene extract ion fo r cancer diagno sis by support v ector machines- an impro vemen t. Artificia l Intellige nce in Medicine , 35:185–19 4, 2005. [JTU03] I. T . J ollif fe, N.T . T rend afilov , and M. U ddin. A modified princip al compo nent tec hnique b ased on the LASSO. Jo urnal of Computation al and G raph ical Statist ics , 12:53 1–547, 2003. [MVL03] C. Moler and C. V an Loan. Nineteen dubious ways to compute the exponen tial of a matrix, twenty-fi v e years later . SIAM Revie w , 45(1):3–4 9, 2003. [MW A 06a] B. Moghaddam, Y . W eiss, and S. A vidan. Generaliz ed spectra l bounds for sparse L D A . In Intern ational Confer ence on M achine Learning , 2006. 13 [MW A 06b] B. Moghadd am, Y . W eiss, and S . A vidan. Spectral bounds for sparse PC A: Exact and greedy algori thms. Advance s in Neural Information Pr ocessing Systems , 18, 2006. [Nes83] Y . Nesterov . A method of solving a con ve x progr amming problem with con ver gence rate O (1 /k 2 ) . Sovi et Mathematics Doklady , 27(2) :372–3 76, 1983. [Nes05] Y . Nestero v . Smooth minimization of non-s mooth functi ons. Mathema tical Pr o gramming , 103(1 ):127–1 52, 2005. [Pat98 ] G. Pataki. On the rank of extreme matrices in semidefinite programs and the multiplicity of optimal eigen valu es. Mathematics of Opera tions Resear ch , 23(2):33 9–358, 1998. [SMP + 03] Y . Su, T . M . Murali, V . Pa vlo vic, M. Schaffe r , an d S. K asif. Rankgene: iden tification of diagno stic genes based on express ion data. Bioinformati cs , 19:1578–1 579, 2003. [SSR06] N. Srebro, G. S hakhn arovi ch, and S . Roweis. A n in vestig ation of computation al and informa- tional limits in gaussian mixture clustering. Pr ocee dings of the 23r d internat ional confer ence on Machine learning , pages 865–872 , 2006. [Stu99] J. Sturm. Using SED UMI 1.0x, a MA TLAB toolbox for optimizatio n over symmetric cones. Optimizatio n Methods and Softwar e , 11:625–65 3, 1999. [T ib96] R. Ti bshiran i. R egre ssion shrinkag e and selection via the LASSO. Journ al of the Royal statis- tical society , series B , 58(1): 267–28 8, 1996. [V ap95] V . V apn ik. The Natur e of Statistical Learning T heory . S pringe r , 1995. [ZH05] H. Z ou and T . Hastie. R egu larizatio n and var iable selection via the elastic net. Jour nal of the Royal Statistical Society Series B(Statistic al Methodolo gy) , 67(2):301– 320, 2005. [ZHT06] H. Zou, T . H astie, and R. Tib shirani . Sparse Principal C omponen t Analysis. J ourn al of Com- putati onal & Graphica l Statistics , 15(2):265– 286, 2006. [ZZS02] Z. Z hang, H. Z ha, and H . S imon. Low rank approximati ons with sparse facto rs I: basic algo- rithms and error analysis. SIAM journal on matrix analysis and its applic ations , 23(3): 706– 727, 2002. [ZZS04] Z. Zhang, H . Zha, and H . Simon. Low rank approximat ions with sparse fact ors II: penal- ized methods with discrete Newton –lik e iteration s. SIAM journal on matrix analysis and its applic ations , 25(4):901– 920, 2004. 14
Original Paper
Loading high-quality paper...
Comments & Academic Discussion
Loading comments...
Leave a Comment